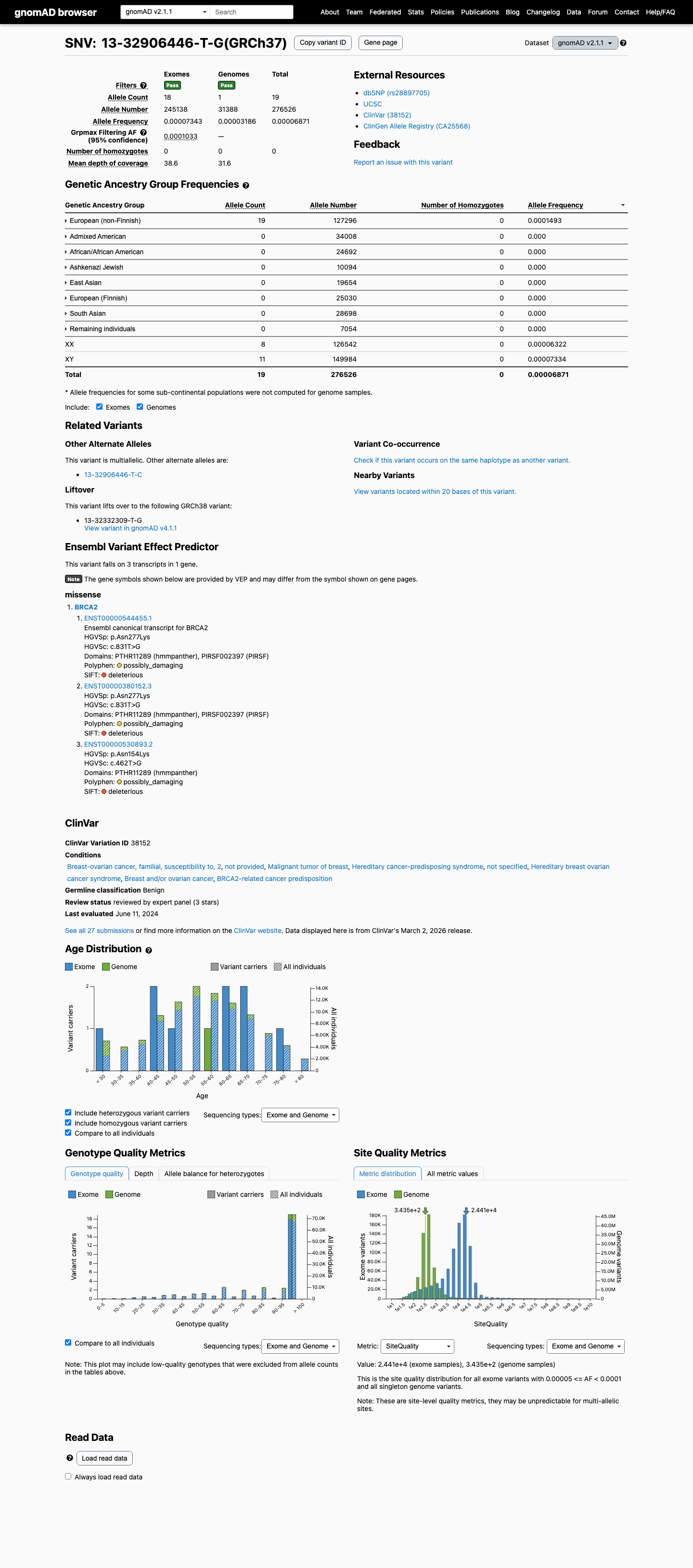
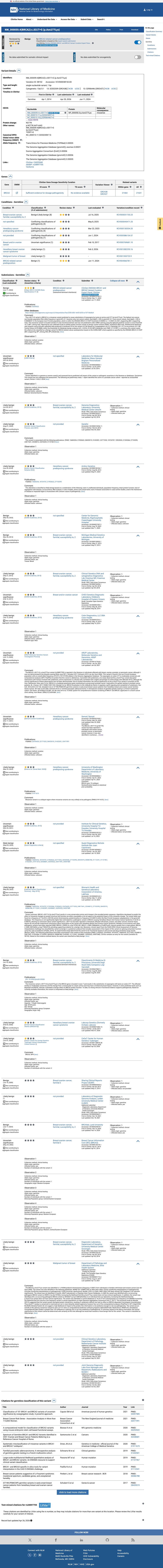
The BRCA2 c.831T>G (p.Asn277Lys) variant has been reported in ClinVar and is classified as Benign by the ClinGen ENIGMA BRCA1/2 expert panel.
