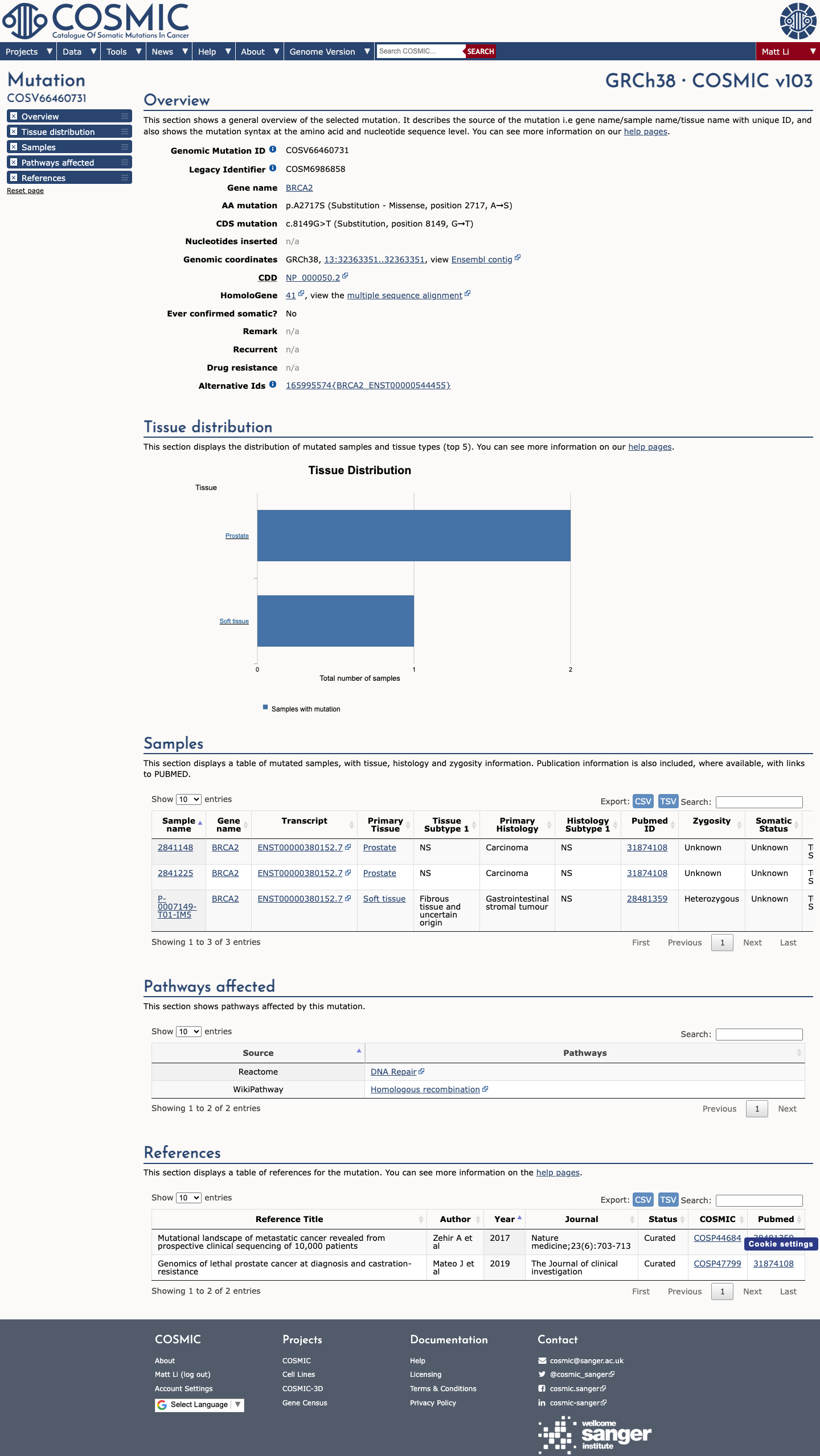
The BRCA2 c.8149G>T (p.Ala2717Ser) variant has been observed in somatic cancers in COSMIC (COSV66460731, n=3) and has also been reported in ClinVar, where the ClinGen ENIGMA BRCA1/2 expert panel classifies it as benign.
clinvar ↗This variant is common in population databases, with gnomAD v2.1 group-maximum filter allele frequency 0.00162114 and gnomAD v4.1 group-maximum filter allele frequency 0.00163682, both above the ENIGMA BA1 threshold of 0.001.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Calibrated BRCA2 functional evidence supports a benign effect: ENIGMA Table 9 assigns BS3 Strong based on three studies showing function similar to benign controls, with no aberrant RNA result listed.
