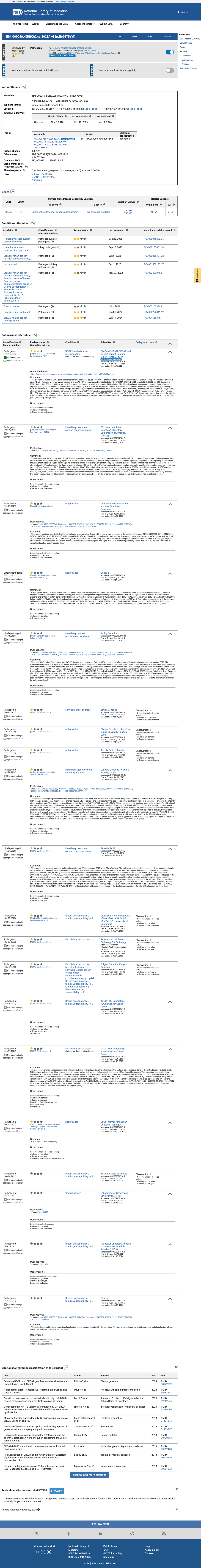
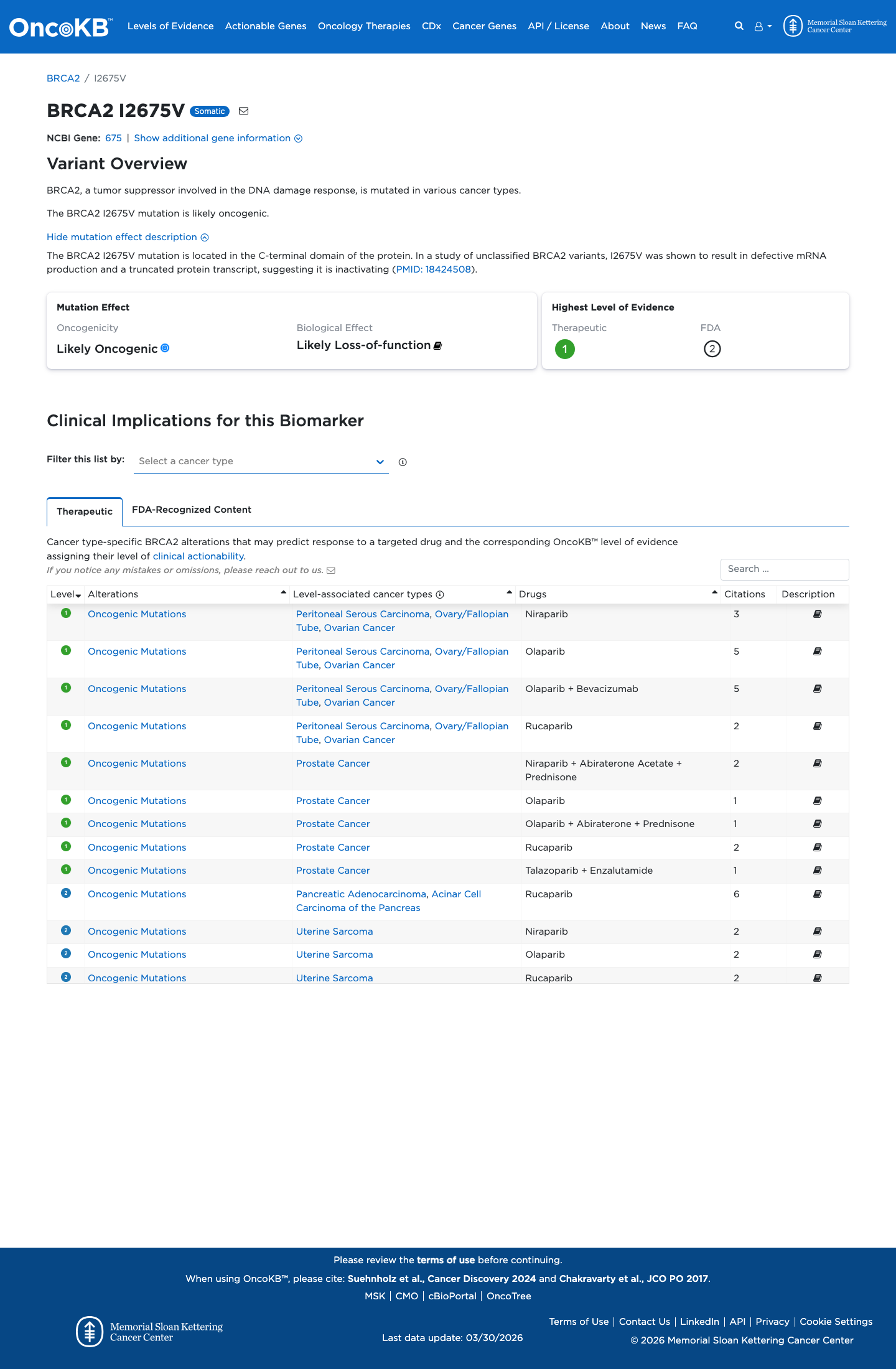
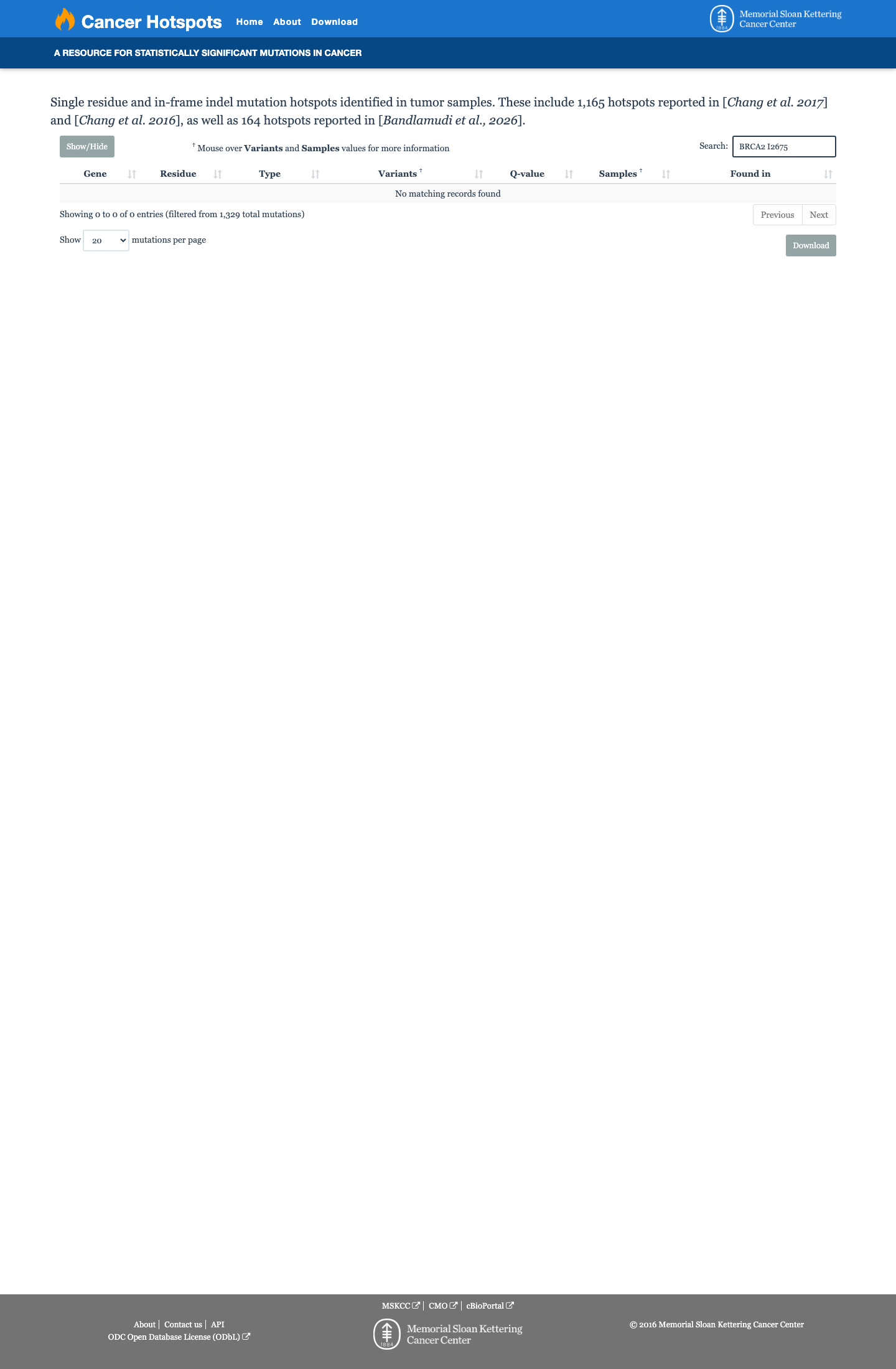
The BRCA2 c.8023A>G (p.Ile2675Val) variant has not been observed in COSMIC and has been reported in ClinVar as pathogenic, including an expert-panel pathogenic classification from the ClinGen ENIGMA BRCA1/2 Variant Curation Expert Panel.
clinvar ↗This variant is present at very low frequency in gnomAD, with 1/251076 alleles in v2.1 and 1/1614050 alleles in v4.1; the highest observed population frequencies are 5.44e-05 in East Asian individuals in v2.1 and 2.23e-05 in East Asian individuals in v4.1, so the variant is rare but not absent from controls.
gnomad_v2 ↗ gnomad_v4 ↗BRCA2 multifactorial data show strong pathogenic evidence for this variant, including a segregation likelihood ratio of 605.14 and a posterior probability of 0.999651, and multiple splicing studies reported complete 309-nt skipping of exon 18 with no full-length transcript detected from the variant allele.
PMID:18424508 ↗ PMID:22505045 ↗Computational splicing analysis supports a deleterious effect, with a SpliceAI maximum delta score of 0.99, which is above the ENIGMA PP3 threshold of 0.2 and well above the benign BP4 threshold of 0.1.
spliceai ↗ cspec ↗