The TP53 c.215C>G (p.Pro72Arg, p.P72R) variant has not been observed in COSMIC as a recurrent somatic cancer variant and is reported in ClinVar as benign, including a benign ClinGen TP53 Variant Curation Expert Panel classification.
clinvar ↗This variant is very common in population databases, with gnomAD v4.1 total allele frequency 0.707882 and grpmax filtering allele frequency 0.746273, far above the TP53 BA1 benign threshold of 0.001.
gnomad_v4 ↗ gnomad_v2 ↗ cspec ↗Available studies of the common TP53 codon 72 polymorphism describe biologic differences between alleles, but no TP53 VCEP-eligible functional code assignment for p.Pro72Arg was identified in the TP53 functional worksheet.
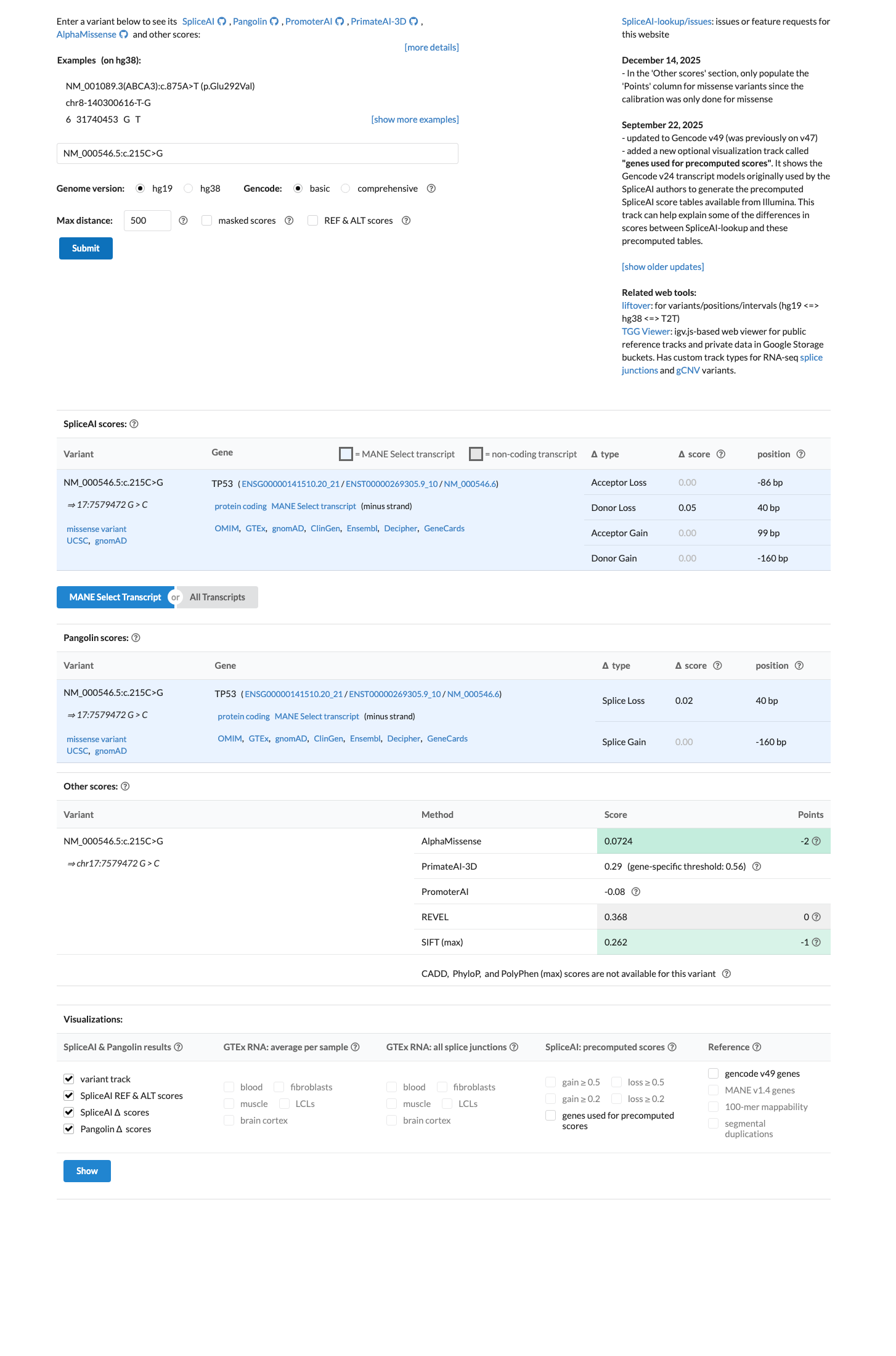
PMID:10802655 ↗ PMID:12567188 ↗ PMID:16964264 ↗TP53 VCEP in silico data support a benign interpretation because c.215C>G is assigned BP4_moderate in the TP53 PP3/BP4 worksheet with BayesDel -0.108475, and SpliceAI predicts no significant splice effect with a maximum delta score of 0.05.
spliceai ↗ cspec ↗