Classification rationale
1
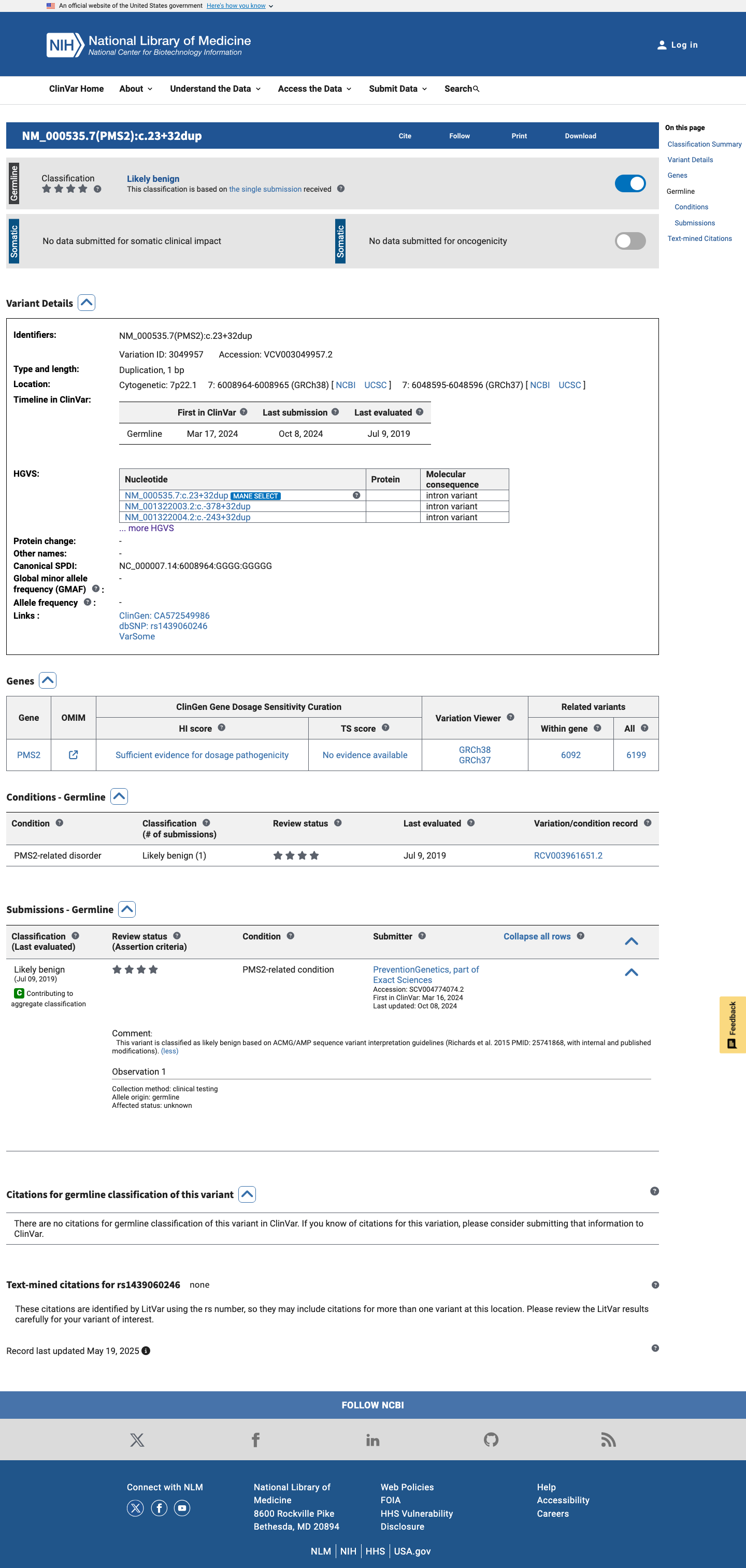
The PMS2 NM_000535.7:c.23+32dup (NP_000526.2:p.?) variant has been reported in ClinVar as likely benign by one clinical laboratory.