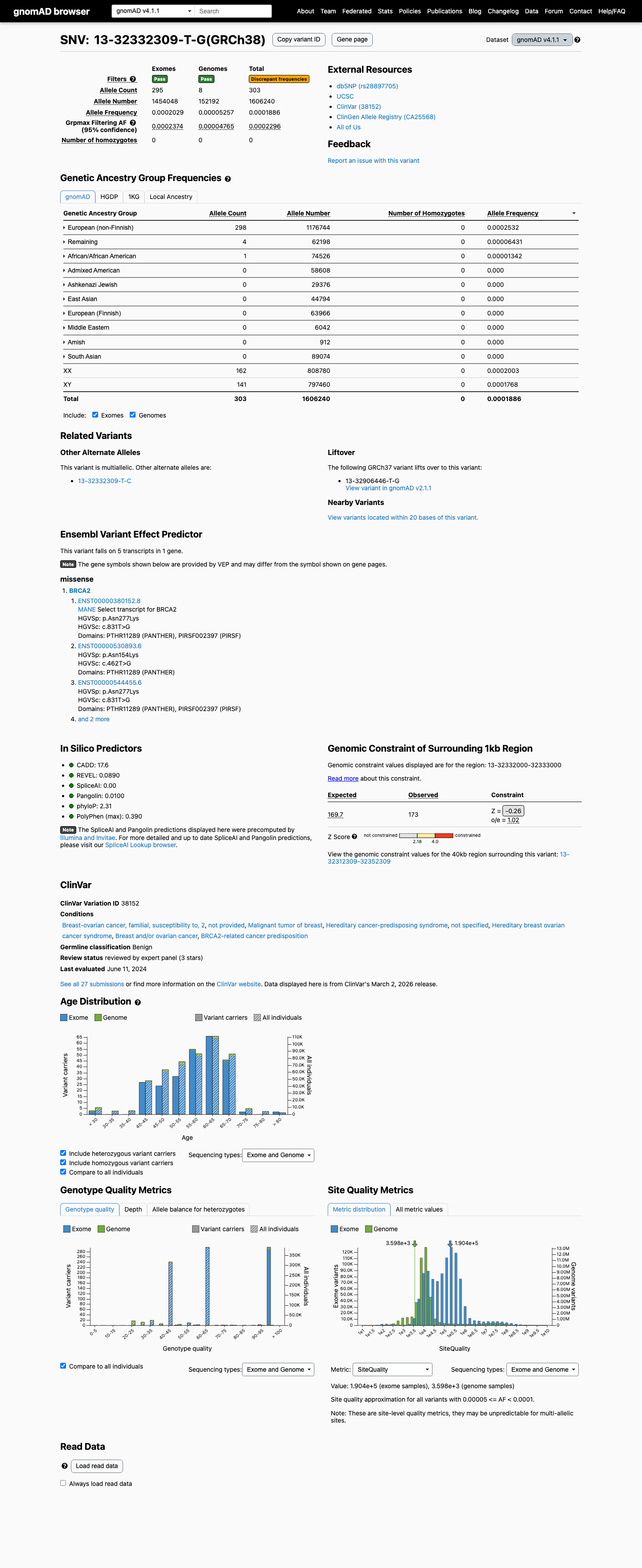
clinvar ↗This variant is present in population databases, including gnomAD v2.1 at AF 0.0000687 with a highest observed filter allele frequency of 0.0001033 and gnomAD v4.1 at AF 0.0001886 with a highest observed filter allele frequency of 0.0002296, supporting BS1.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Calibrated functional data in the ENIGMA BRCA1/2 specification support no damaging effect on protein function for p.(Asn277Lys), supporting BS3.
