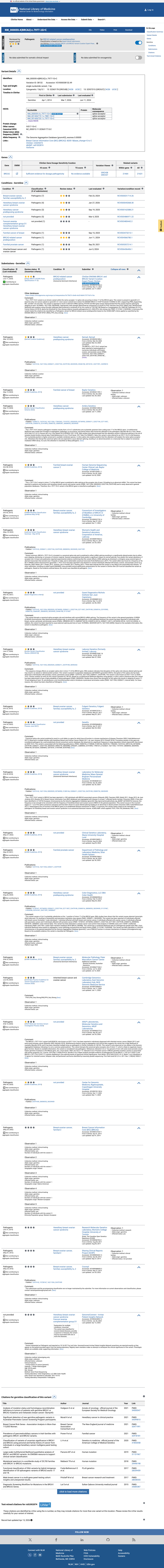
The BRCA2 c.7977-1G>C (p.?) variant has been reported in ClinVar as pathogenic, including review by the ClinGen ENIGMA BRCA1/2 expert panel.
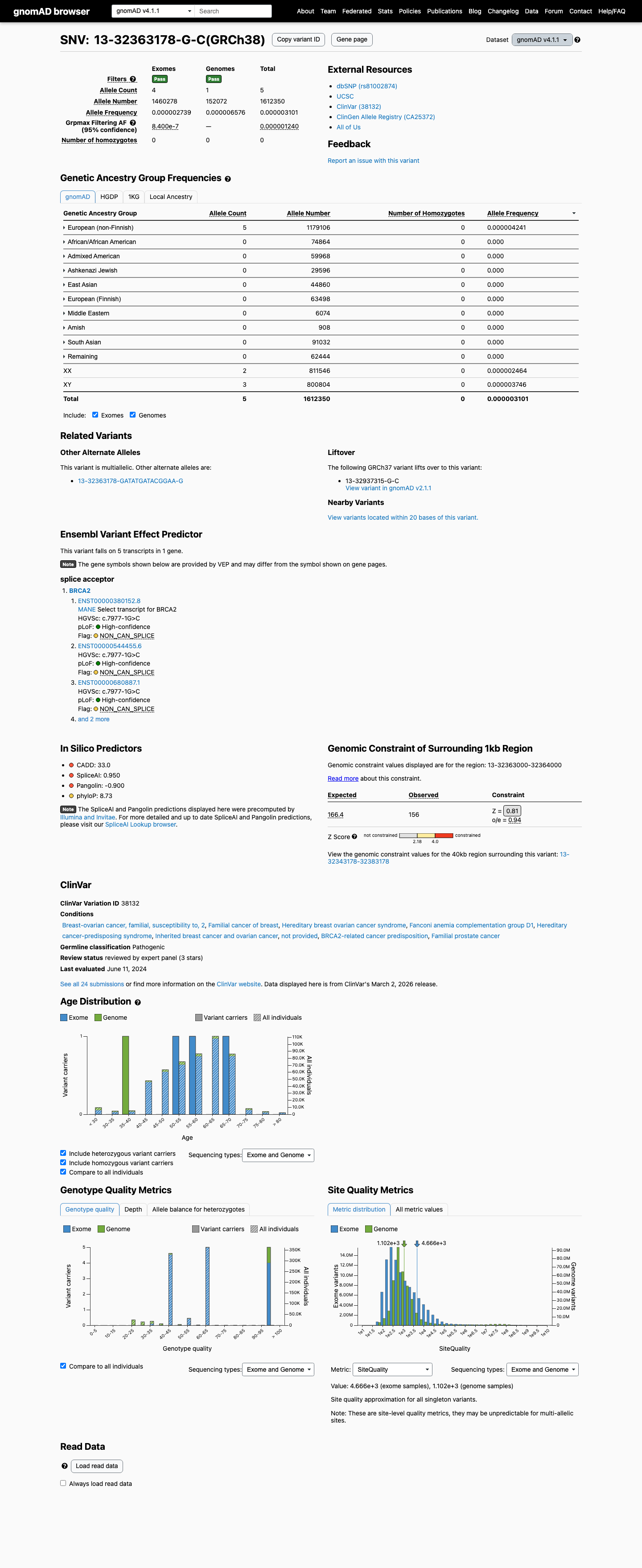
clinvar ↗This variant is present at very low frequency in population databases, with 2/281032 alleles in gnomAD v2.1 (AF 7.12e-06) and 5/1612350 alleles in gnomAD v4.1 (AF 3.10e-06; grpmax FAF 1.24e-06), which is too low for BS1 or BA1 but means PM2 is not met because the variant is not absent from controls.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗A BRCA2 clinical-history likelihood-ratio analysis reported LR 7.22 across 7 probands, which meets ENIGMA PP4_Moderate.
