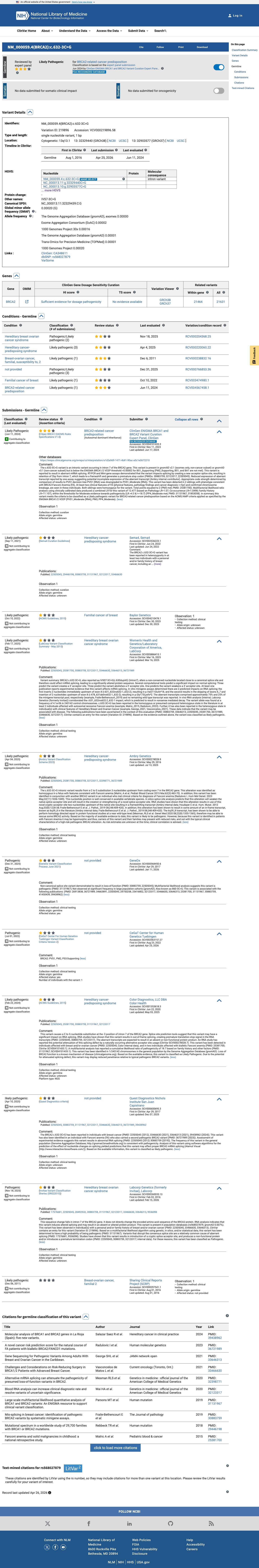
The BRCA2 c.632-3C>G (NP_000050.3:p.?) variant has been reported in ClinVar and is classified as Likely Pathogenic by the ClinGen ENIGMA BRCA1/2 expert panel.
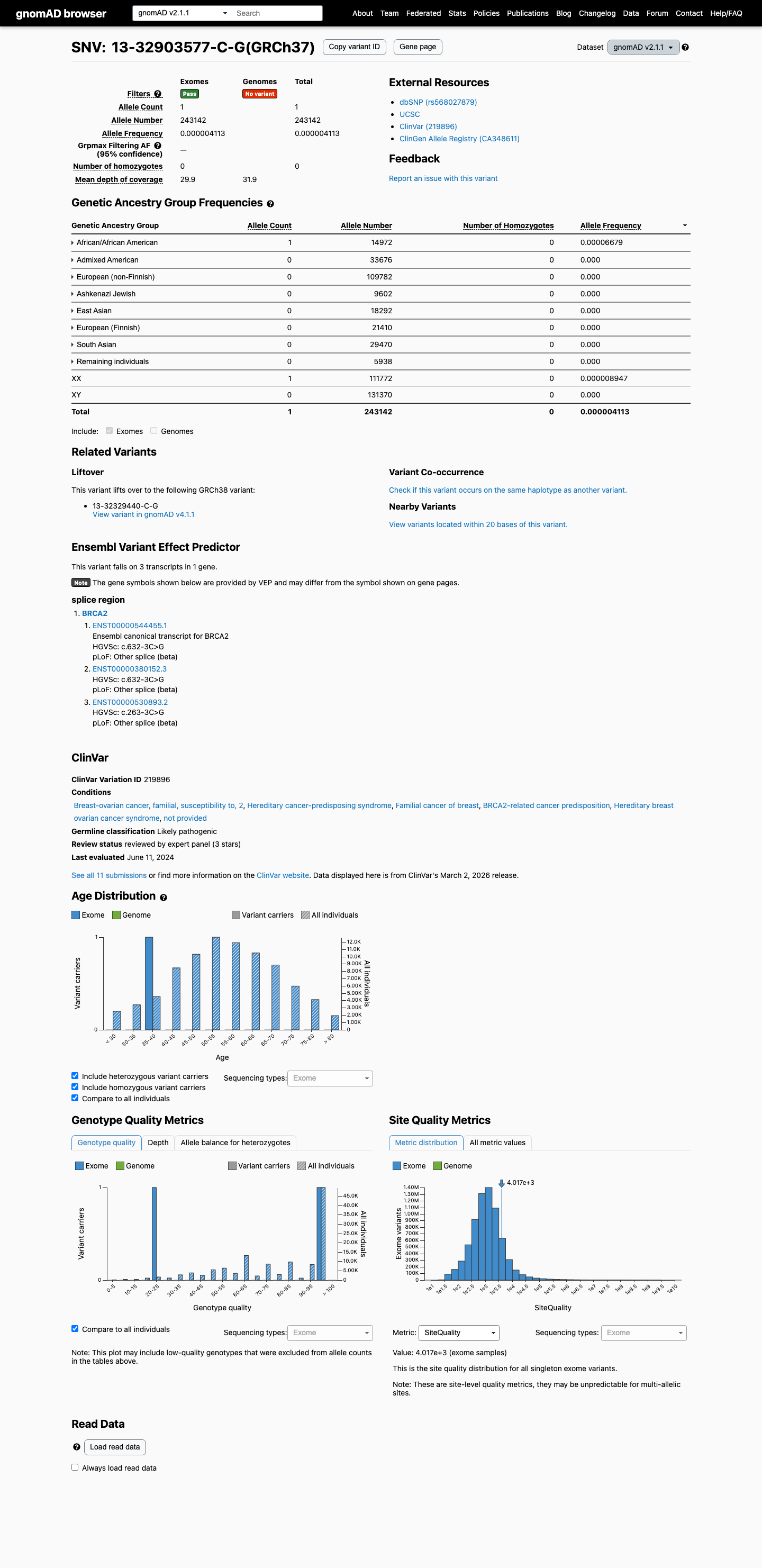
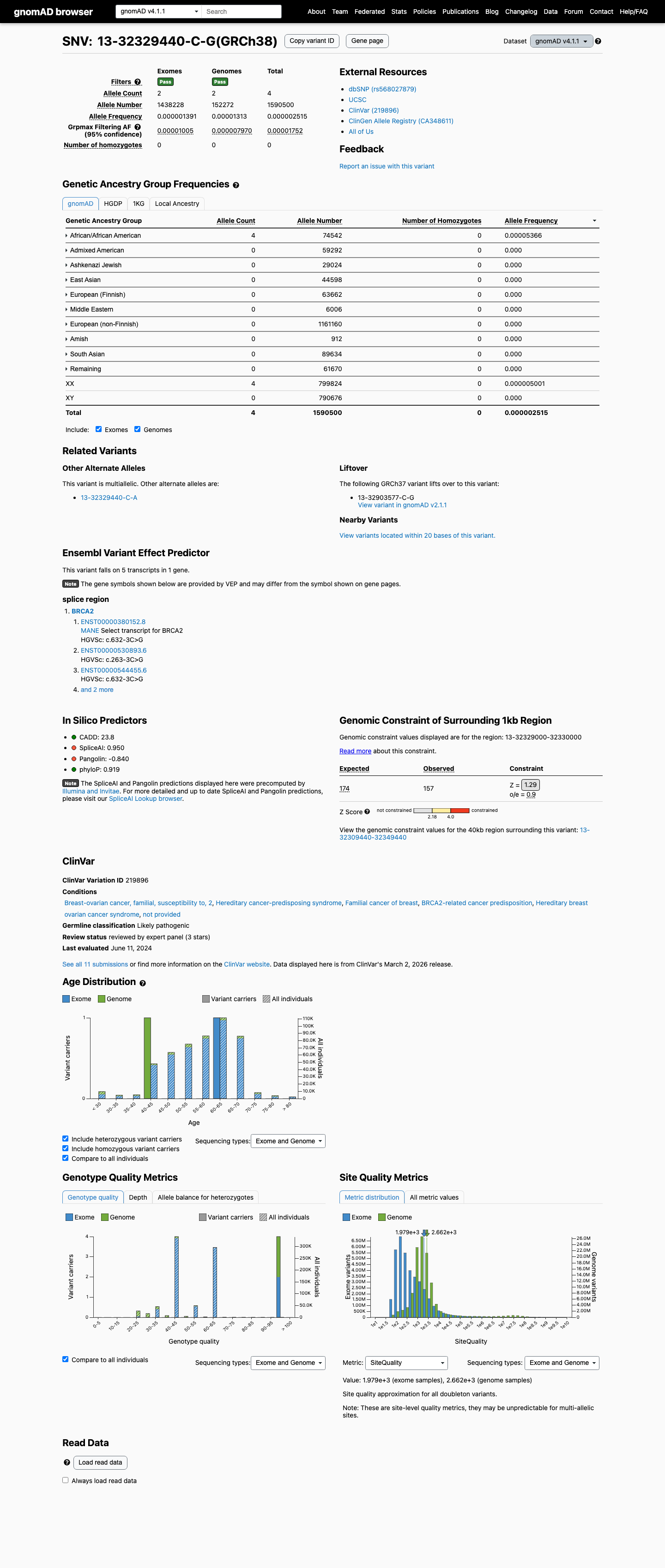
clinvar ↗This variant is rare in population databases, with 1/243142 alleles in gnomAD v2.1 (AF 0.00041%) and 4/1590500 alleles in gnomAD v4.1 (AF 0.00025%), which is below ENIGMA BA1 and BS1 thresholds but does not meet the absence requirement for PM2.
gnomad_v2 ↗ gnomad_v4 ↗RNA splicing studies reported complete splice disruption with retention of 2 intronic bases and no wild-type transcript detected from the variant allele, which is consistent with an RNA-based loss-of-function effect and supports PVS1_Strong (RNA) under the ENIGMA BRCA2 framework.
