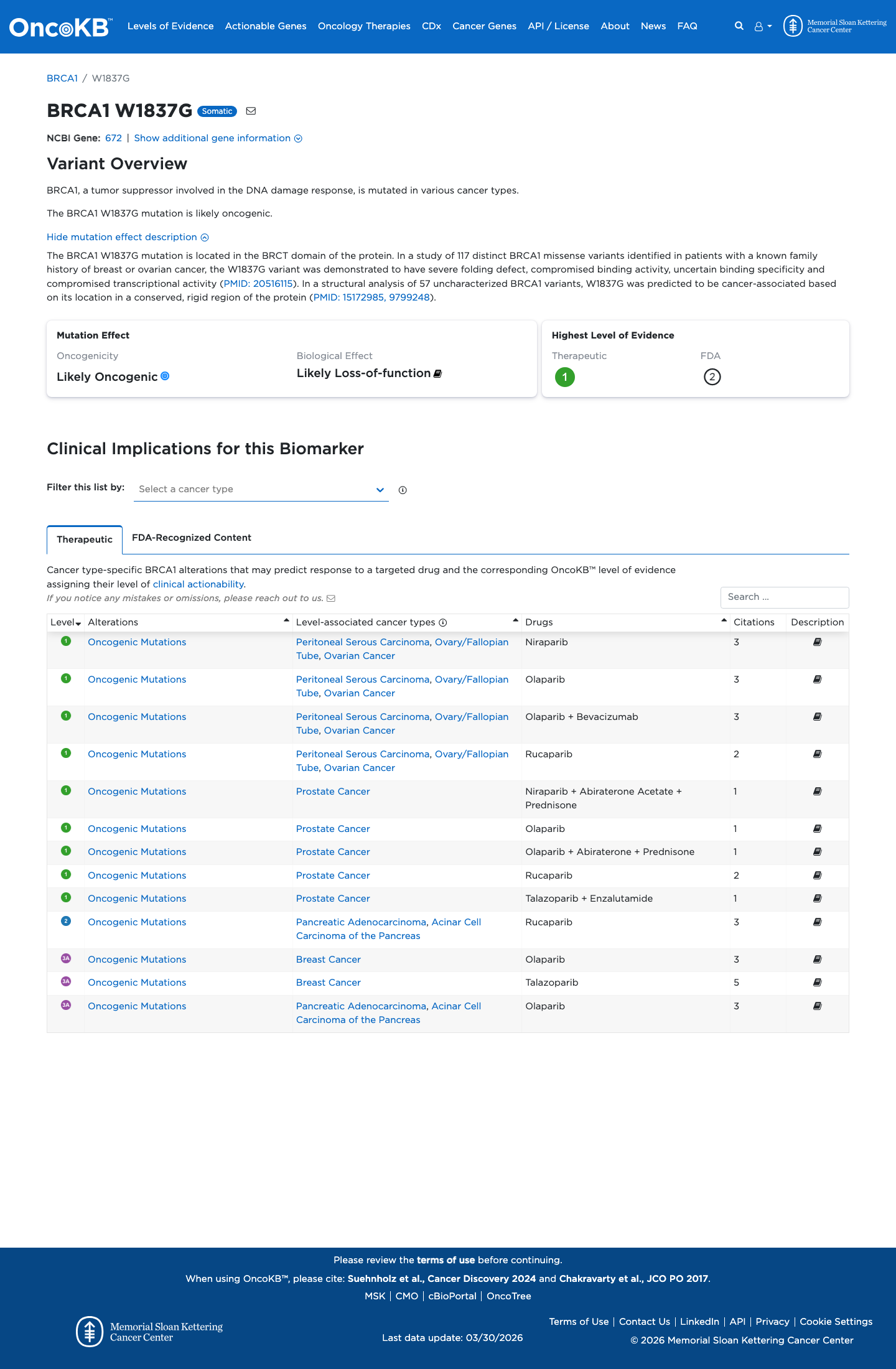
The BRCA1 c.5509T>G (p.Trp1837Gly; W1837G) variant has been reported in ClinVar with an expert-panel Pathogenic classification by the ClinGen ENIGMA BRCA1 and BRCA2 Variant Curation Expert Panel, and OncoKB classifies the variant as Likely Oncogenic with likely loss-of-function biological effect.
clinvar ↗ oncokb ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, giving an observed population frequency of 0 in the available population datasets and supporting PM2_Supporting under the ENIGMA framework.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗ENIGMA curated functional data for c.5509T>G are discordant across calibrated studies, so neither PS3 nor BS3 is met for this variant.
The variant lies in the BRCA1 BRCT repeats and has an ENIGMA supplementary BayesDel no-AF score of approximately 0.533, above the PP3 threshold of ≥0.28, while SpliceAI predicts no significant splice impact with a max delta score of 0.00.
spliceai ↗ cspec ↗The BRCA1 clinical-history likelihood-ratio table reports c.5509T>G in 2 probands with LR 8.0617, exceeding the PP4_Moderate threshold of LR ≥4.3 and falling below the PP4_Strong threshold of LR ≥18.7.
PMID:31853058 ↗ cspec ↗