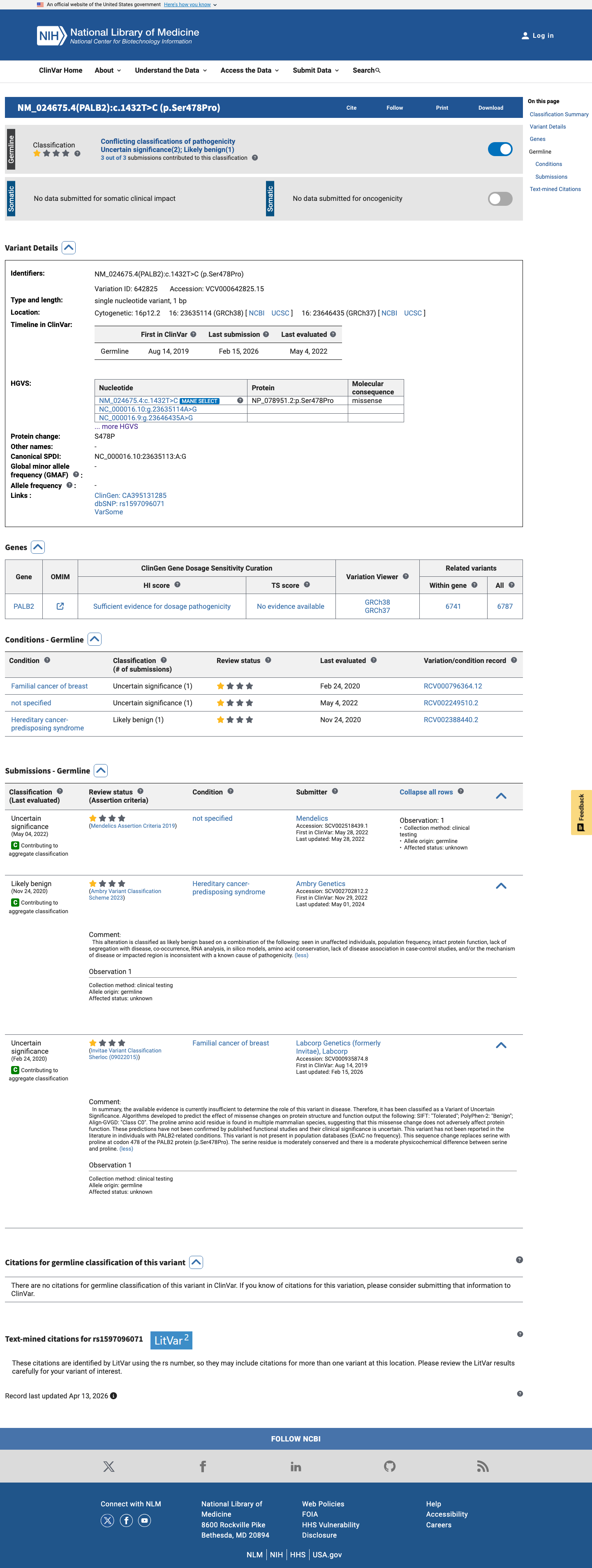
The PALB2 c.1432T>C (p.Ser478Pro) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar with conflicting germline submissions, including uncertain significance and likely benign interpretations.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1; the observed frequency is therefore below the PALB2 PM2_Supporting threshold of ≤0.000333% and below the benign population thresholds for BS1 and BA1.
