Classification rationale
1
2
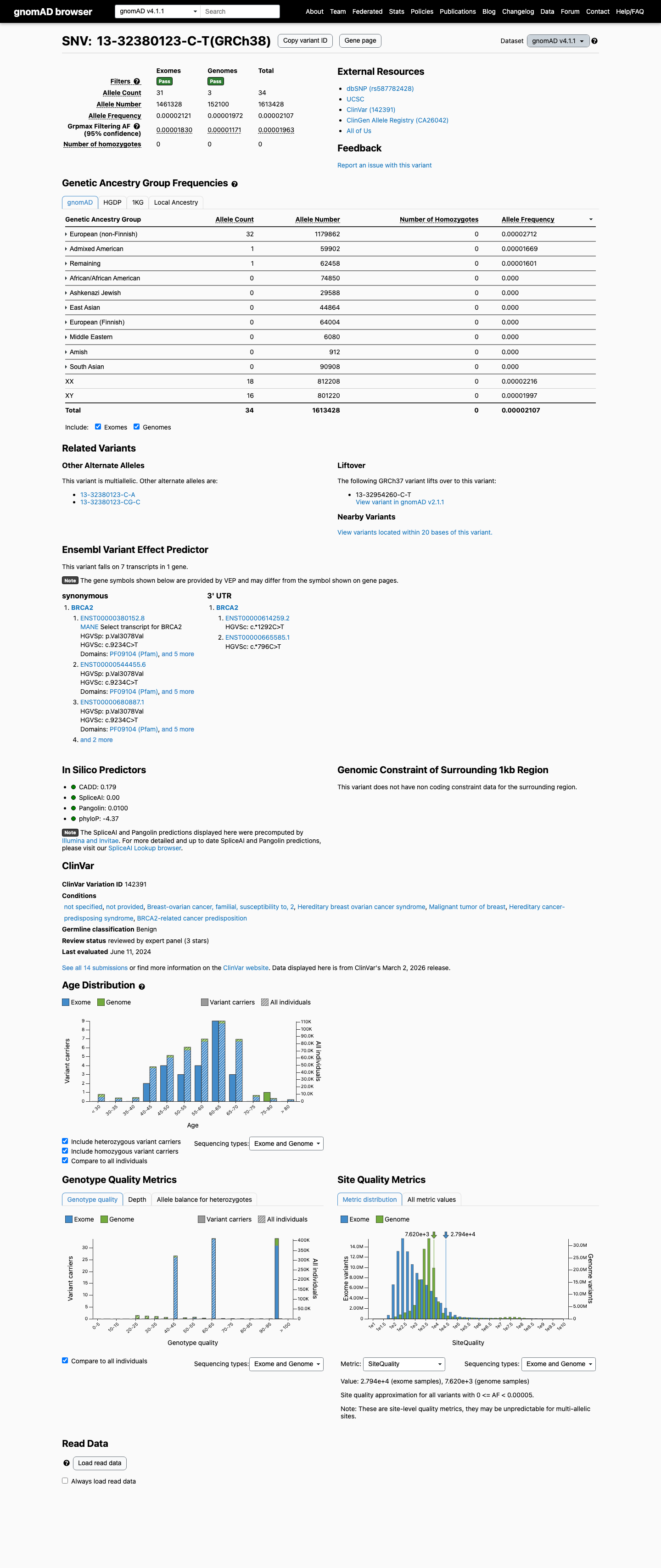
This variant is rare but not absent in population databases, with gnomAD v2.1 AF 1.77165e-05 (5/282222 alleles) and gnomAD v4.1 AF 2.10731e-05 (34/1613428 alleles); the highest observed filter allele frequency is 1.963e-05, which is below the BS1 supporting threshold of greater than 0.00002 and inconsistent with the PM2 requirement for absence.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
BRCA2 clinical-history likelihood data show an LR of 0.284 in 12 probands, which is at or below the BP5 supporting threshold of 0.48 and supports a benign clinical-history code.
