Classification rationale
1
2
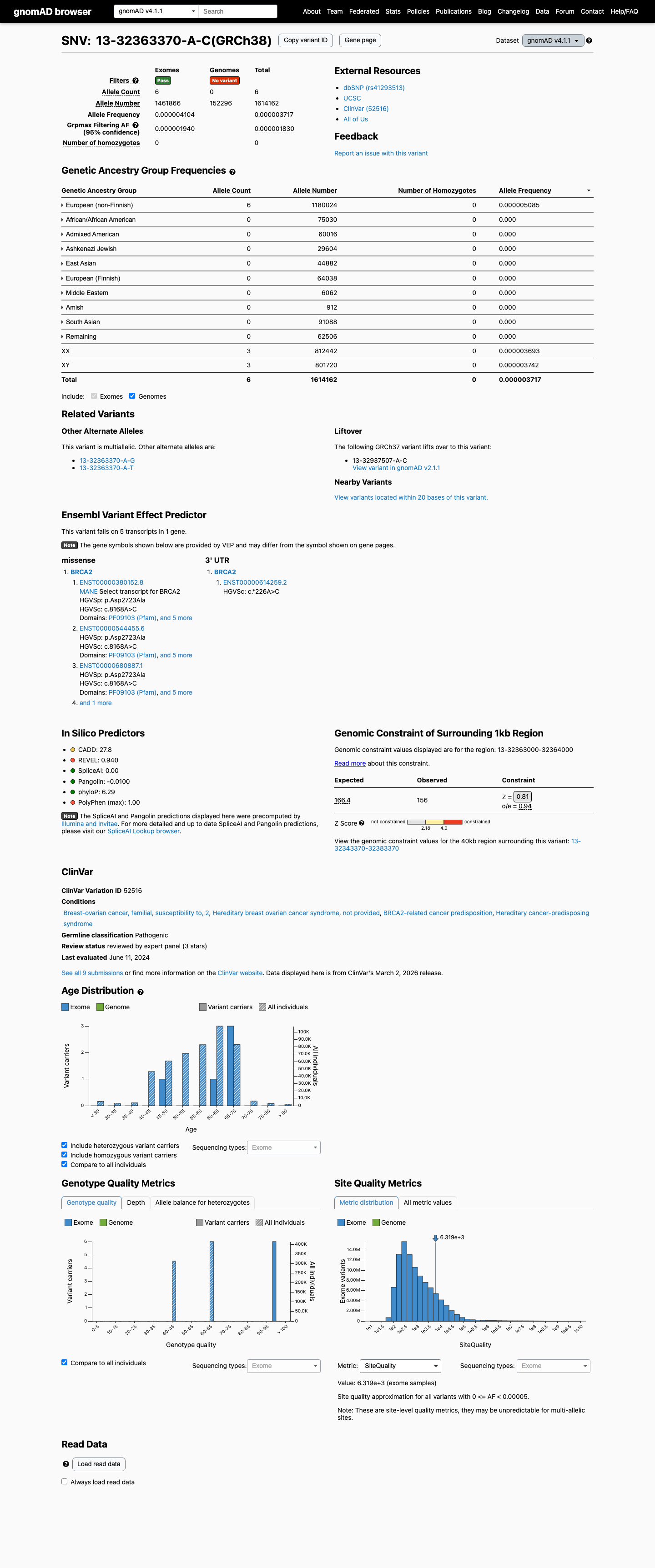
This variant is present at very low frequency in population databases, with 1/251098 alleles in gnomAD v2.1 and 6/1614162 alleles in gnomAD v4.1, which is too low to support benign frequency criteria but means PM2 is not met because the variant is not absent from controls.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
Calibrated BRCA2 functional evidence supports a damaging effect, and the ENIGMA curated functional table assigns PS3 at Strong strength for p.Asp2723Ala.
