Classification rationale
1
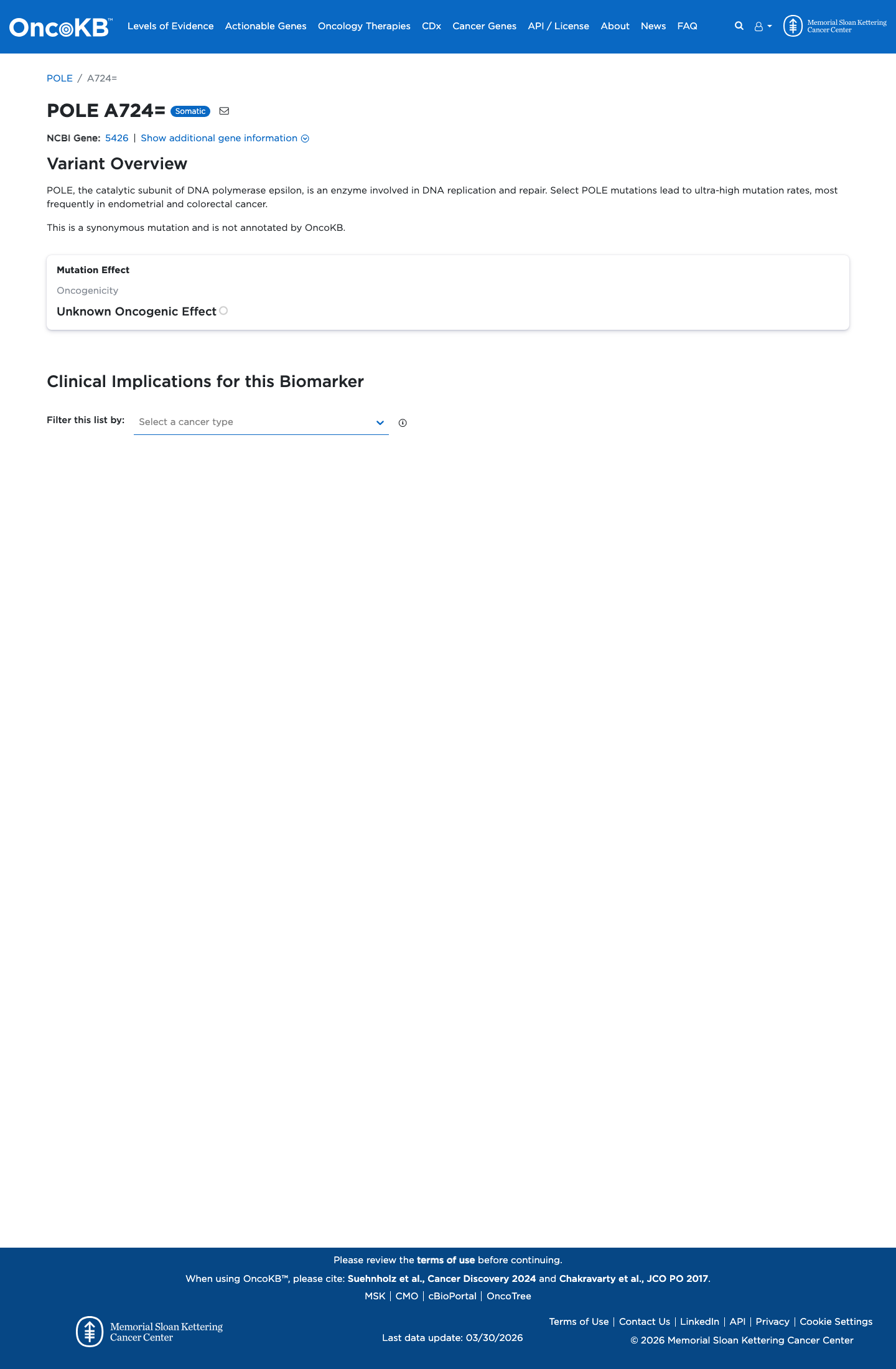
The POLE c.2172G>A (p.Ala724=) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar predominantly as likely benign, with five likely benign submissions and one submission of uncertain significance.
