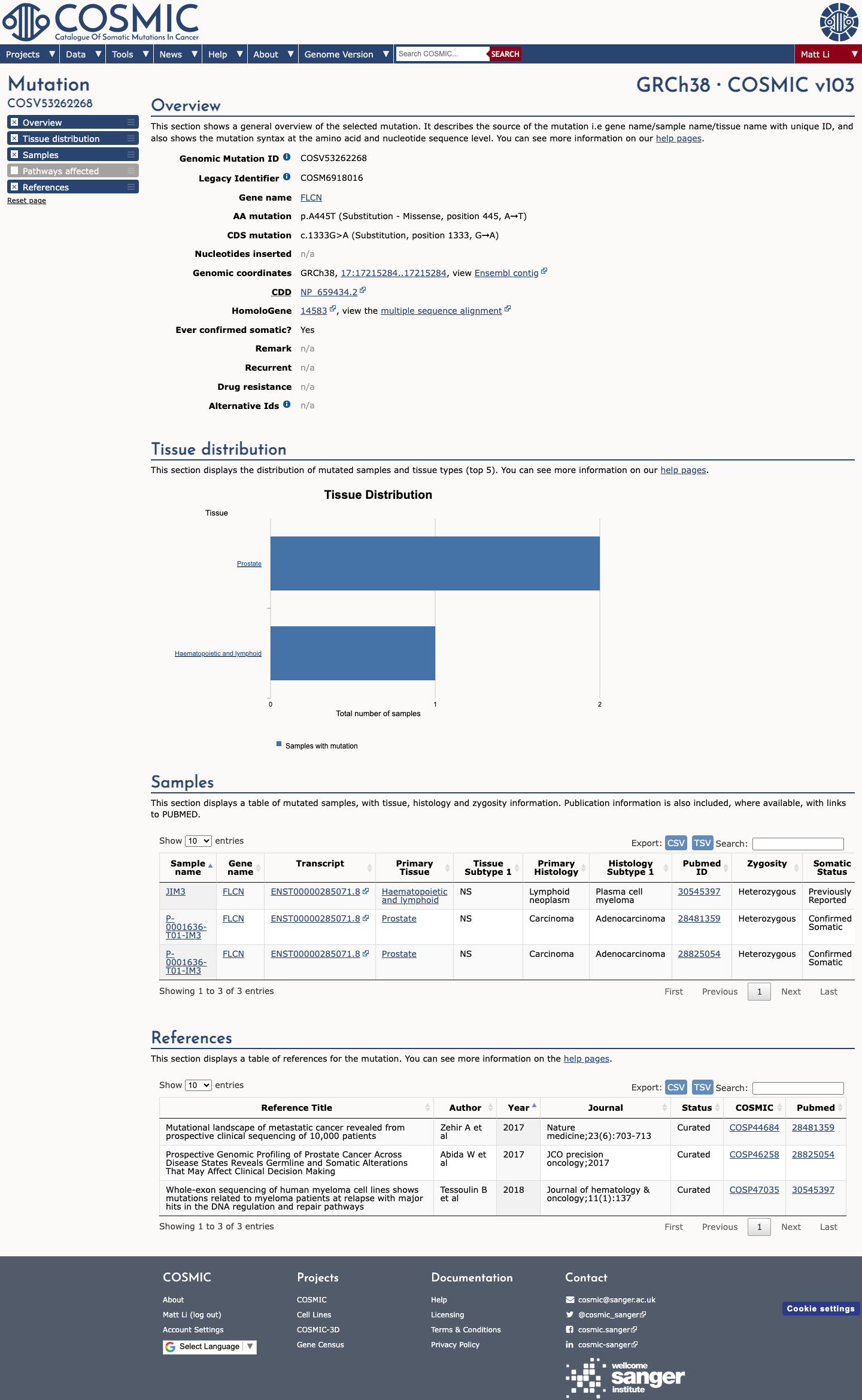
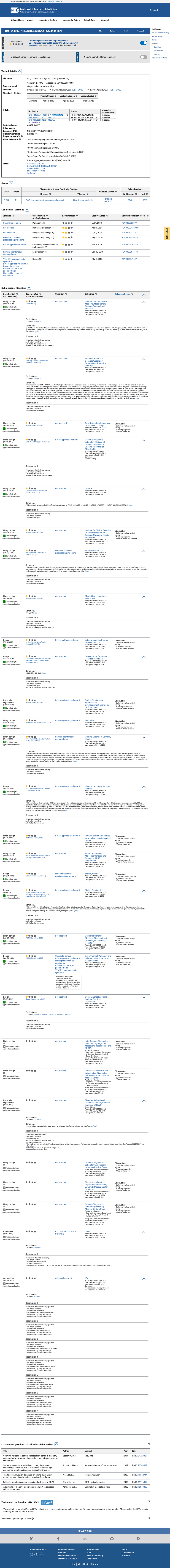
The FLCN c.1333G>A (p.Ala445Thr) variant has been reported in ClinVar predominantly as likely benign or benign, with only a single uncertain significance submission.
clinvar ↗This variant is common in population databases, including gnomAD v2.1 with a total allele frequency of 0.25848% and a highest observed subpopulation frequency of 0.37088%, and gnomAD v4.1 with a total allele frequency of 0.29305% and a Middle Eastern population frequency of 1.35269%, which exceeds the usual benign stand-alone threshold of 1%.
gnomad_v2 ↗ gnomad_v4 ↗FLCN-related disease is primarily associated with loss-of-function variants, and the reviewed FLCN mutation database reported only 2 missense variants among 53 unique germline mutations, which supports a benign interpretation for this missense change.
PMID:19562744 ↗SpliceAI predicts no significant splice effect for this variant, with a maximum delta score of 0.00.
spliceai ↗