cspec ↗4
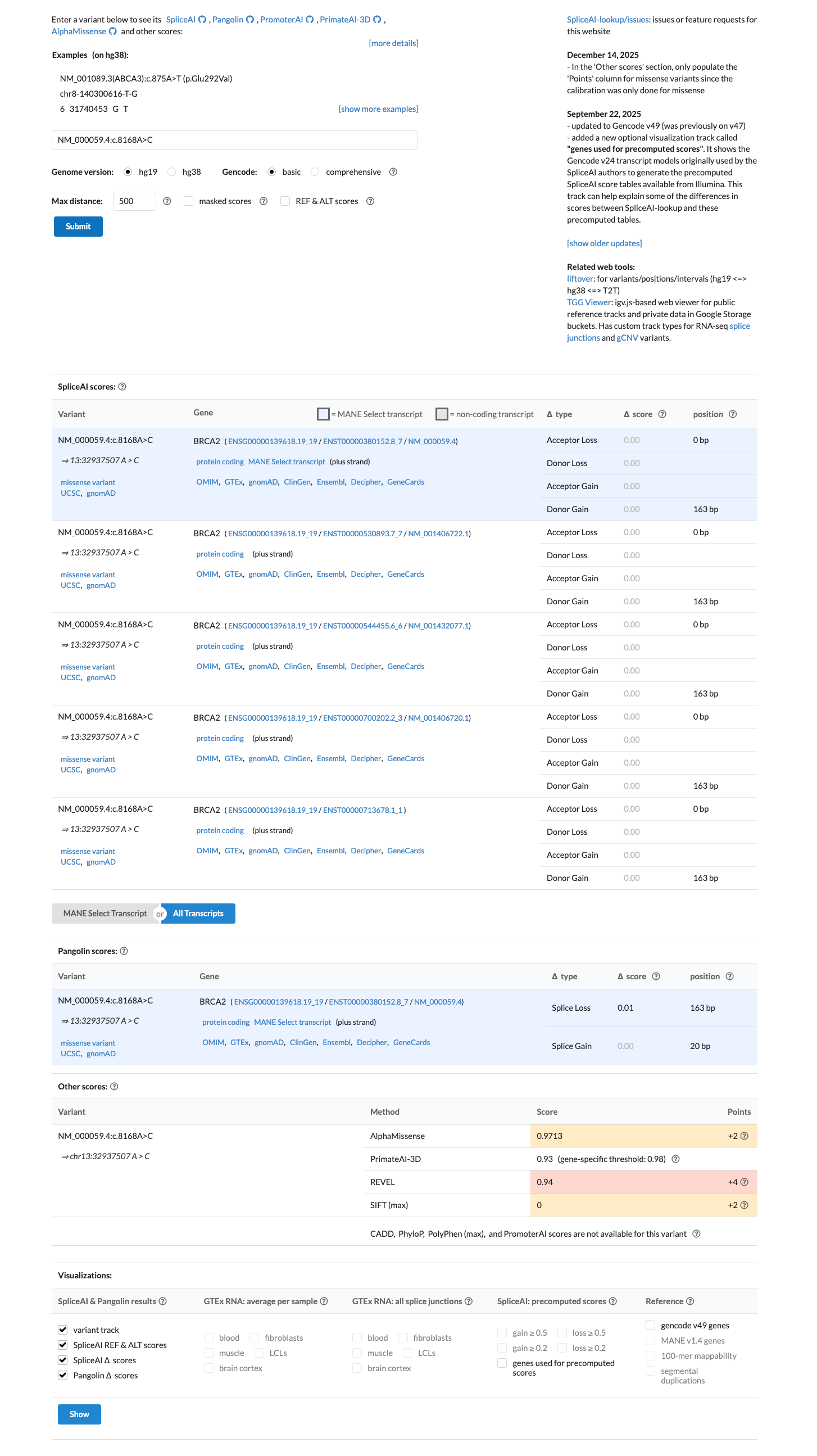
This missense change occurs within the BRCA2 DNA-binding domain, SpliceAI predicts no splice disruption with a max delta score of 0.00, and the computational profile supports PP3 rather than BP4.
cspec ↗ spliceai ↗