Classification rationale
1
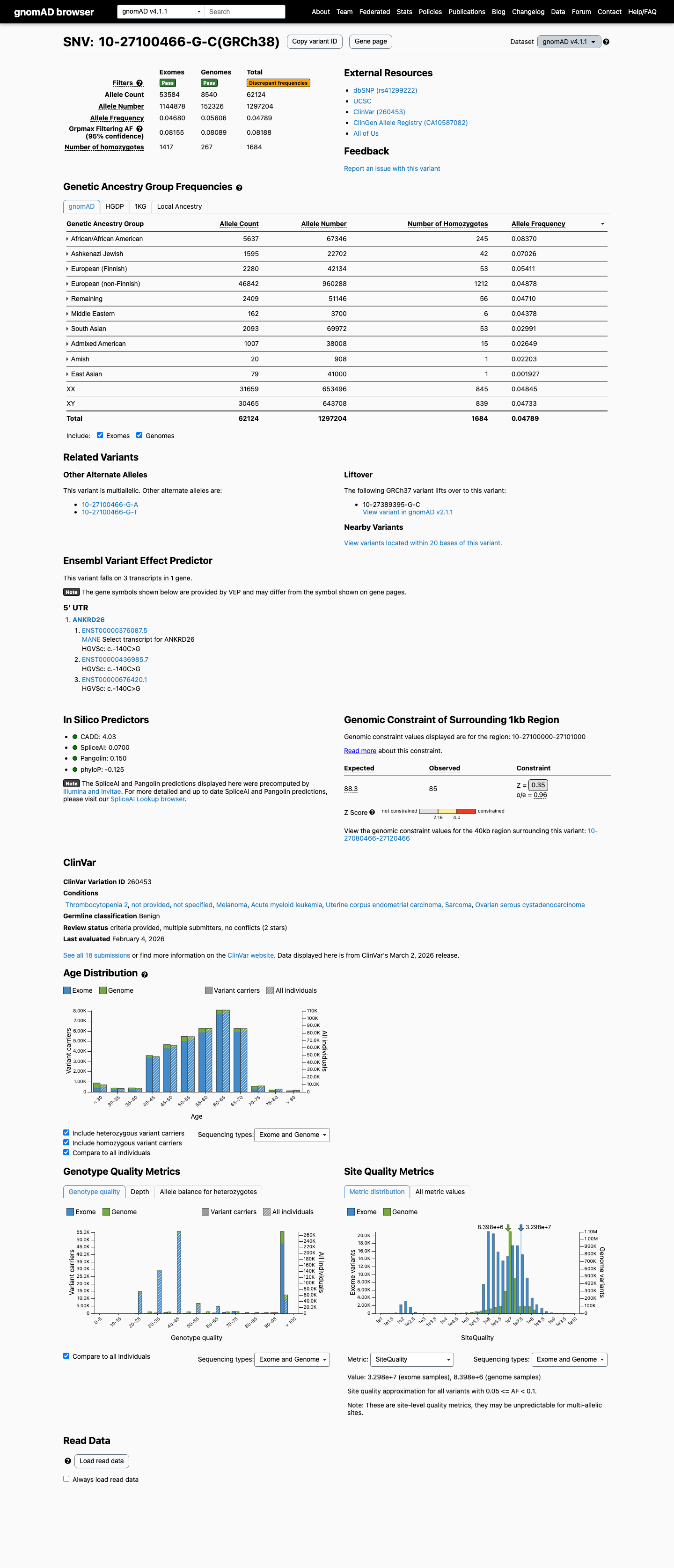
The ANKRD26 c.-140C>G (NP_055730.2:p.?) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar as Benign.
clinvar ↗2
This variant is common in population databases, with allele frequencies of 0.0640054 (6.40054%) in gnomAD v2.1 and 0.0478907 (4.78907%) in gnomAD v4.1, which are well above benign thresholds.