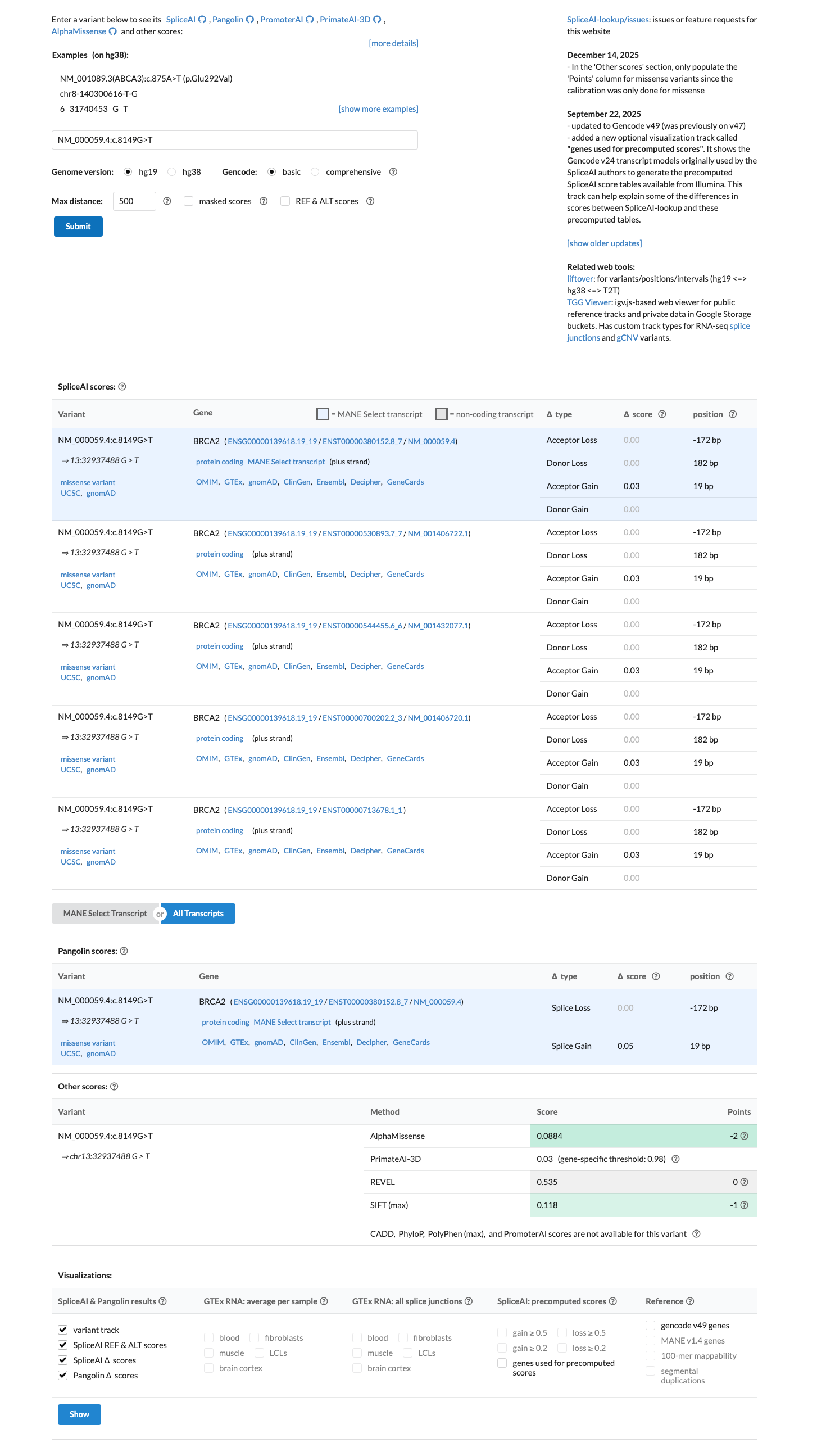
cspec ↗SpliceAI predicts no significant splice effect for this variant, with a maximum delta score of 0.03, which argues against splice disruption.
spliceai ↗