Classification rationale
1
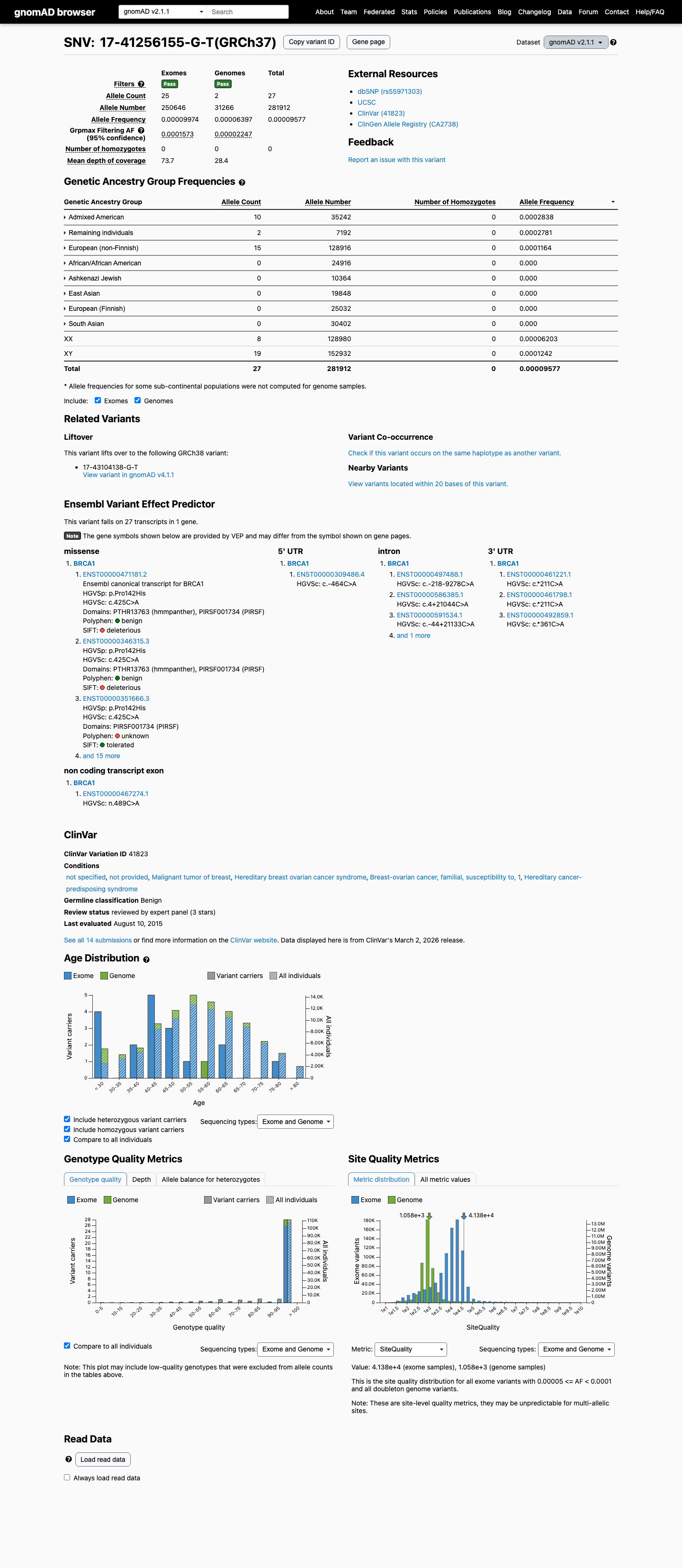
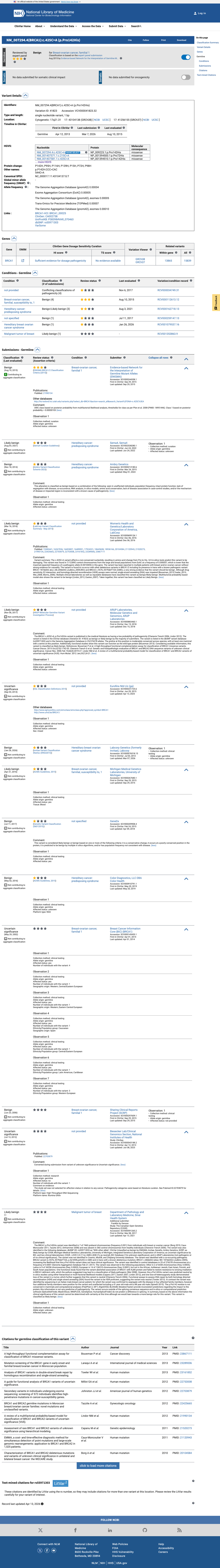
The BRCA1 c.425C>A (p.Pro142His) variant has been reported in ClinVar with an ENIGMA expert panel benign classification.
clinvar ↗2
This variant is present in gnomAD, and the highest observed filter allele frequencies exceed the ENIGMA BS1_Strong threshold of 0.0001 (gnomAD v2.1 grpmax FAF 0.00015726; gnomAD v4.1 joint grpmax FAF 0.00015364).
cspec ↗ gnomad_v2 ↗ gnomad_v4 ↗3
Calibrated BRCA1 functional evidence supports no damaging effect, with ENIGMA Table 9 assigning BS3_Strong and supplementary functional data describing no functional impact.
4
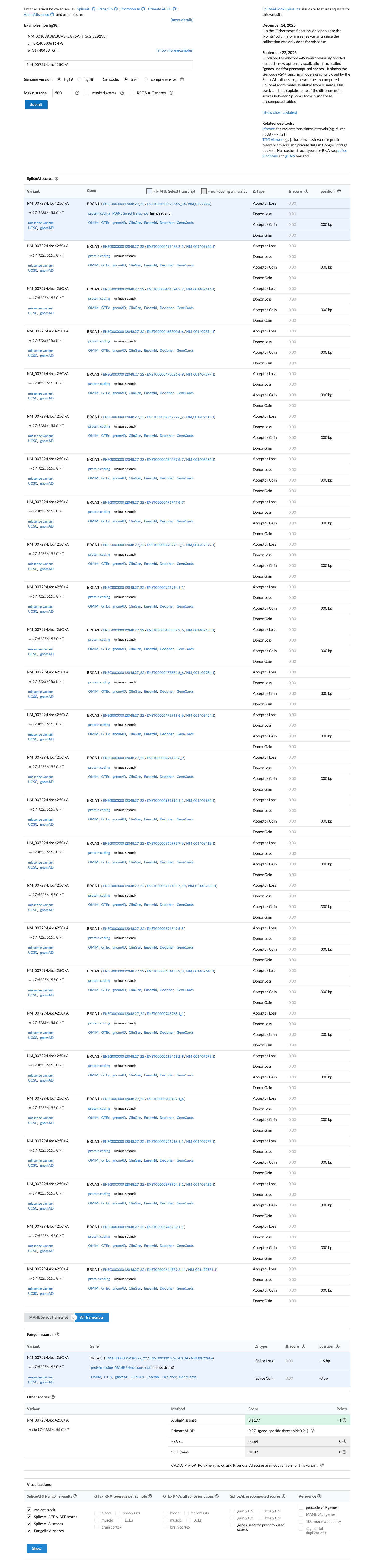
This missense change is outside the BRCA1 clinically important functional domains, and SpliceAI predicts no significant splice effect with a maximum delta score of 0.00, which is consistent with BP1_Strong and does not support PP3.
cspec ↗ spliceai ↗