PMID:31853058 ↗ cspec ↗4
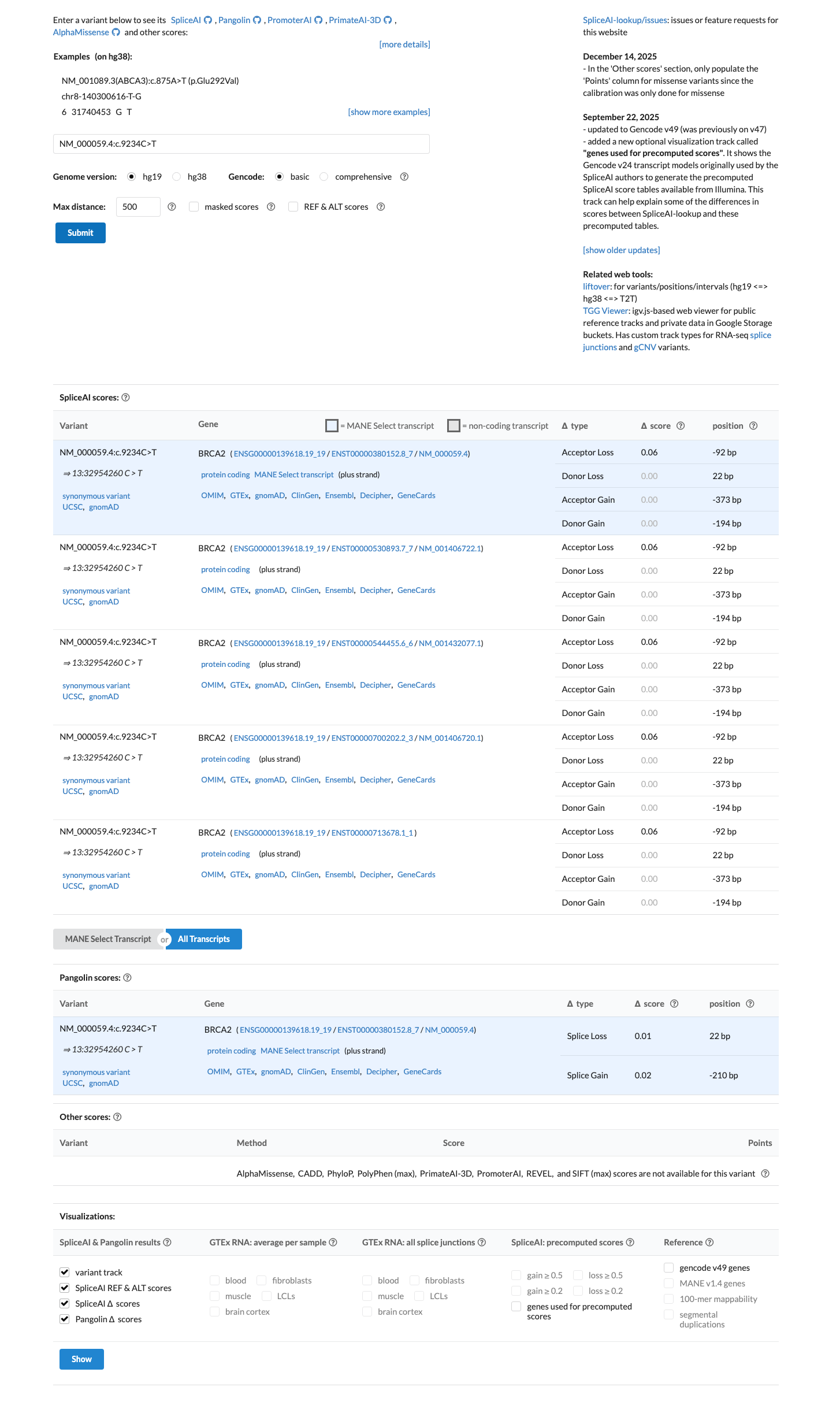
This synonymous variant lies within the BRCA2 DNA-binding domain (amino acids 2481-3186), and SpliceAI predicts no significant splice effect with a maximum delta score of 0.06, supporting BP4 and BP7 and arguing against PP3.
spliceai ↗ cspec ↗