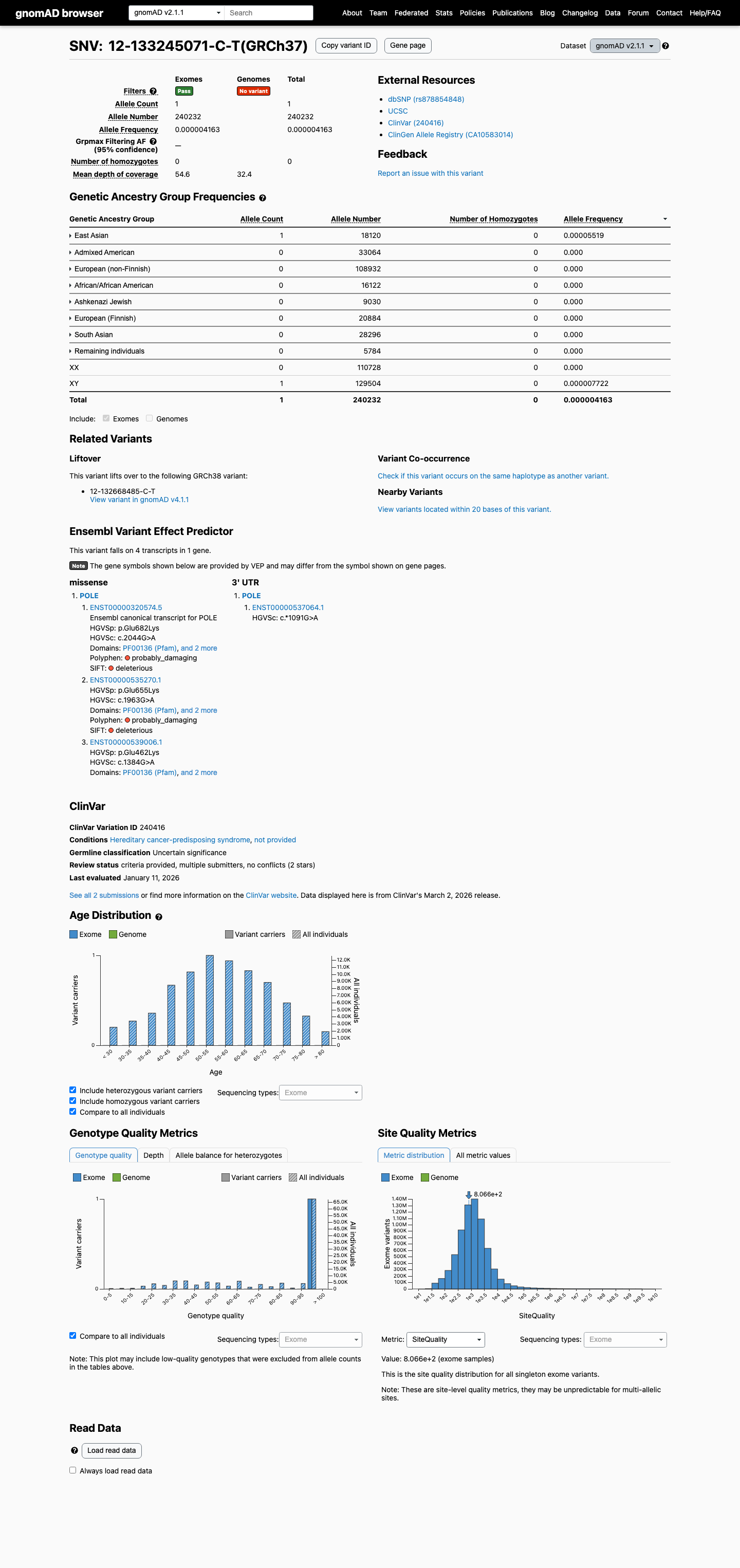
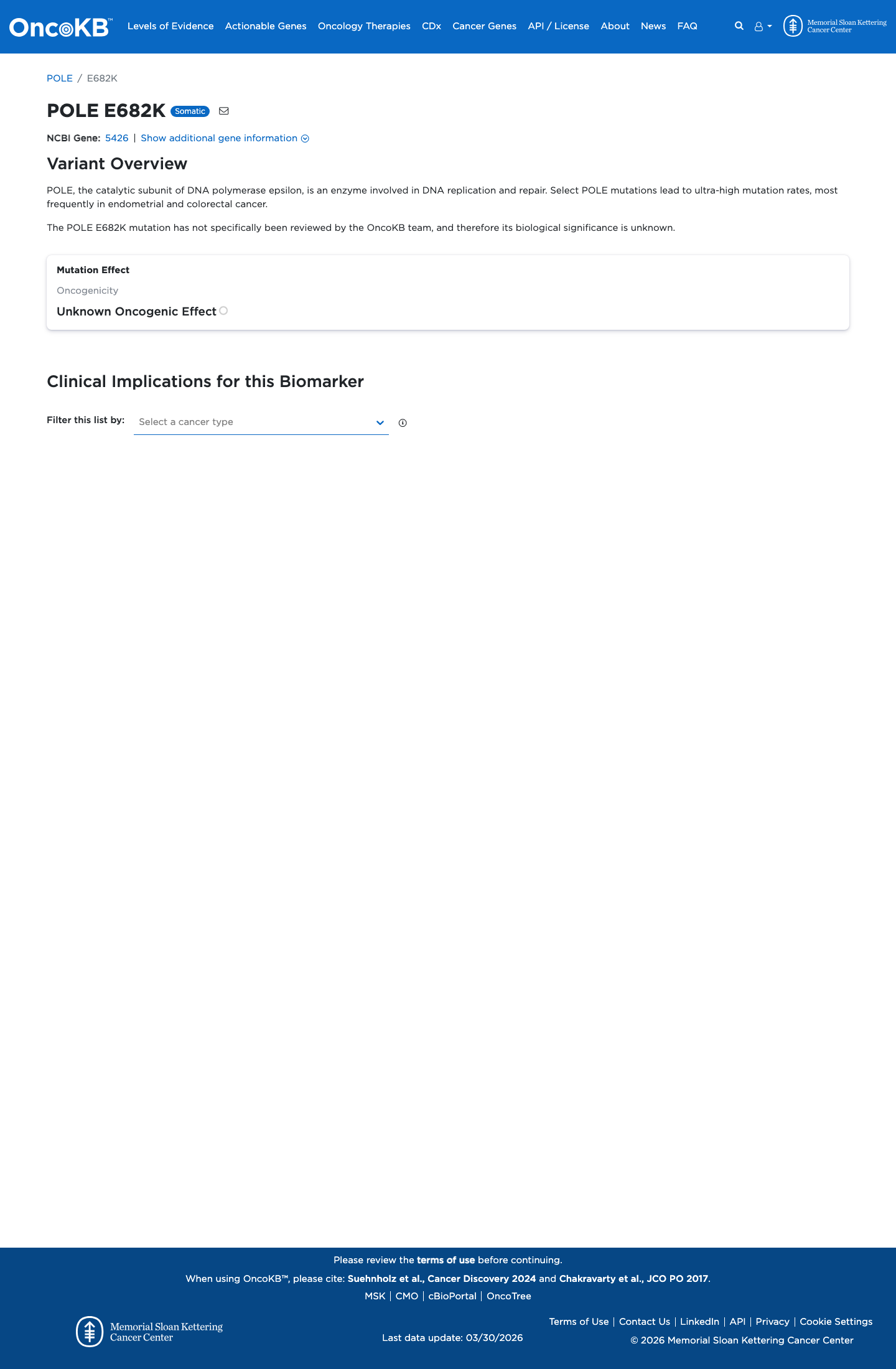
The POLE c.2044G>A (p.Glu682Lys; p.E682K) variant has not been identified in the León-Castillo recurrent COSMIC/TCGA endometrial carcinoma POLE variant table and has been reported in ClinVar as a variant of uncertain significance.