clinvar ↗2
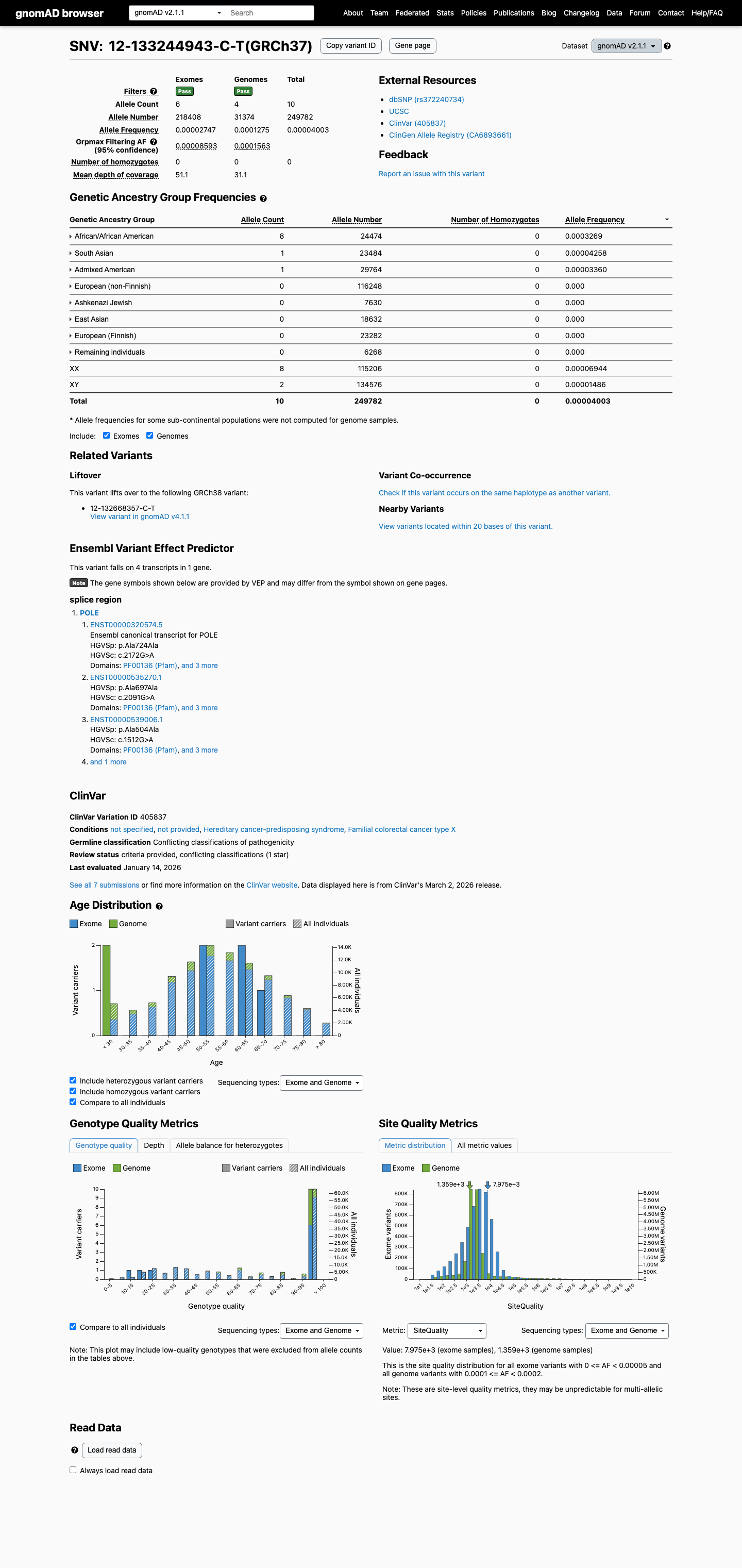
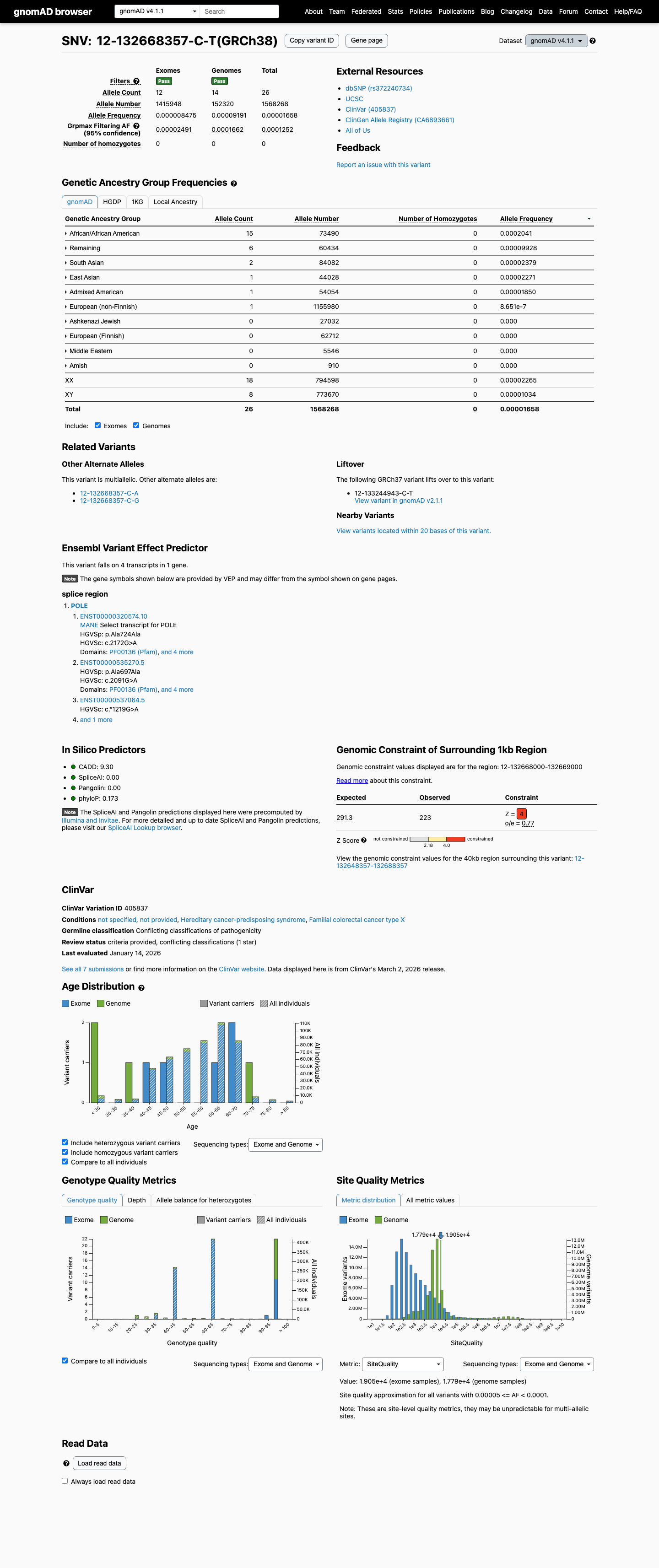
This variant is present at low frequency in gnomAD, with total allele frequencies of 0.00400% in v2.1 and 0.00166% in v4.1, both below the project's PM2 threshold of 0.1% and below the BS1 threshold of 0.3%.
gnomad_v2 ↗ gnomad_v4 ↗3
This is a synonymous change, and SpliceAI predicts no significant splice impact with a maximum delta score of 0.01; it is also not among the missense hotspot or computationally selected variants used for the custom POLE PM1, PS4, PP3, and BP4 rules.
spliceai ↗