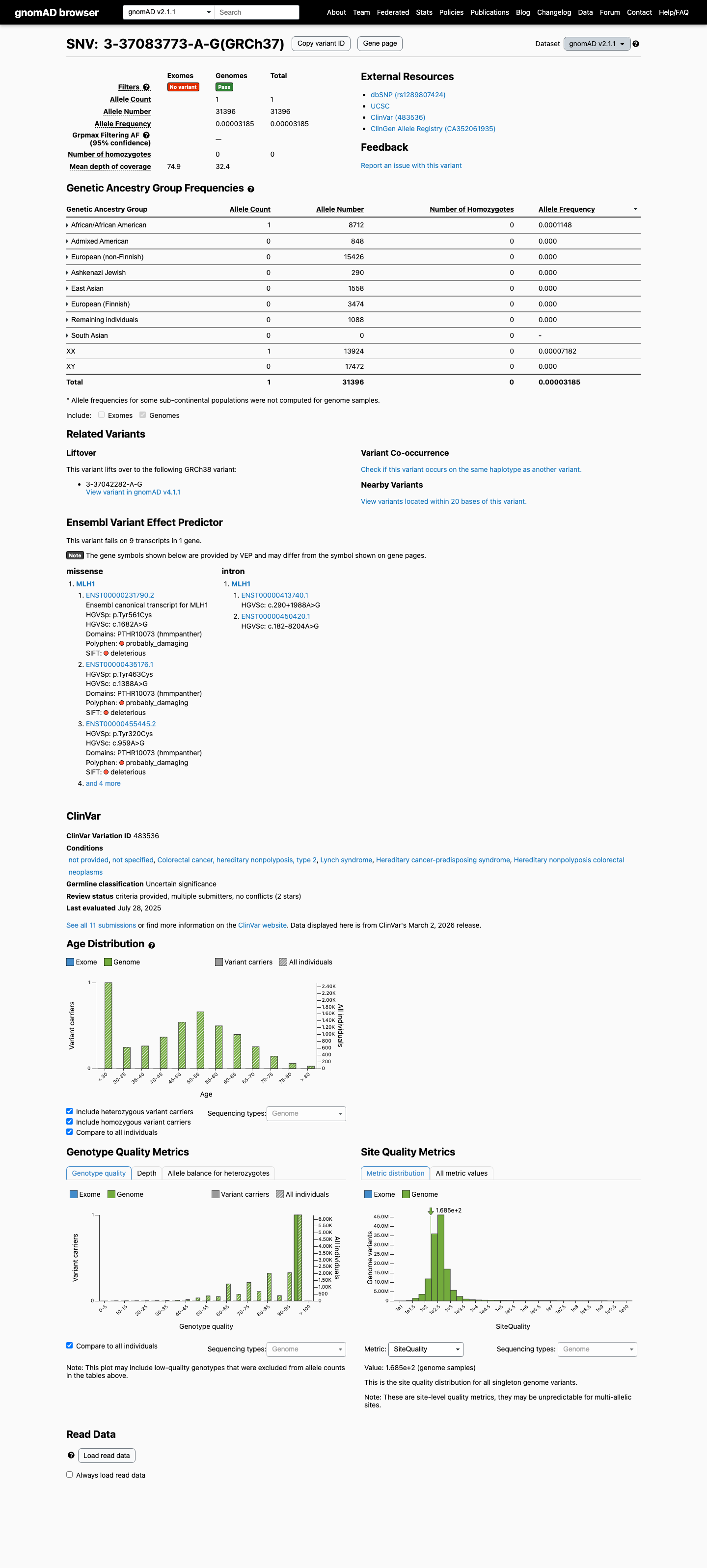
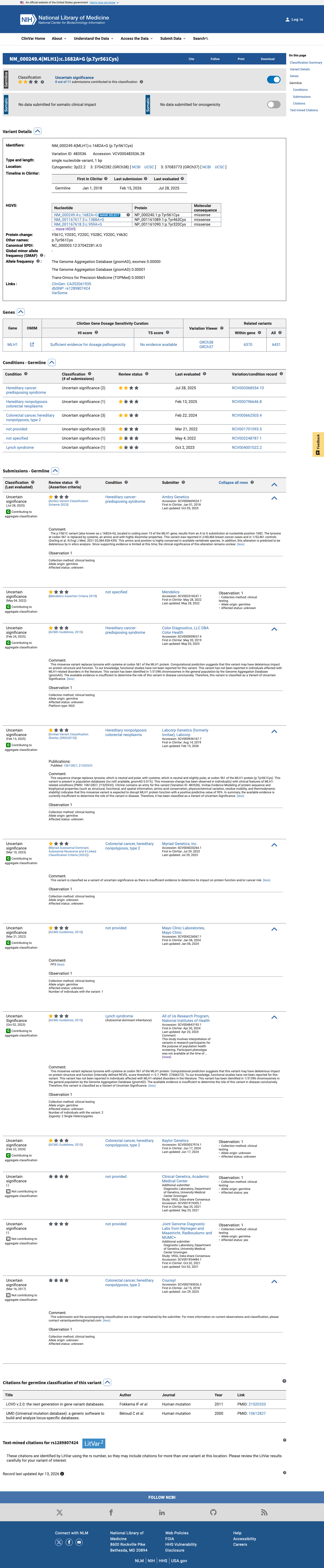
The MLH1 c.1682A>G (p.Tyr561Cys; p.Y561C) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar with uncertain significance submissions.