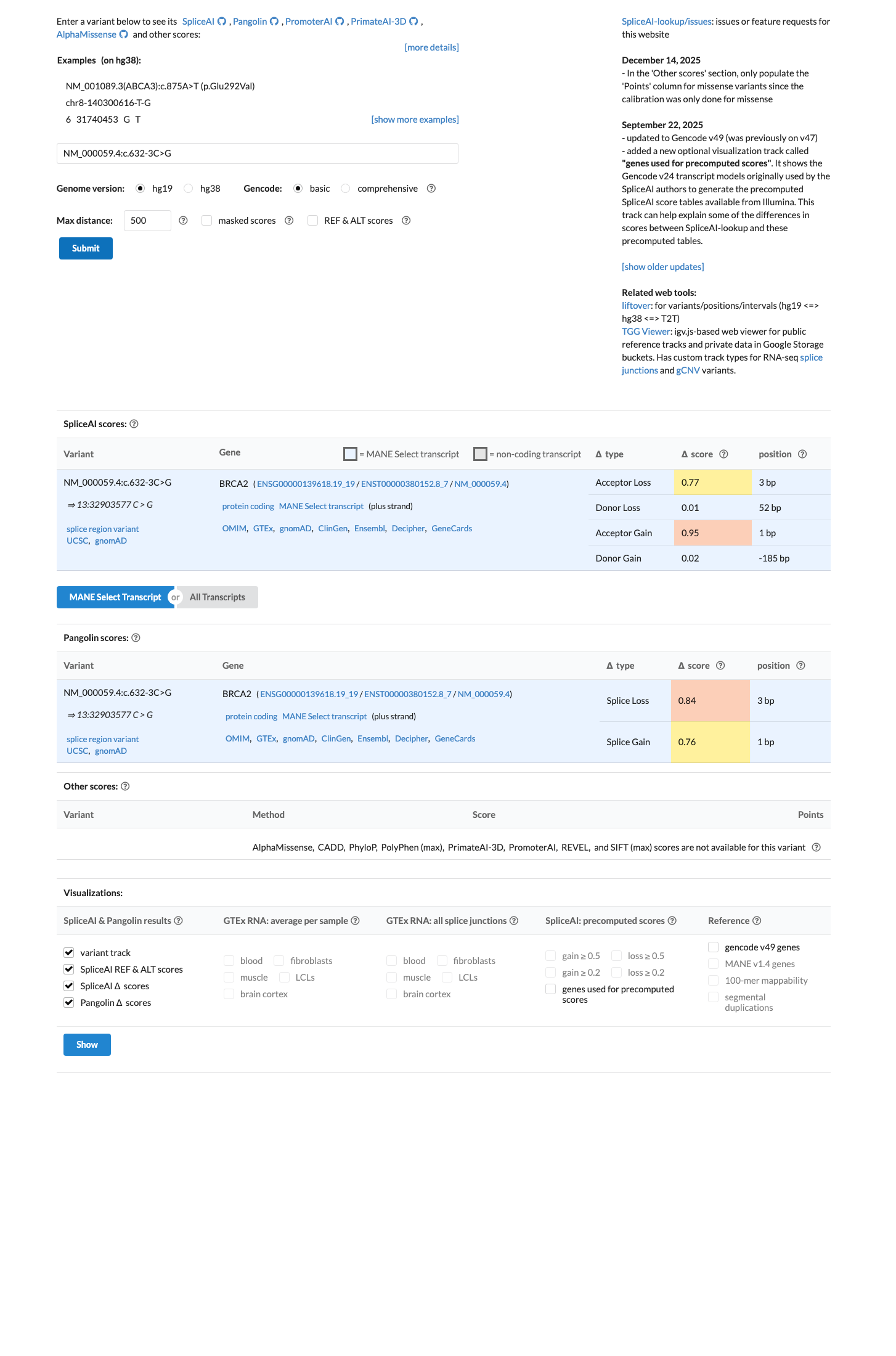
PMID:22505045 ↗In silico splicing prediction also supports a spliceogenic effect, with SpliceAI max delta score 0.95, well above the ENIGMA PP3 splice threshold of 0.2, although PP3 is not scored separately when PVS1 is met.
spliceai ↗