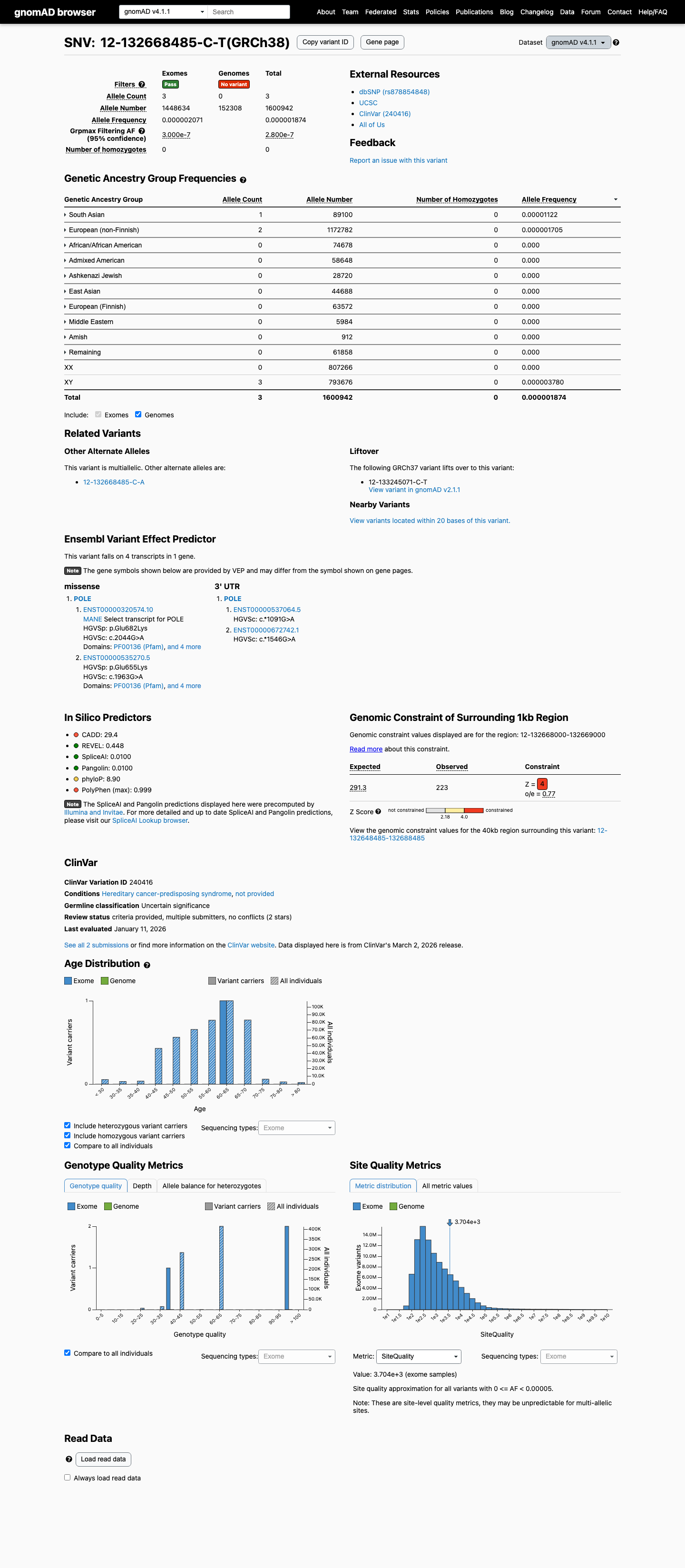
clinvar ↗This variant is present at very low frequency in gnomAD, with AF 4.16e-06 in v2.1 (1/240232 alleles) and AF 1.87e-06 in v4.1 (3/1600942 alleles), which is below the project's PM2 threshold of 0.1% and below benign frequency thresholds.
gnomad_v2 ↗ gnomad_v4 ↗This missense change is not one of the exact POLE hotspot or recurrent exonuclease-domain substitutions specified by the León-Castillo framework for PM1 or PS4.