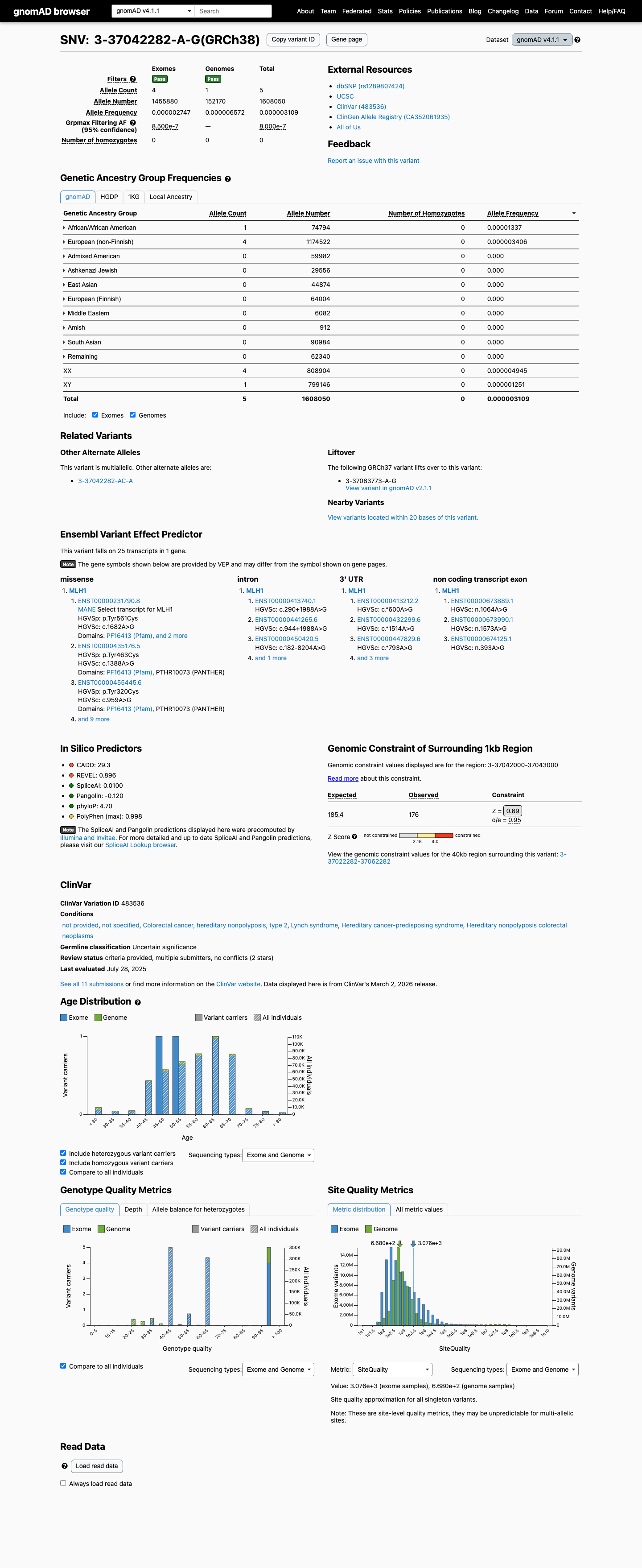
clinvar ↗This variant is present at very low frequency in population databases, with gnomAD v4.1 total allele frequency 3.10936e-06 and grpmax filtering allele frequency 8e-07, which is below the MLH1 PM2_Supporting threshold of 0.00002.
cspec ↗ gnomad_v4 ↗ gnomad_v2 ↗No variant-specific calibrated functional assay result for this variant was identified in the reviewed MMR functional assay materials, so functional evidence was not used to support or refute pathogenicity.
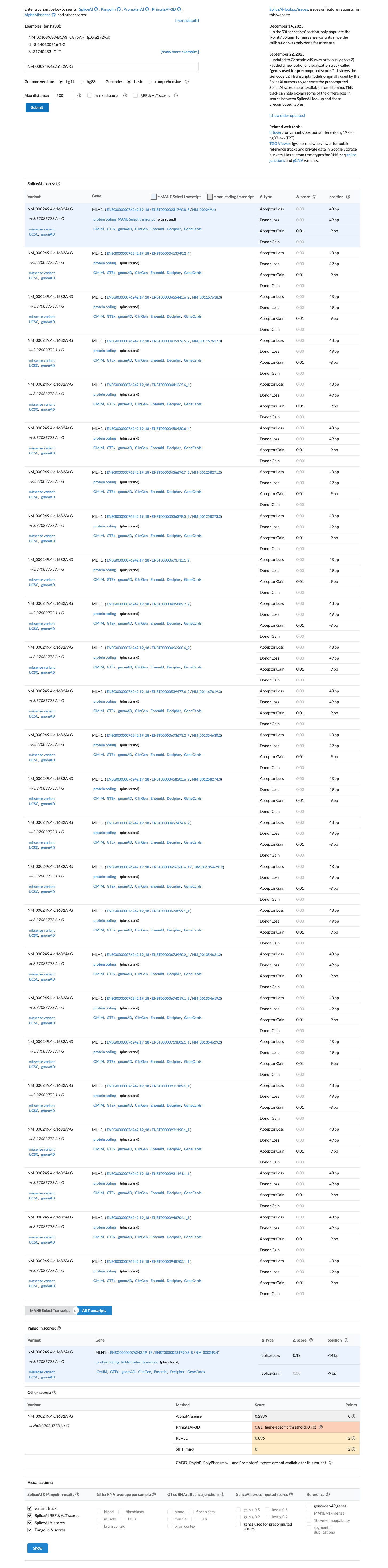
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.01, and available in silico evidence was insufficient to assign the MLH1 missense PP3 or BP4 rules without an HCI-prior value.
cspec ↗ spliceai ↗