clinvar ↗2
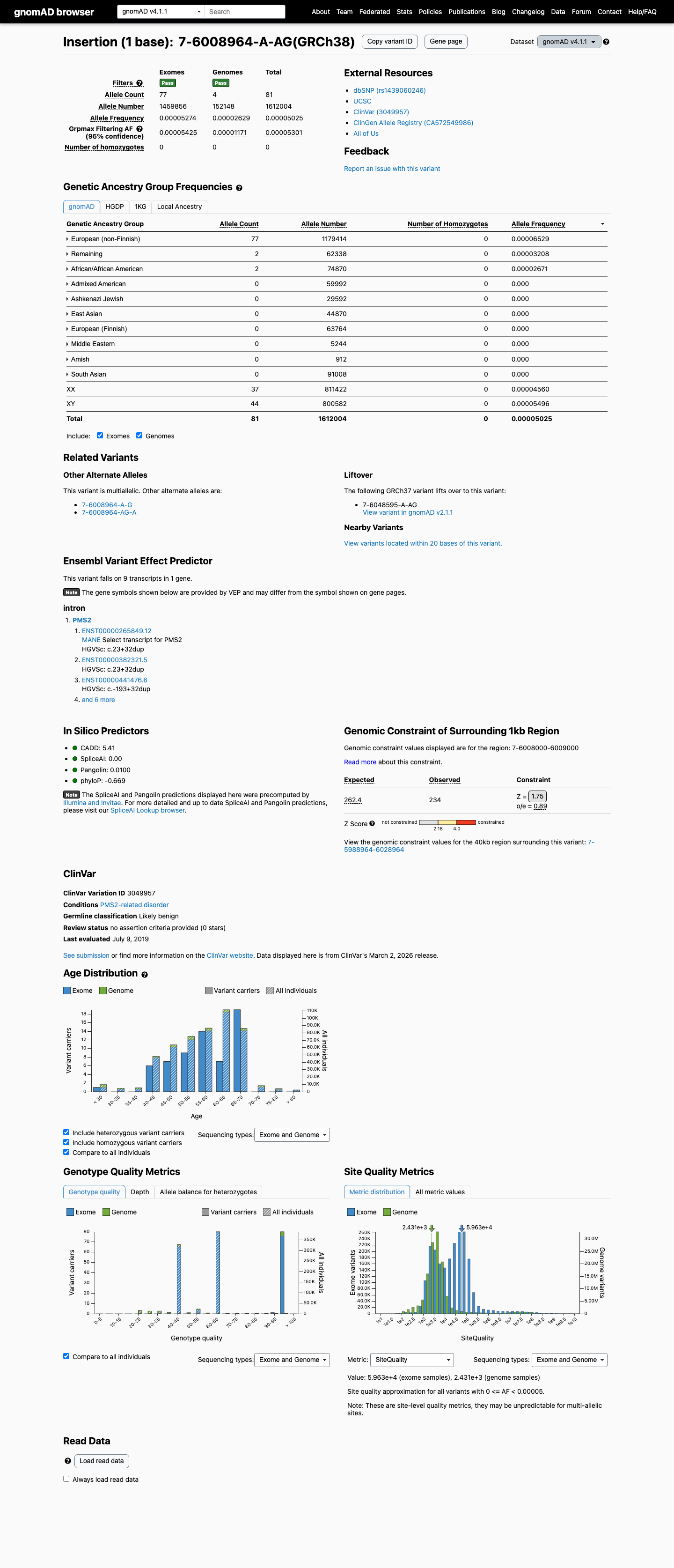
In gnomAD, this variant has a v4.1 total allele frequency of 5.0248e-05 (81/1612004 alleles) and a grpmax filtering allele frequency of 5.301e-05, which is above the PMS2 PM2 threshold of 0.00002 but below the BS1 threshold of 0.00028.
gnomad_v4 ↗ cspec ↗3
This intronic duplication is located at c.23+32, beyond the PMS2 BP7 boundary of +7, and SpliceAI predicts no significant splice impact with a maximum delta score of 0.02, supporting BP7 and BP4 while arguing against PP3 and PVS1.
cspec ↗ spliceai ↗