PMID:31853058 ↗RNA evidence cited by ENIGMA indicates that variants at this splice acceptor site cause leaky abnormal splicing, and ENIGMA specifically assigns c.7977-1G>C as PVS1_Strong (RNA).
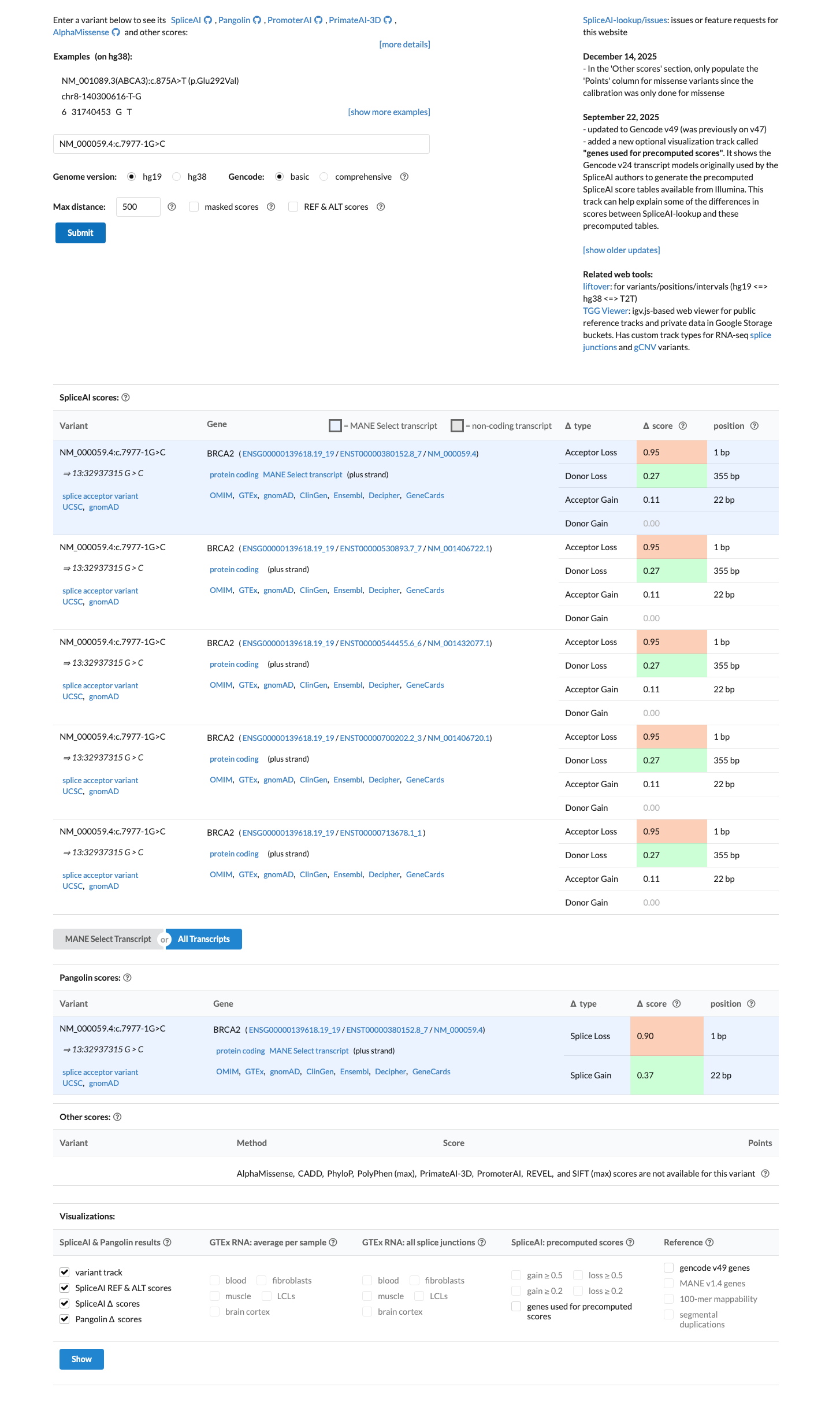
PMID:16211554 ↗SpliceAI predicts a strong splice effect for this variant with a maximum delta score of 0.95, which supports splice disruption but is not separately counted as PP3 because the variant is at a canonical splice position already captured by PVS1.
spliceai ↗ cspec ↗