gnomad_v2 ↗ gnomad_v4 ↗3
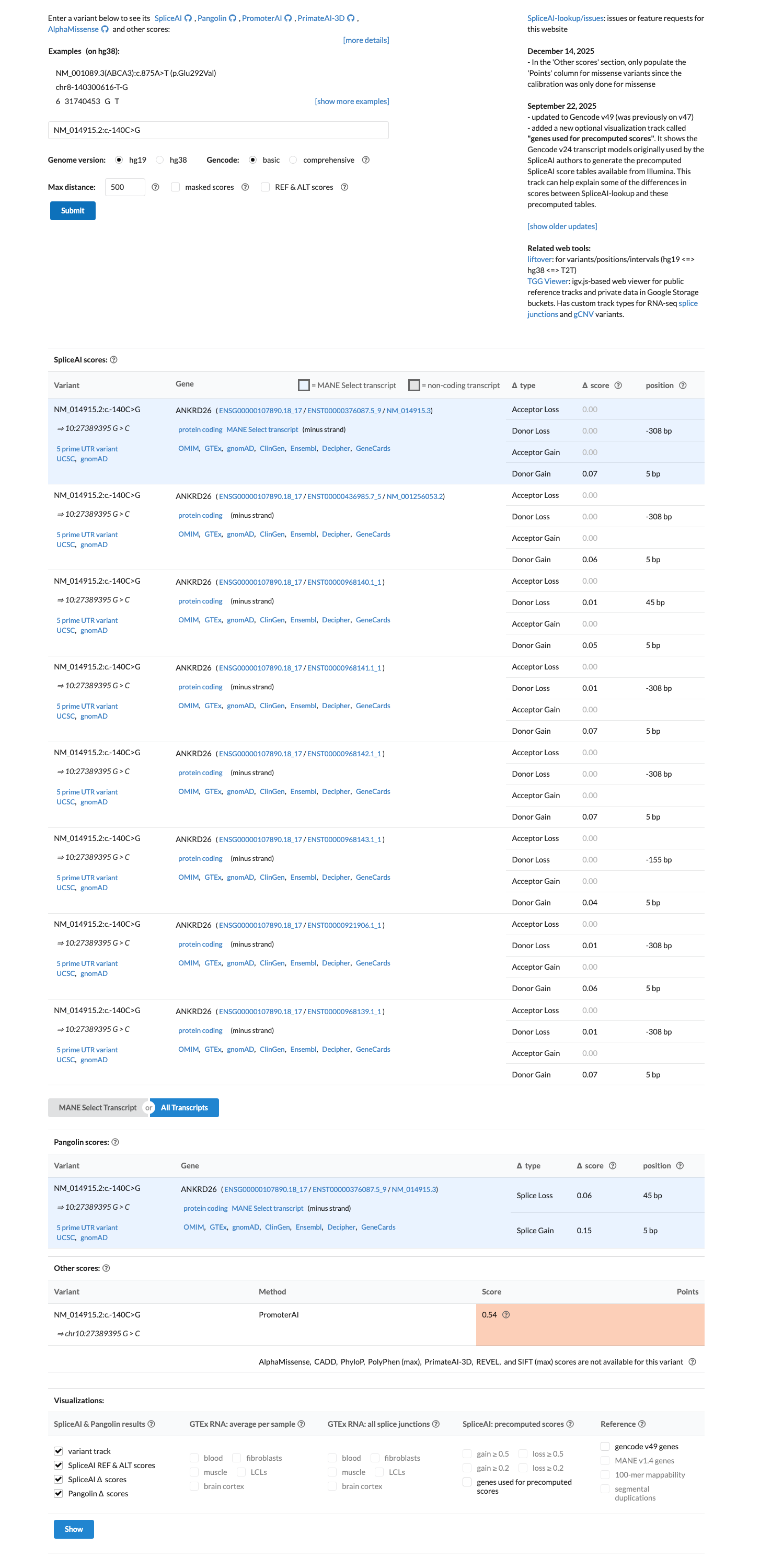
In silico splice prediction does not support a splice-altering effect, with a SpliceAI maximum delta score of 0.07.
spliceai ↗