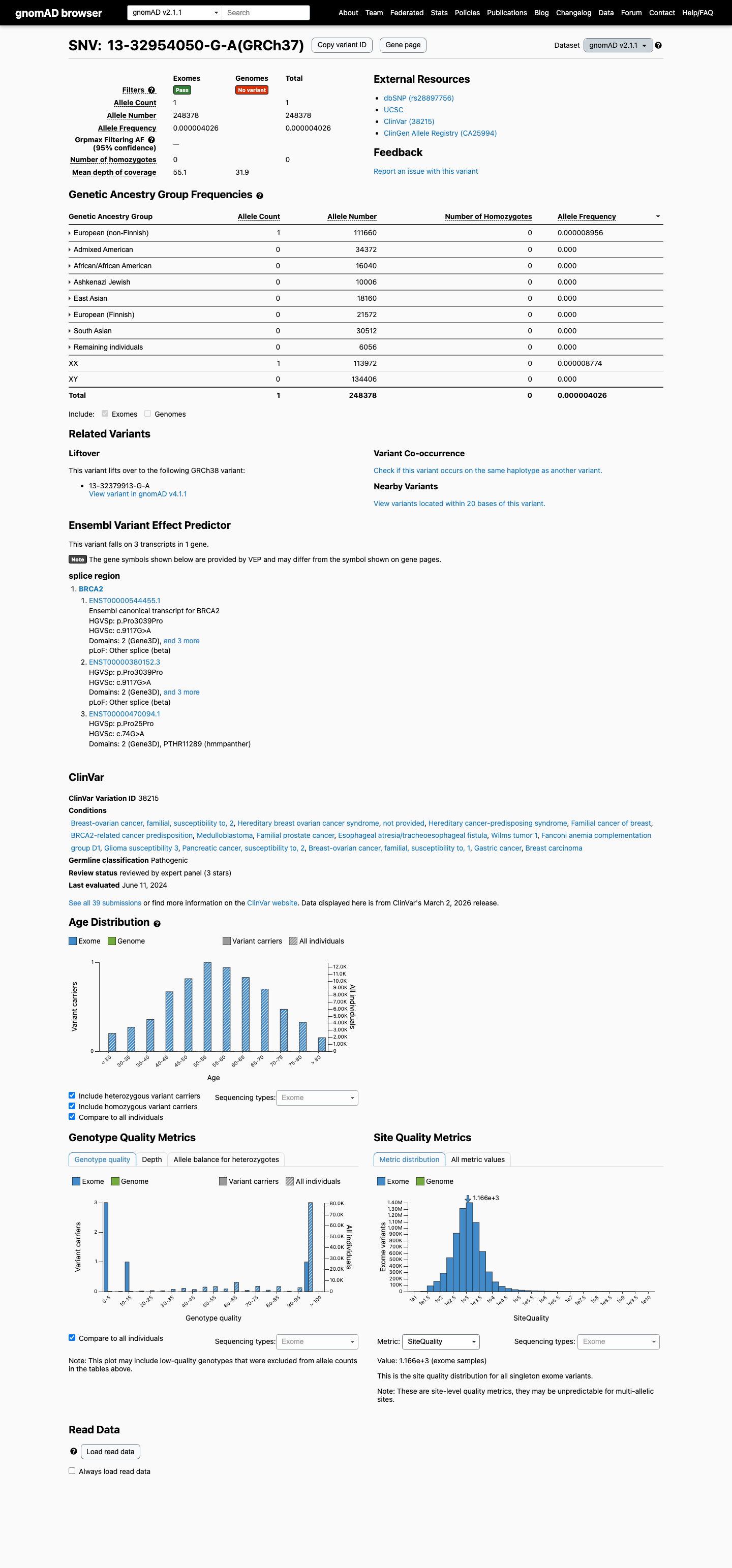
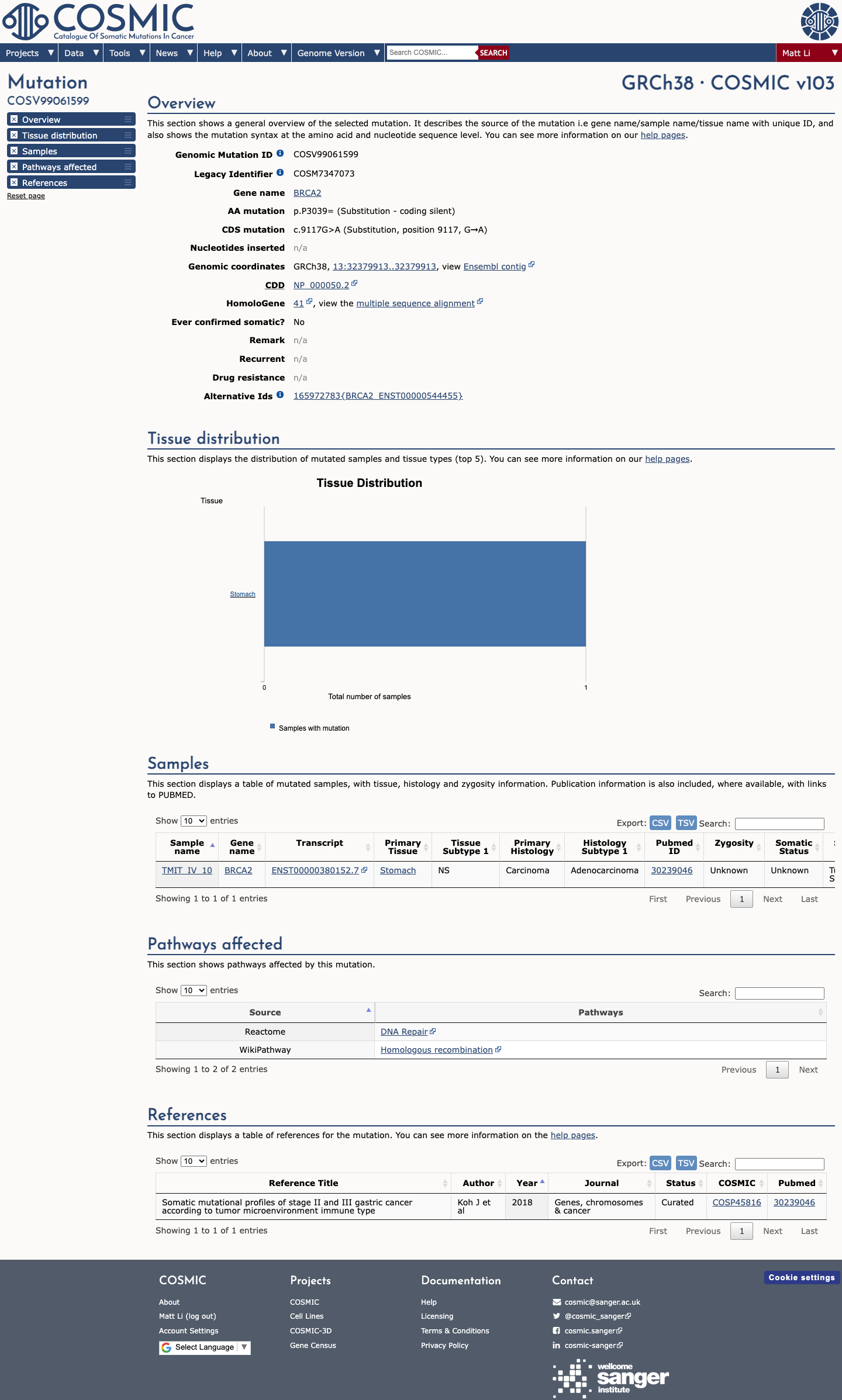
The BRCA2 c.9117G>A (p.(Pro3039=), p.(P3039=)) variant has been reported in ClinVar as pathogenic, including an expert-panel pathogenic classification from the ClinGen ENIGMA BRCA1/2 Variant Curation Expert Panel.
clinvar ↗This variant is rare in population databases but is not absent from controls, with 1/248378 alleles in gnomAD v2.1 (AF 0.00000403) and 6/1613032 alleles in gnomAD v4.1 (AF 0.00000372), so PM2 is not met and BA1/BS1 thresholds are not reached.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Clinical-history likelihood-ratio analysis for BRCA2 c.9117G>A showed LR 3.91 in 12 probands, which exceeds the ENIGMA PP4 supporting threshold of 2.08 and supports PP4 at supporting strength.
