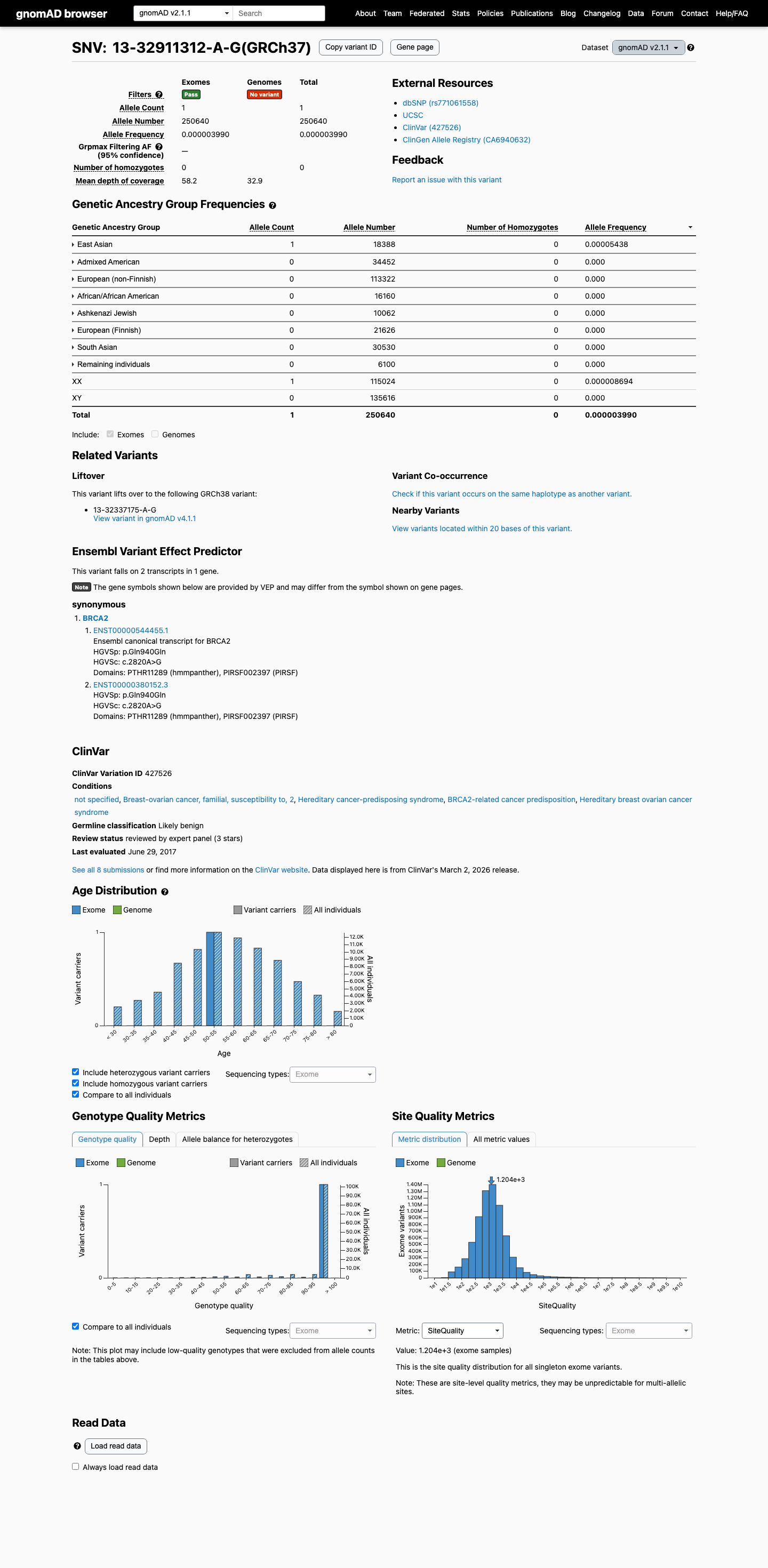
The BRCA2 NM_000059.4:c.2820A>G (p.(Gln940=), p.(Q940=)) variant has been reported in ClinVar as likely benign, including review by the ENIGMA expert panel and multiple clinical laboratories.