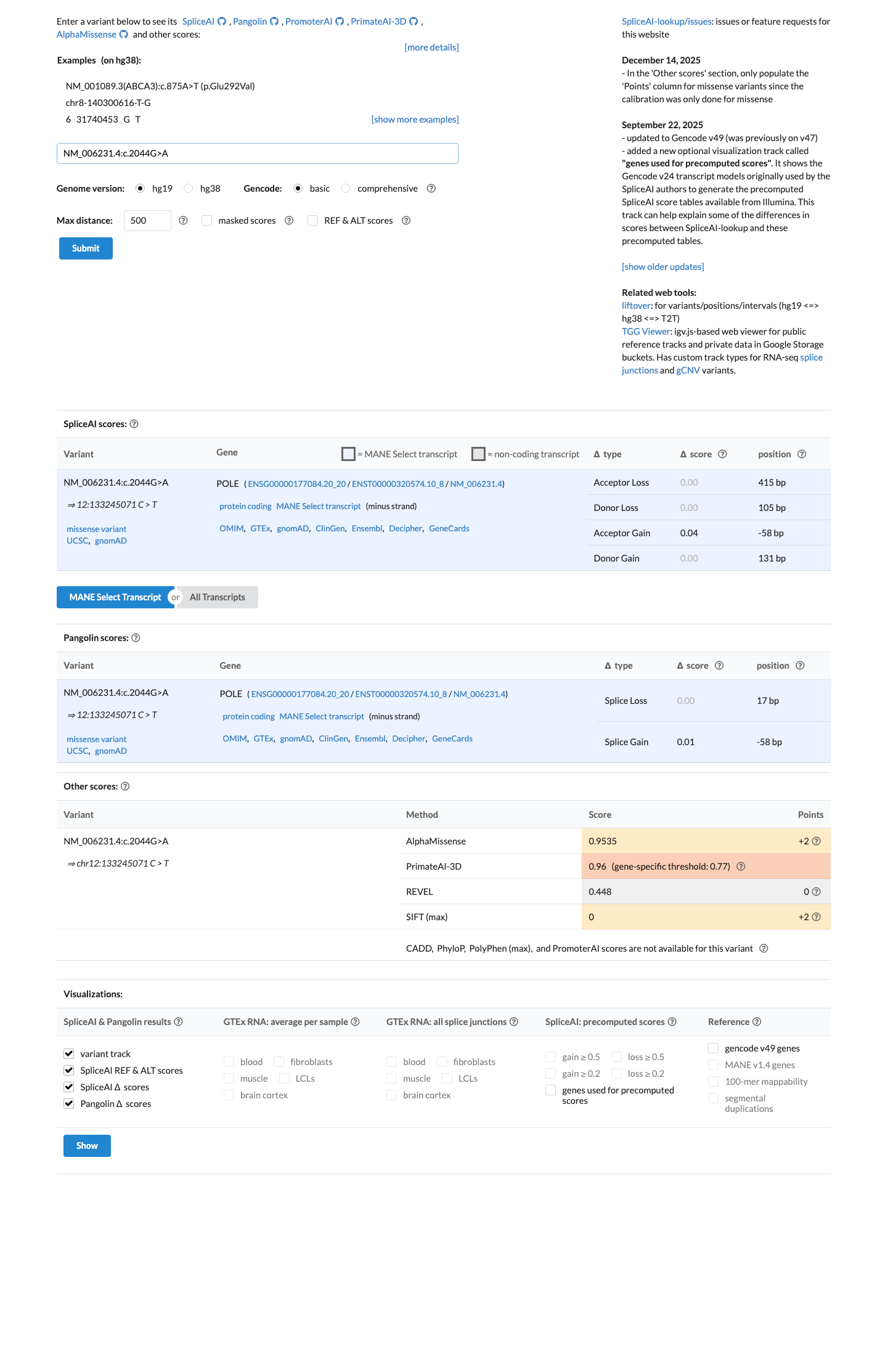
Available computational evidence does not establish a POLE-specific deleterious or benign pattern for this exact variant because it was not identified in the León-Castillo supplementary computational tables, and SpliceAI predicts no significant splice impact (max delta score 0.04).
spliceai ↗