Classification rationale
1
2
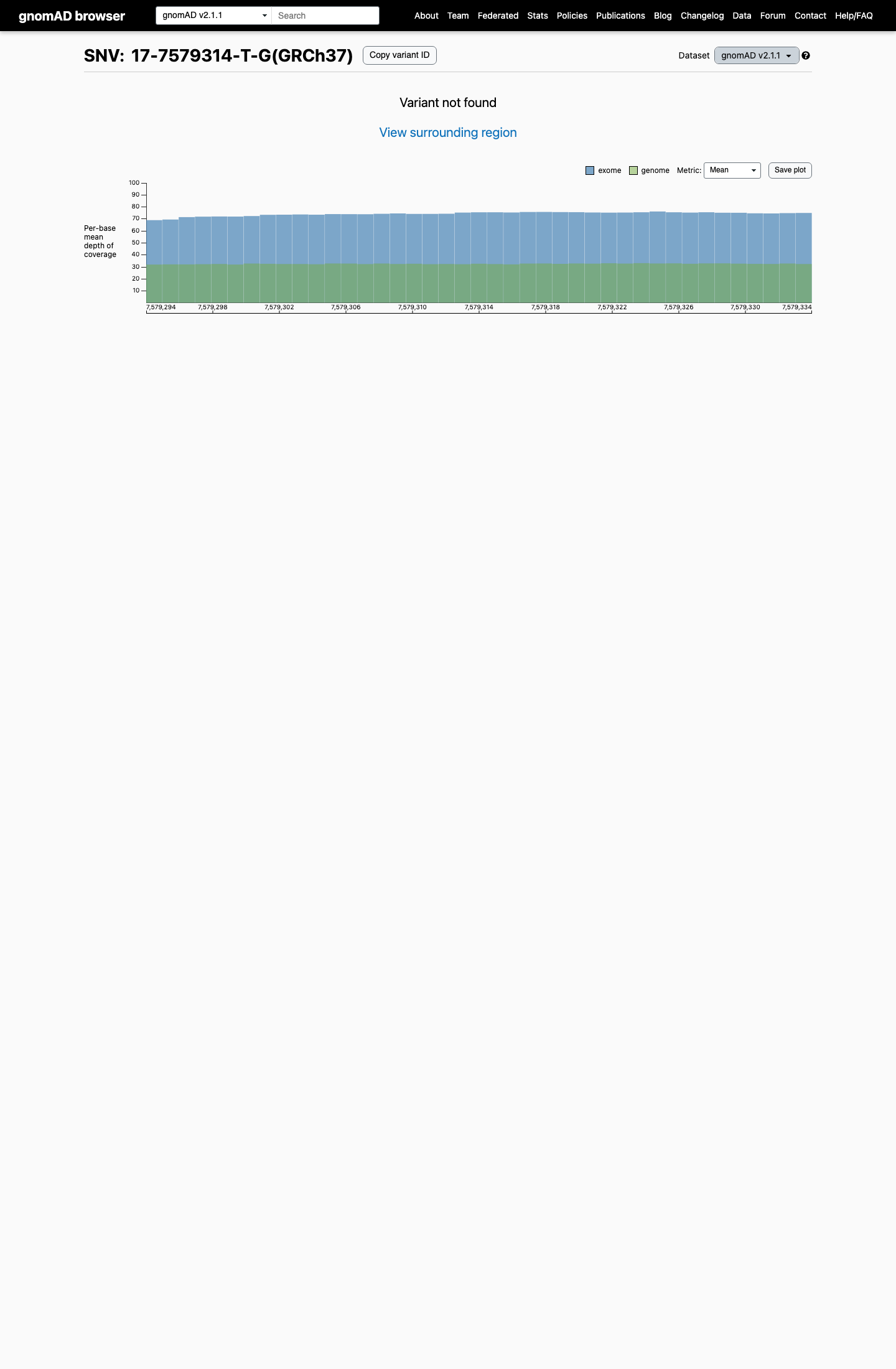
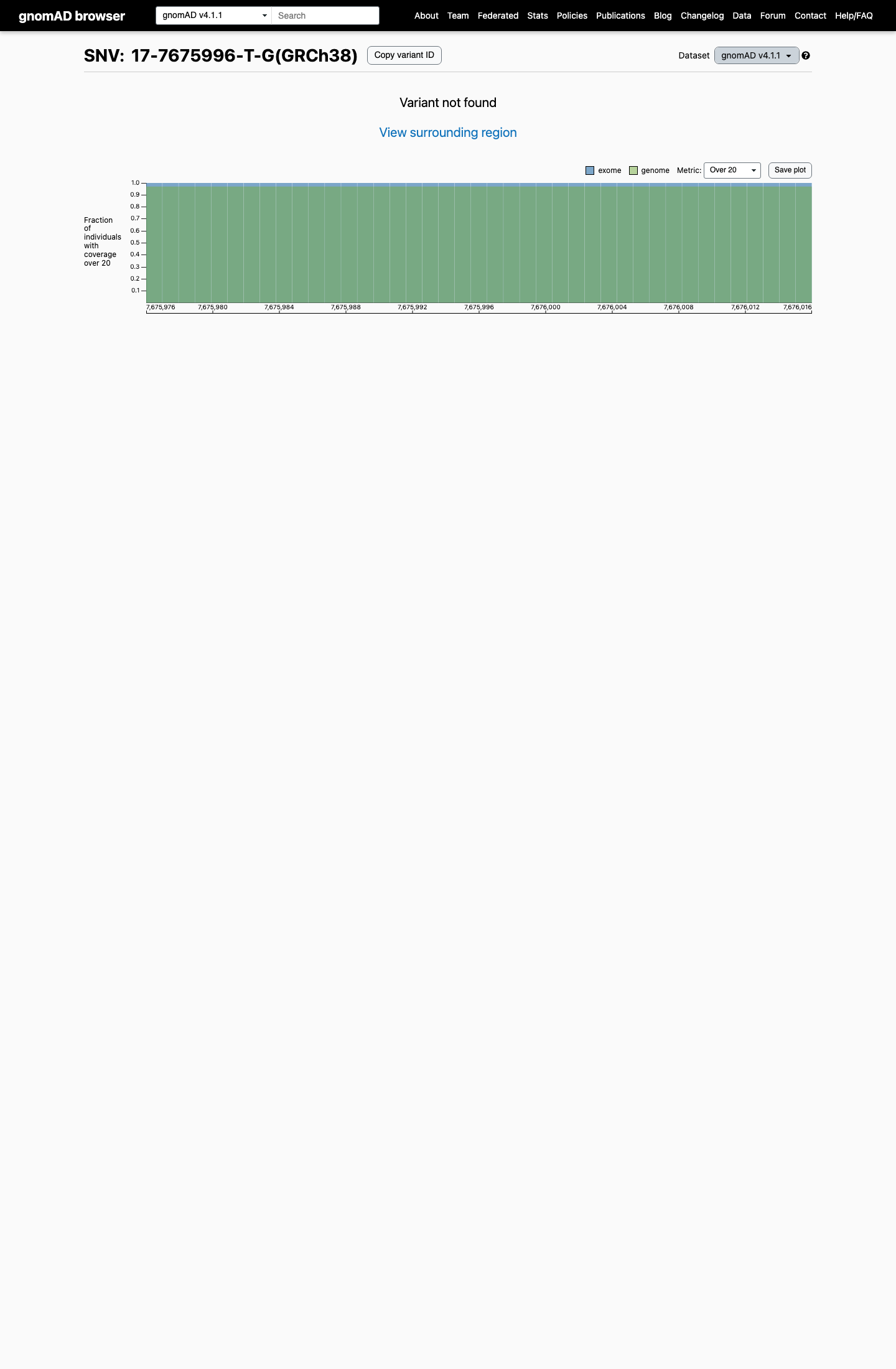
This