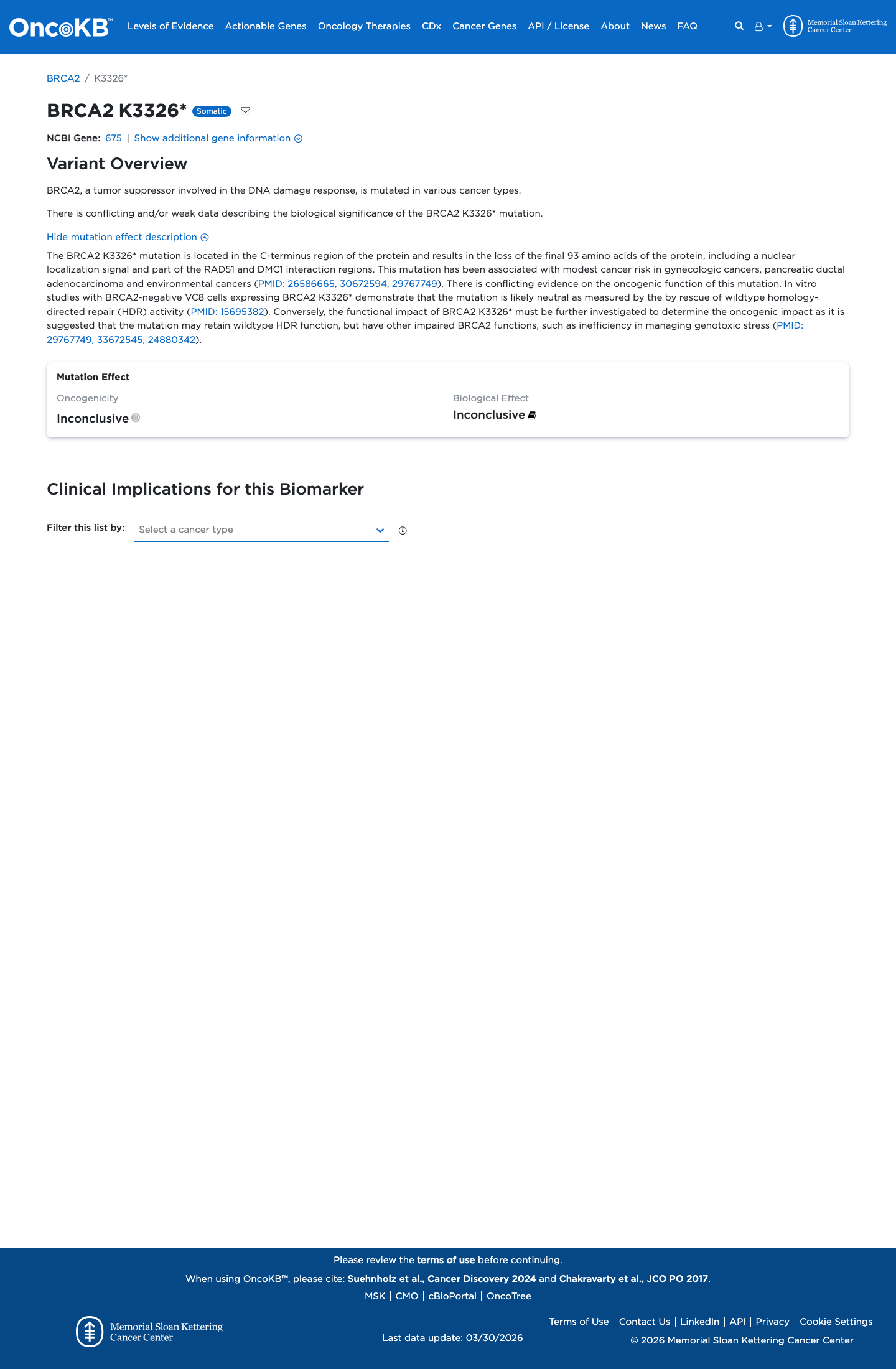
The BRCA2 c.9976A>T (p.Lys3326Ter, K3326*) variant has been observed in somatic cancers in COSMIC (18 occurrences) and has been reported in ClinVar with an expert-panel benign classification.
