Classification rationale
1
2
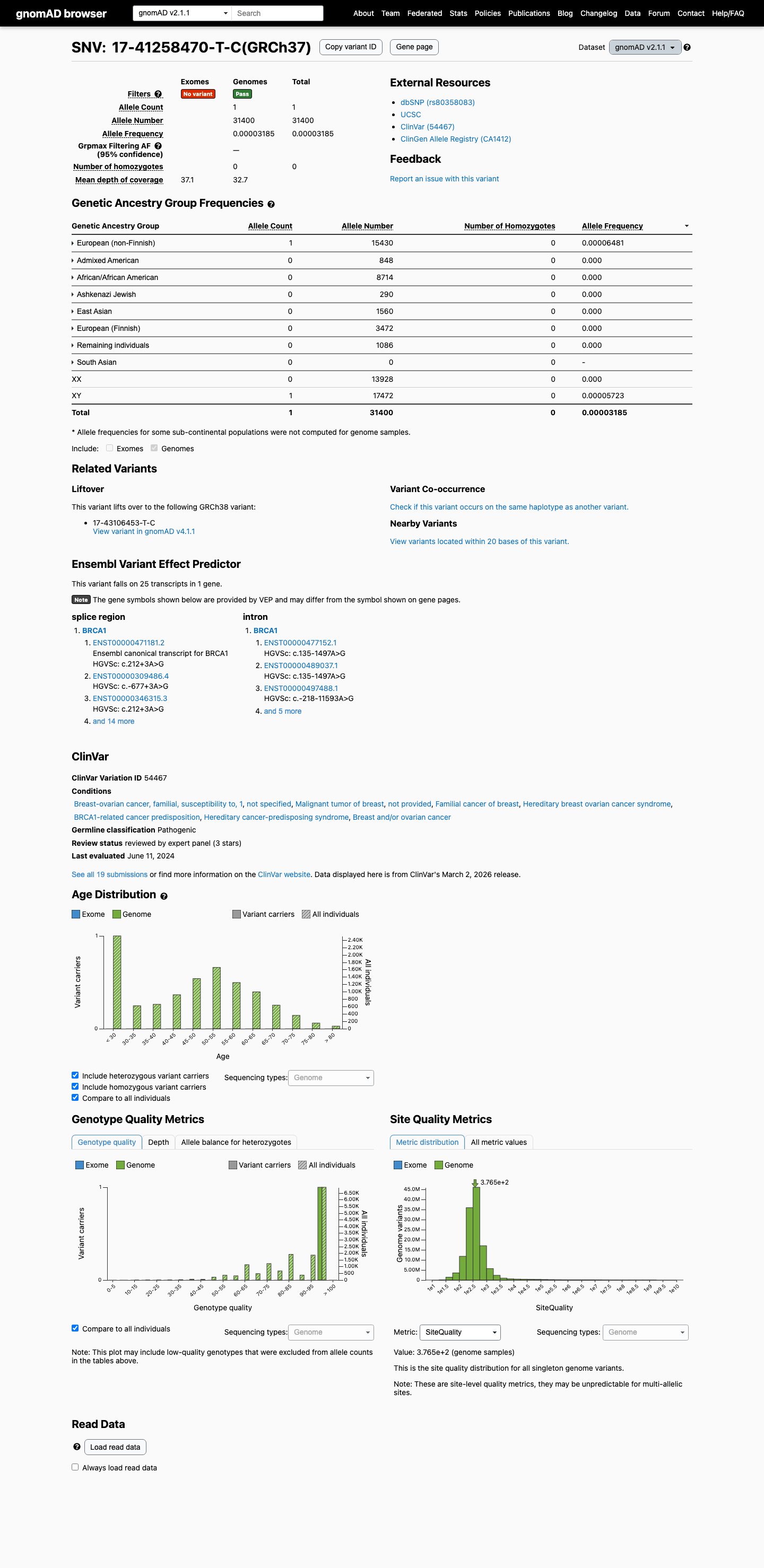
This variant is absent from gnomAD v2.1 and is present in gnomAD v4.1 at a very low frequency of 2/1574274 alleles (AF 1.27043e-06; 0.00013%), which is below the BA1 and BS1 population thresholds.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
ENIGMA-calibrated functional evidence supports a damaging effect: Table 9 assigns PS3_Strong for BRCA1 c.212+3A>G, and supplementary ENIGMA datasets classify the variant as having complete functional impact with multiple RNA assays showing aberrant splicing consistent with loss of function.
PMID:12037674 ↗4
SpliceAI predicts a deleterious splicing effect with a maximum delta score of 0.63, which exceeds the ENIGMA PP3 threshold of 0.2 for intronic variants outside the canonical +/-1,2 splice positions.
spliceai ↗ cspec ↗