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity and meeting the TP53 VCEP PM2_Supporting population threshold of less than 0.00003.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
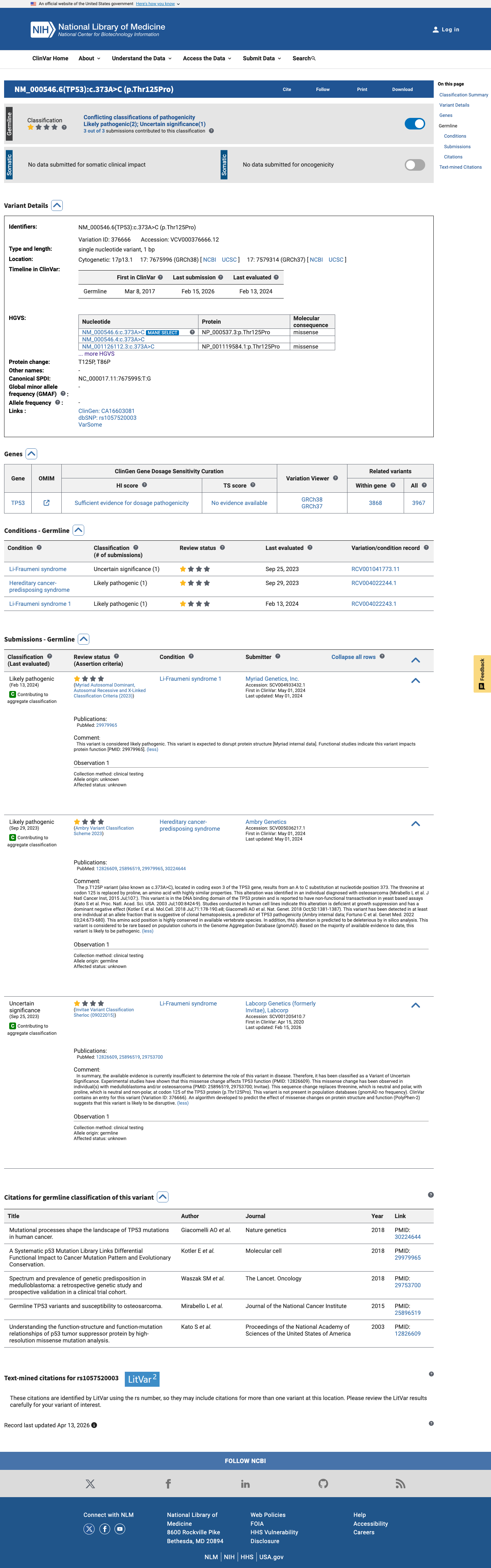
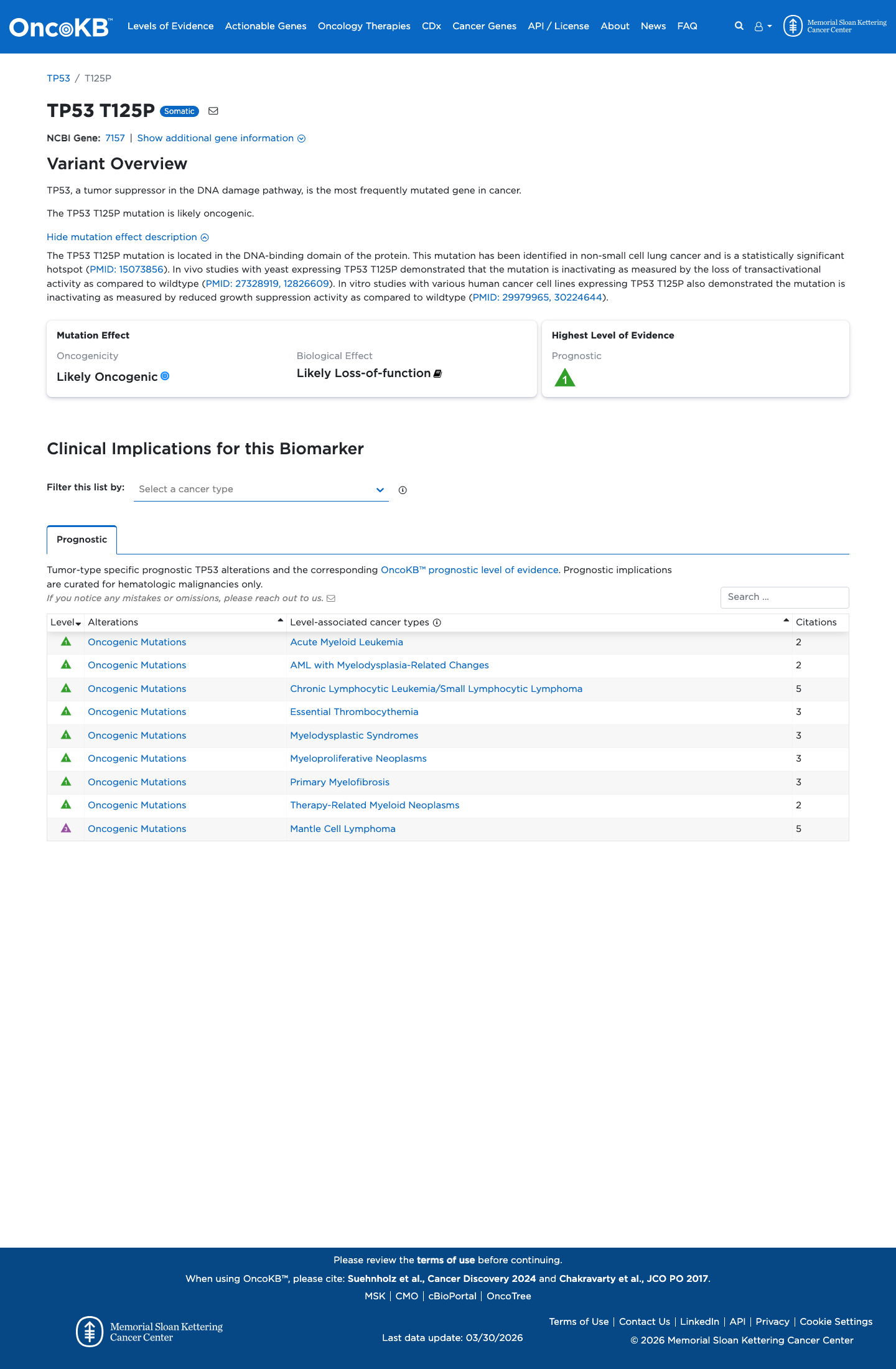