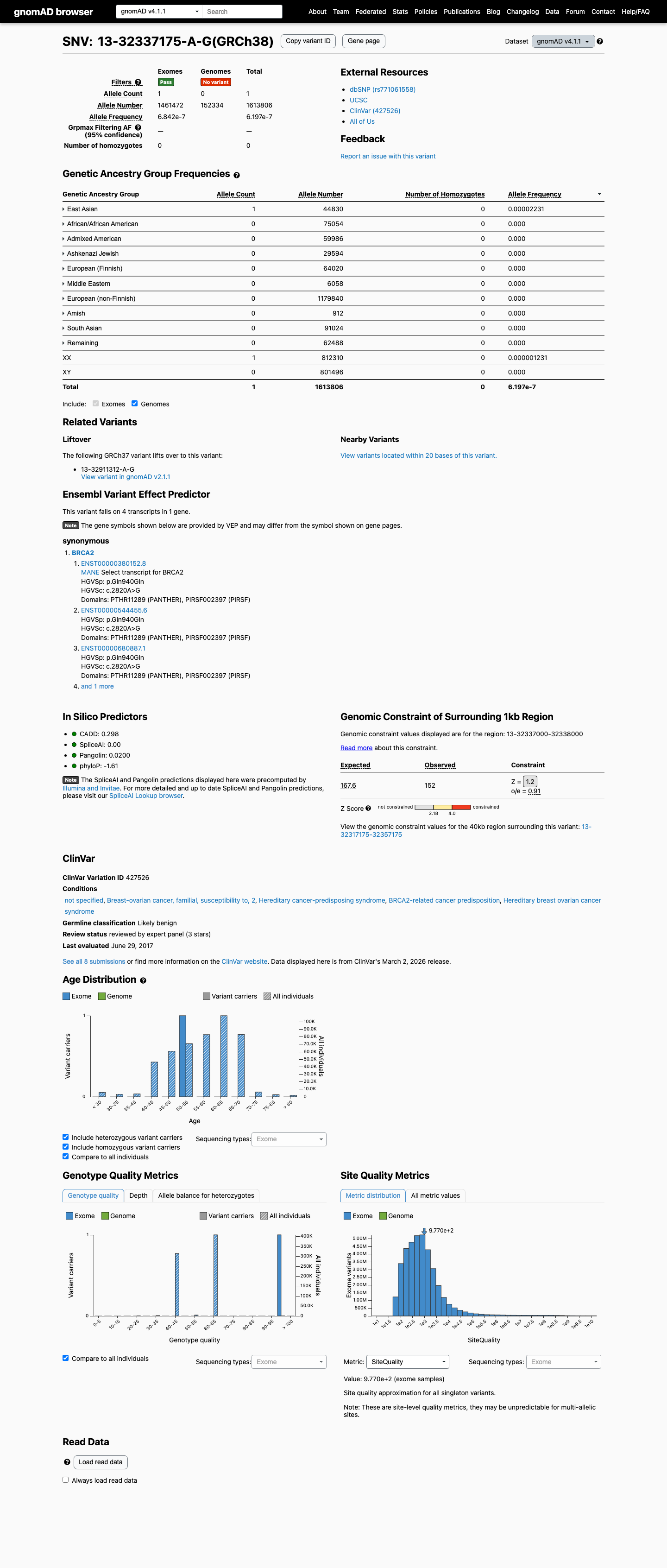
clinvar ↗This variant is present at very low frequency in population databases, with 1/250640 alleles in gnomAD v2.1 (AF 0.00040%) and 1/1613806 alleles in gnomAD v4.1 (AF 0.00006%), which does not meet ENIGMA benign stand-alone or strong population thresholds and does not satisfy PM2 absence criteria.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In silico splicing analysis predicts no significant splice impact, with a SpliceAI maximum delta score of 0.00, and the synonymous p.(Gln940=) change lies outside the BRCA2 clinically important domains defined by ENIGMA, supporting BP1_Strong and arguing against PP3.
spliceai ↗ cspec ↗