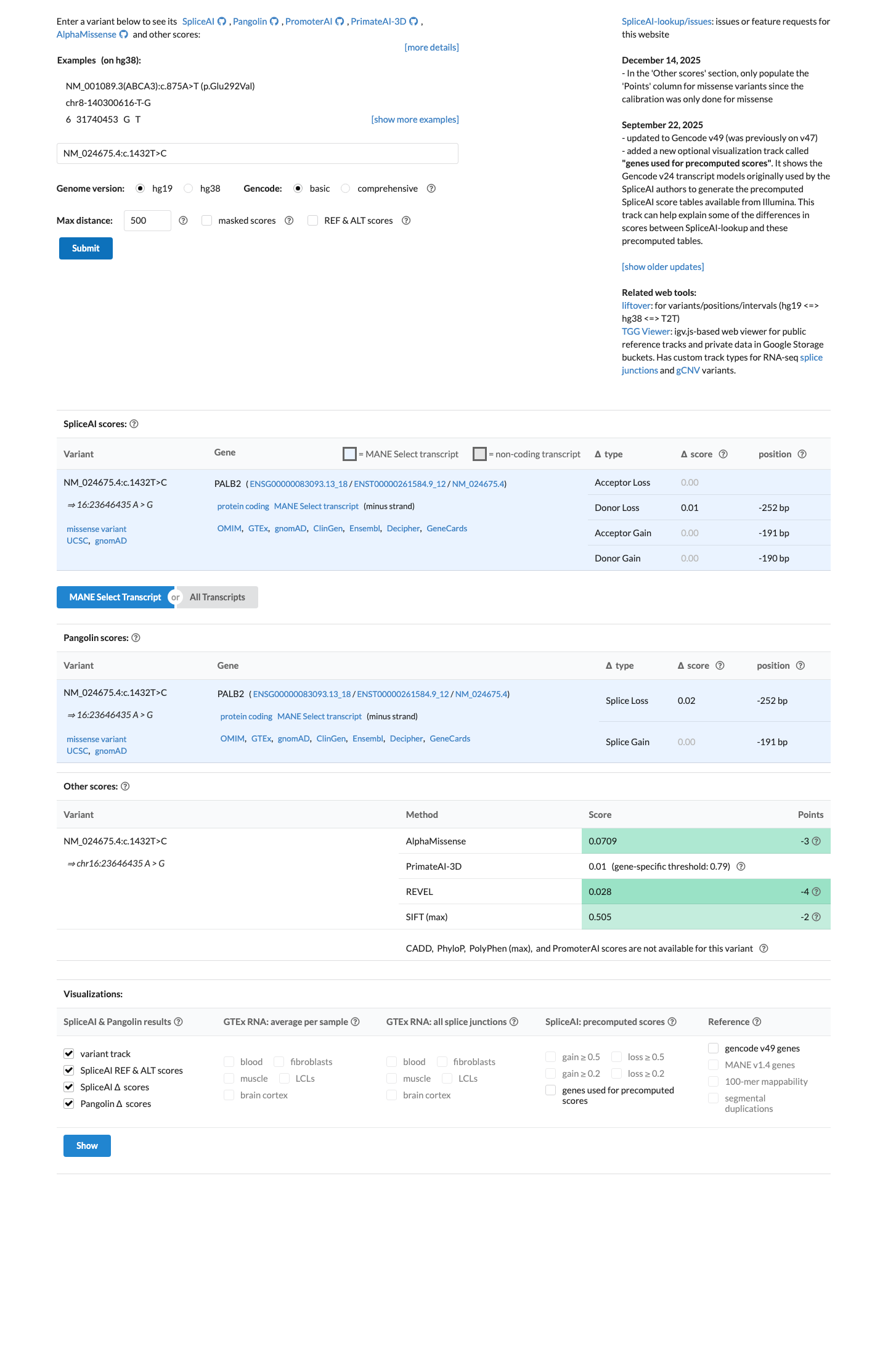
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.01, which is below the PALB2 splice thresholds used for PP3 (≥0.2) and BP4 (≤0.1), although PP3 and BP4 are not applied to missense variants under the PALB2 specification.
spliceai ↗ cspec ↗Under the PALB2 expert specification, BP1_Supporting is met because this is a missense variant in a gene where disease is primarily associated with truncating variants, while PVS1 is not met because this variant is not a qualifying loss-of-function change.
cspec ↗