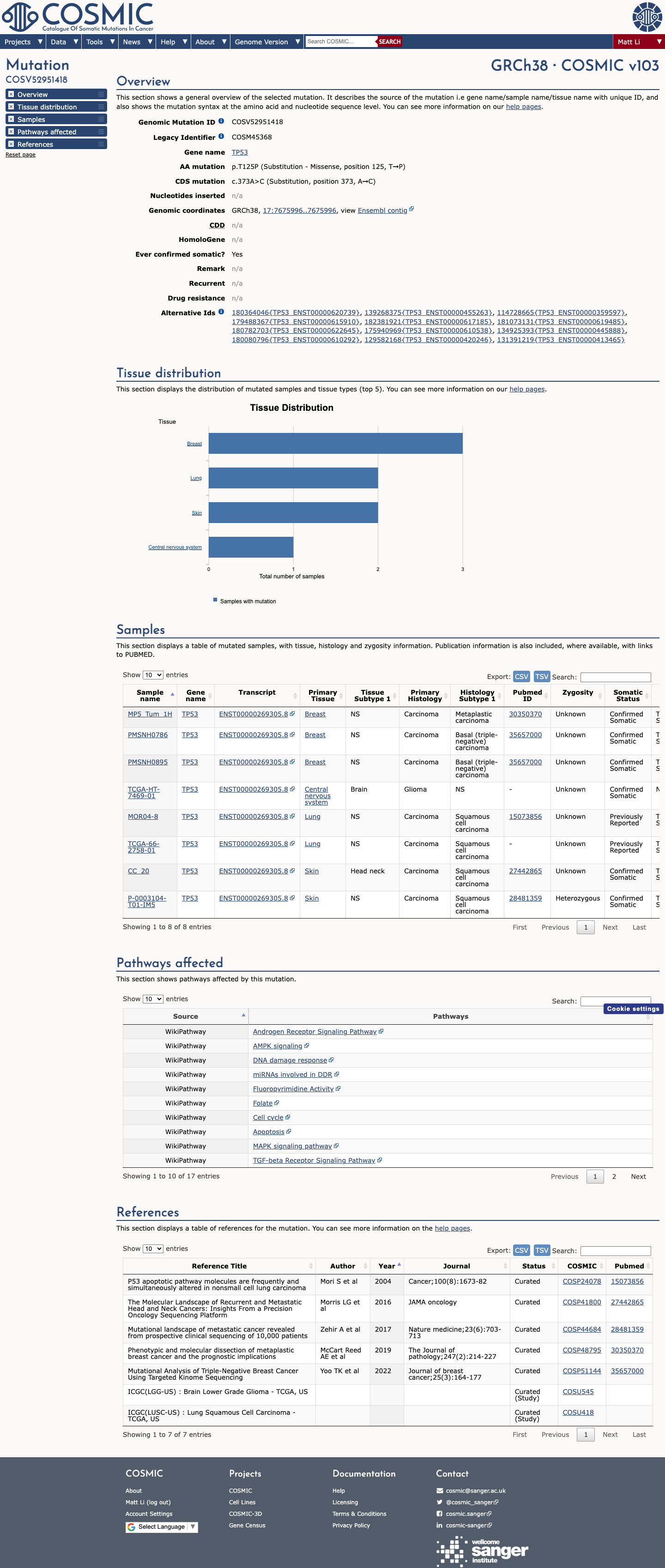
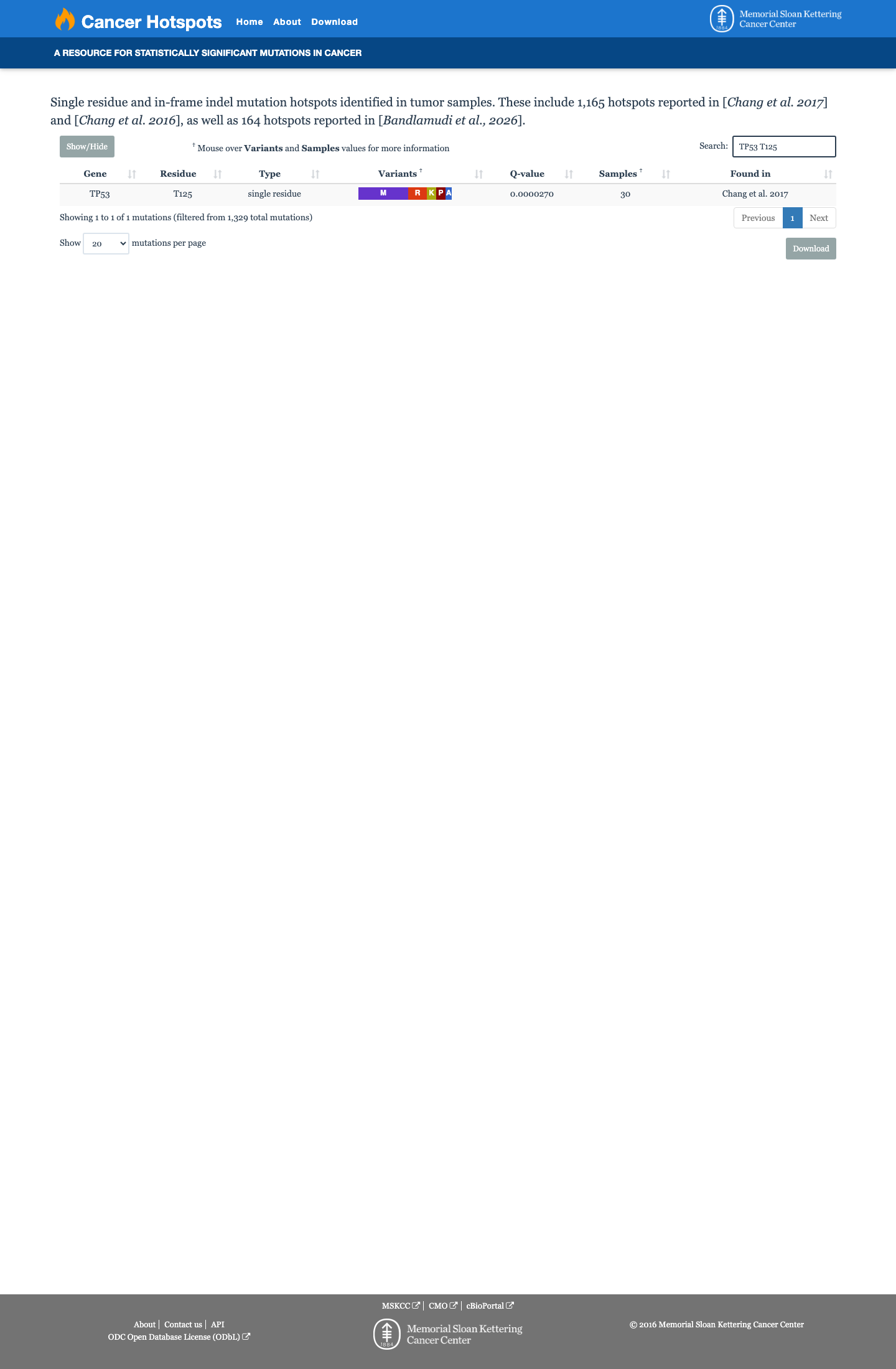
TP53 functional data support a damaging effect, with the TP53 VCEP functional worksheet listing p.(Thr125Pro) as non-functional with loss of function across eligible assays and assigning PS3.
