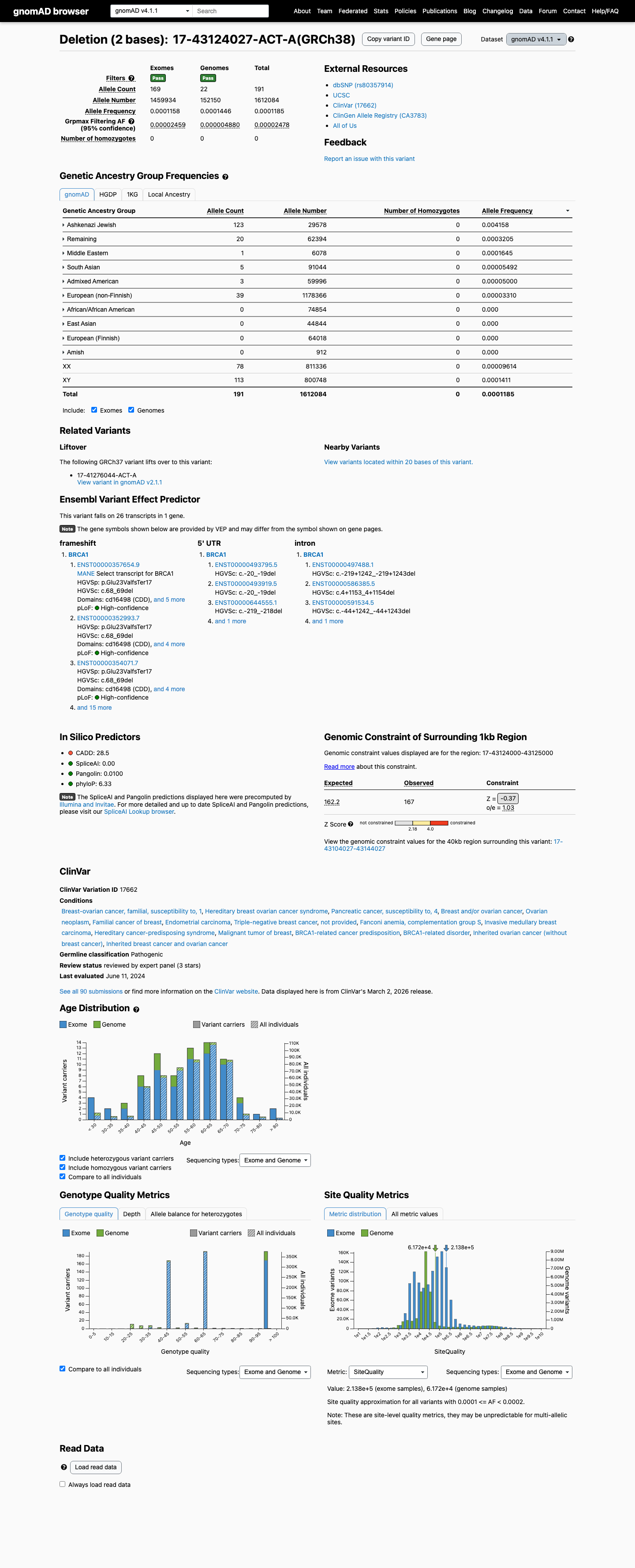
This variant is present in population databases, with gnomAD v2.1 AF 0.000205352 (58/282442 alleles) and gnomAD v4.1 AF 0.00011848 (191/1612084 alleles); the observed filter allele frequencies of 5.395e-05 in v2.1 and 2.478e-05 in v4.1 are below the BA1 threshold of 0.001 but within the ENIGMA BS1_Supporting range above 0.00002 and less than or equal to 0.0001.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗ENIGMA functional data assign PS3 Strong to this variant, with calibrated assay evidence showing a damaging effect consistent with a deleterious control profile.
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.00, which is consistent with the primary effect being protein truncation rather than splice disruption.
spliceai ↗This early exon 2 frameshift introduces a premature termination codon at p.Glu23ValfsTer17, and ENIGMA exon-level rules support PVS1 at Very Strong strength and PM5 for protein-truncating variants in this exon.
cspec ↗