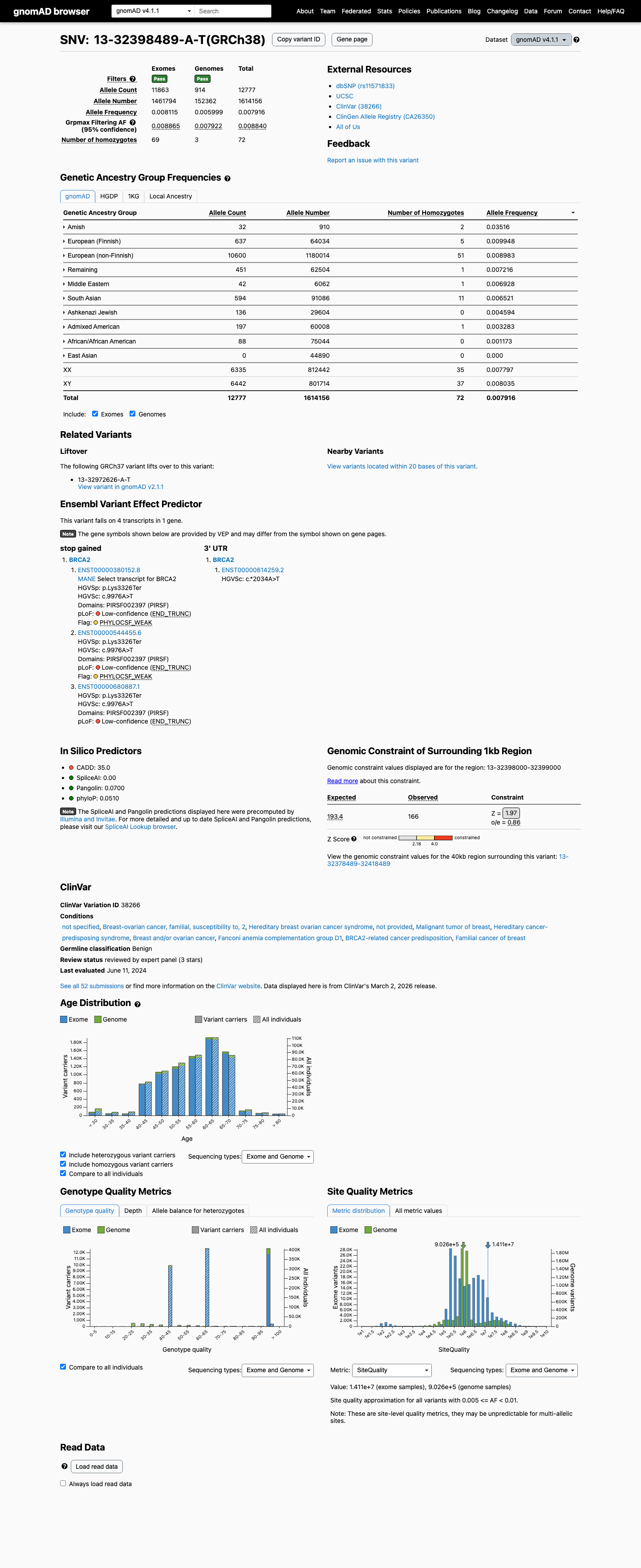
clinvar ↗This variant is common in population databases, including gnomAD v2.1 with an overall allele frequency of 0.64680% and grpmax FAF of 0.849119%, and gnomAD v4.1 with an overall allele frequency of 0.79156%; these values are above the BRCA2 ENIGMA BA1 (>0.1%) and BS1 (>0.01%) thresholds.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Calibrated BRCA2 functional evidence supports a benign effect, with the expert specification assigning BS3 Strong to this exact variant based on protein function similar to benign controls.