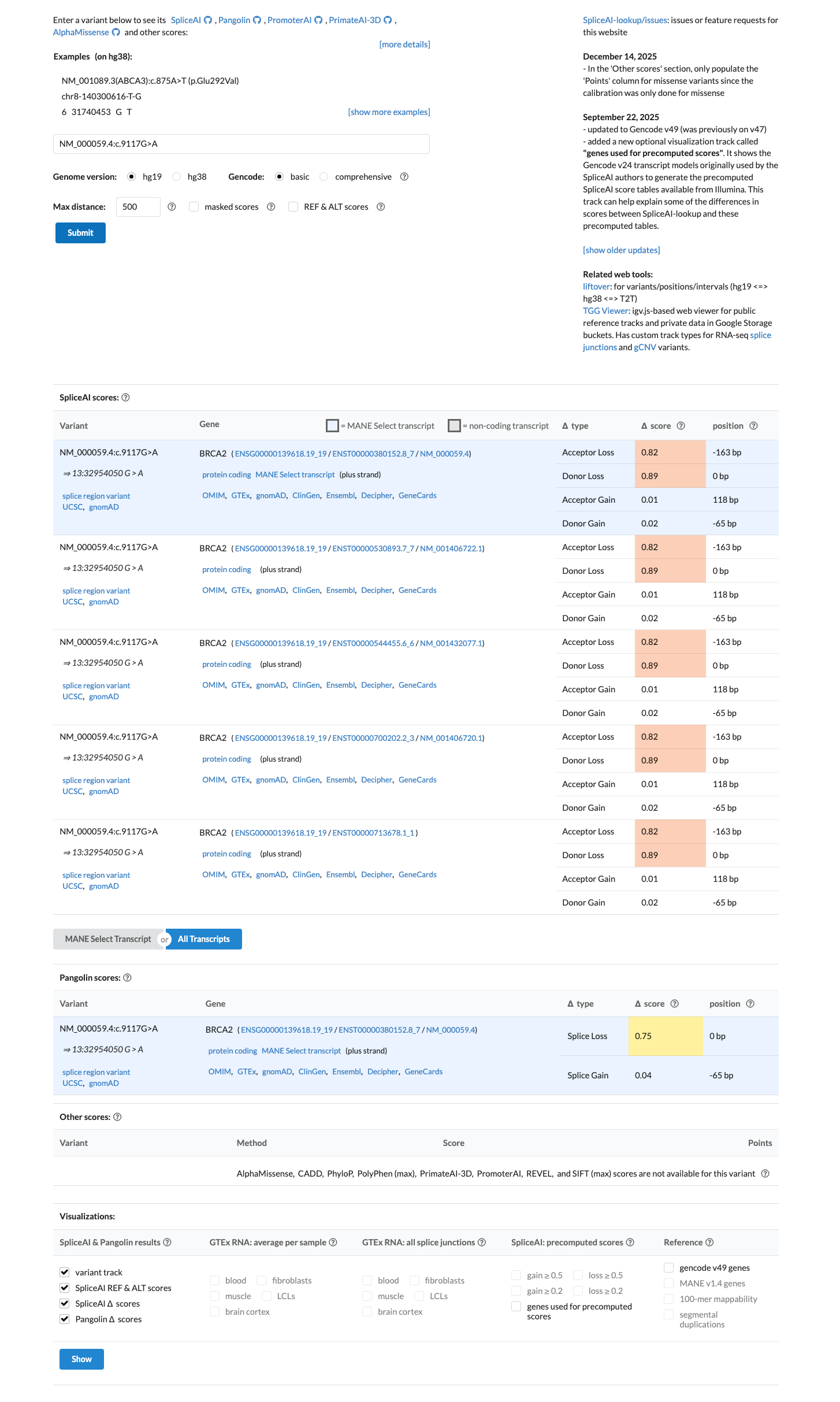
PMID:31853058 ↗ cspec ↗SpliceAI predicts a strong splice effect for this synonymous variant, with a maximum delta score of 0.89, which is above the ENIGMA PP3 threshold of 0.20 for predicted splicing impact and argues against BP4/BP7.
spliceai ↗ cspec ↗