PMID:12826609 ↗ PMID:29979965 ↗ PMID:30224644 ↗4
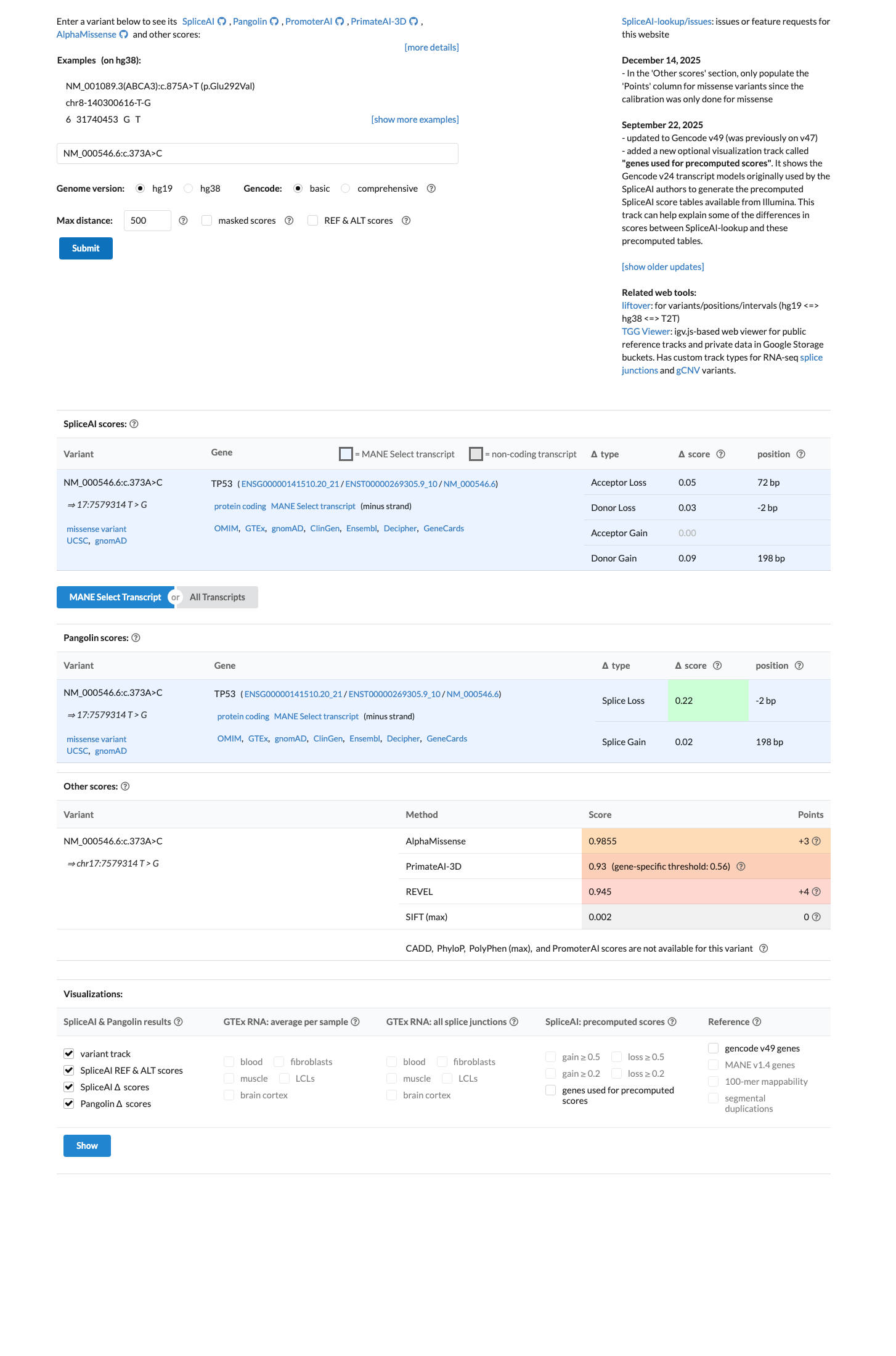
TP53 in silico evidence also supports deleteriousness: the TP53 VCEP PP3/BP4 worksheet assigns PP3 for c.373A>C with aGVGD Class C35 and BayesDel score 0.576078, while SpliceAI predicts no significant splice impact with a max delta score of 0.09.
spliceai ↗