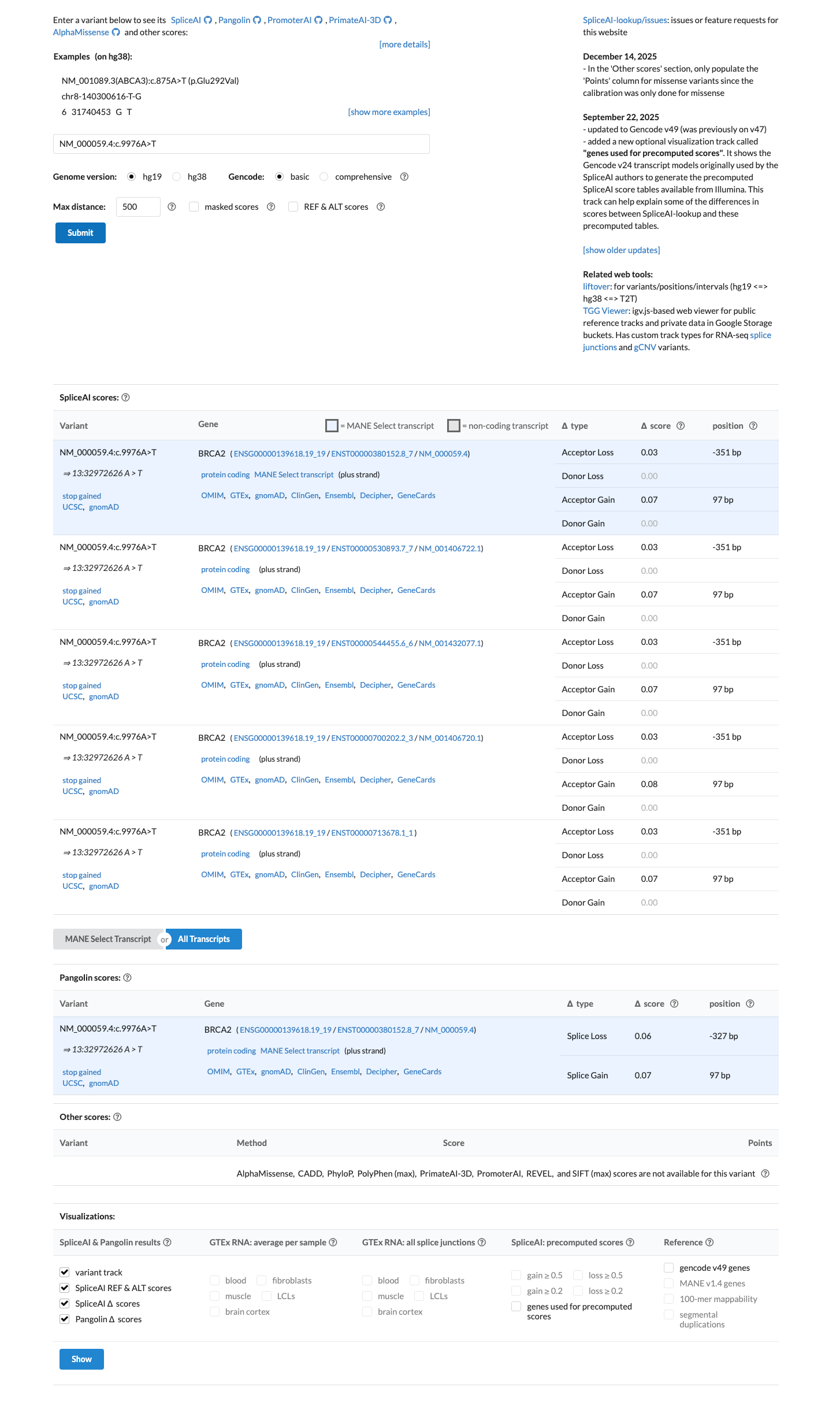
cspec ↗SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.07.
spliceai ↗