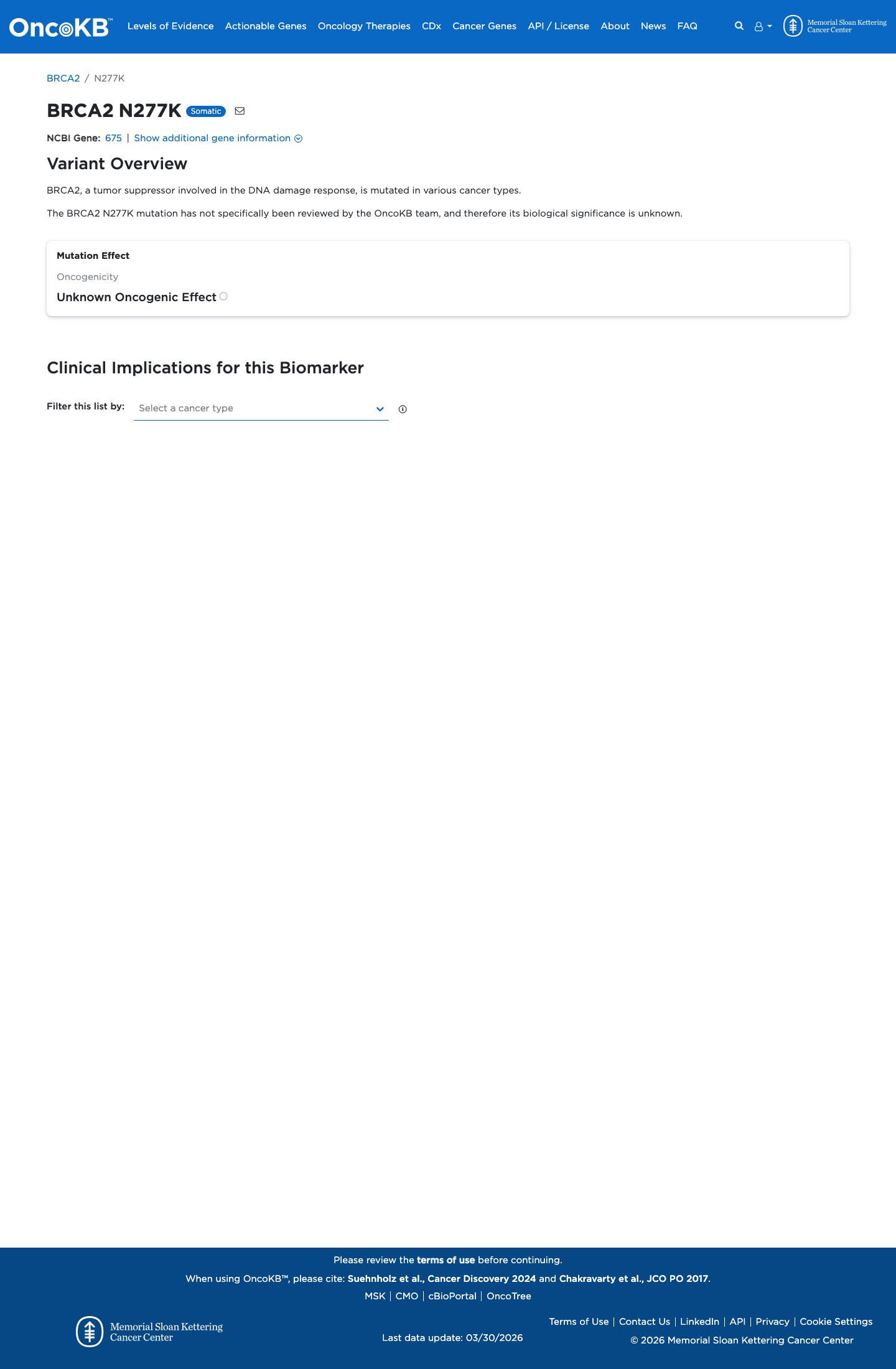
cspec ↗In silico evidence does not suggest splice disruption, with a SpliceAI maximum delta score of 0.00, and the missense change lies outside the BRCA2 clinically important domains used by ENIGMA for missense interpretation, supporting BP1_Strong.
spliceai ↗ cspec ↗