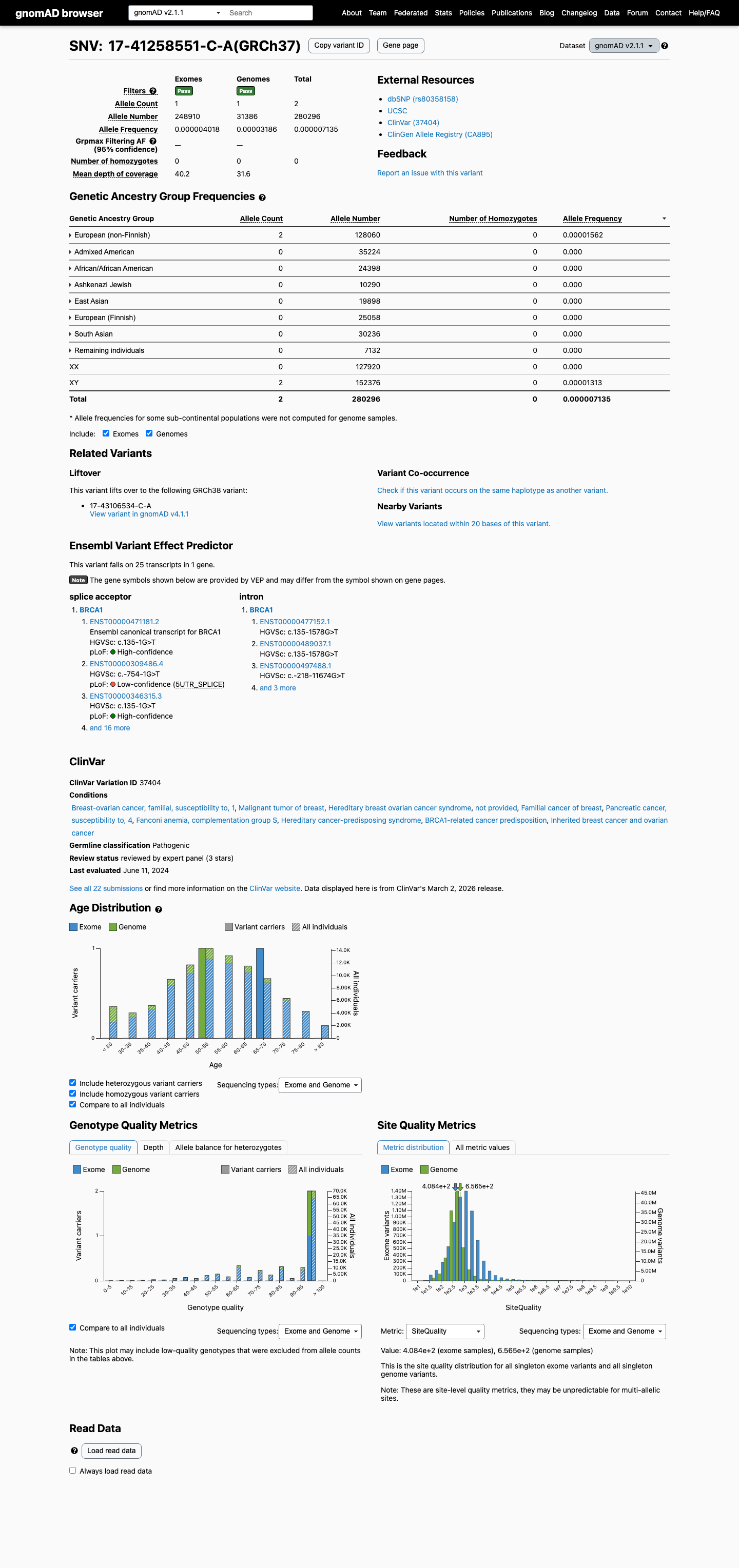
The BRCA1 c.135-1G>T (NP_009225.1:p.?) variant has been observed in somatic cancers in COSMIC and has been reported in ClinVar with a pathogenic expert-panel classification.
clinvar ↗This variant is present at very low frequency in population databases, with gnomAD v2.1 AF 7.14e-06 (2/280296 alleles) and gnomAD v4.1 AF 4.42e-06 (7/1583254 alleles), which is below benign frequency thresholds but does not meet the ENIGMA PM2 requirement for absence from controls.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗RNA and multifactorial evidence support a damaging splice effect for this variant, including exon 5 deletion in splicing studies, ENIGMA assignment of PVS1 (RNA), a reference-set posterior probability of 0.99999838416, and a BRCA1 clinical-history likelihood ratio of 59.69 across 7 probands, supporting PP4_Strong.
PMID:20020529 ↗ PMID:31853058 ↗In silico splicing analysis predicts a strong splice effect, with a SpliceAI maximum delta score of 0.86, which is consistent with disruption of normal splicing but is not counted separately as PP3 because the same splice evidence is already captured under the PVS1 framework.
spliceai ↗ cspec ↗