Classification rationale
1
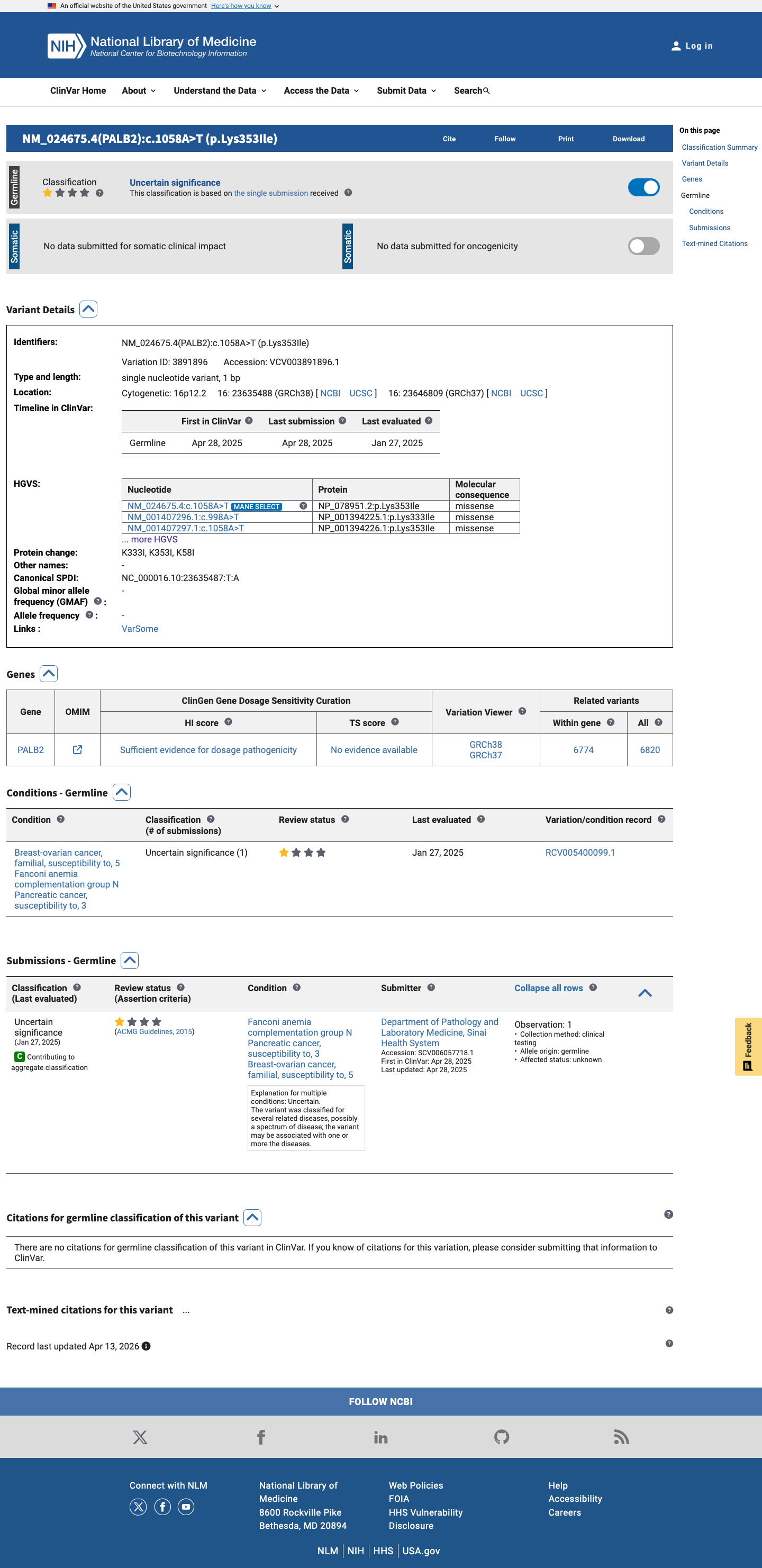
The PALB2 c.1058A>T (p.Lys353Ile) variant has been reported in ClinVar as uncertain significance by a single submitter.
clinvar ↗2
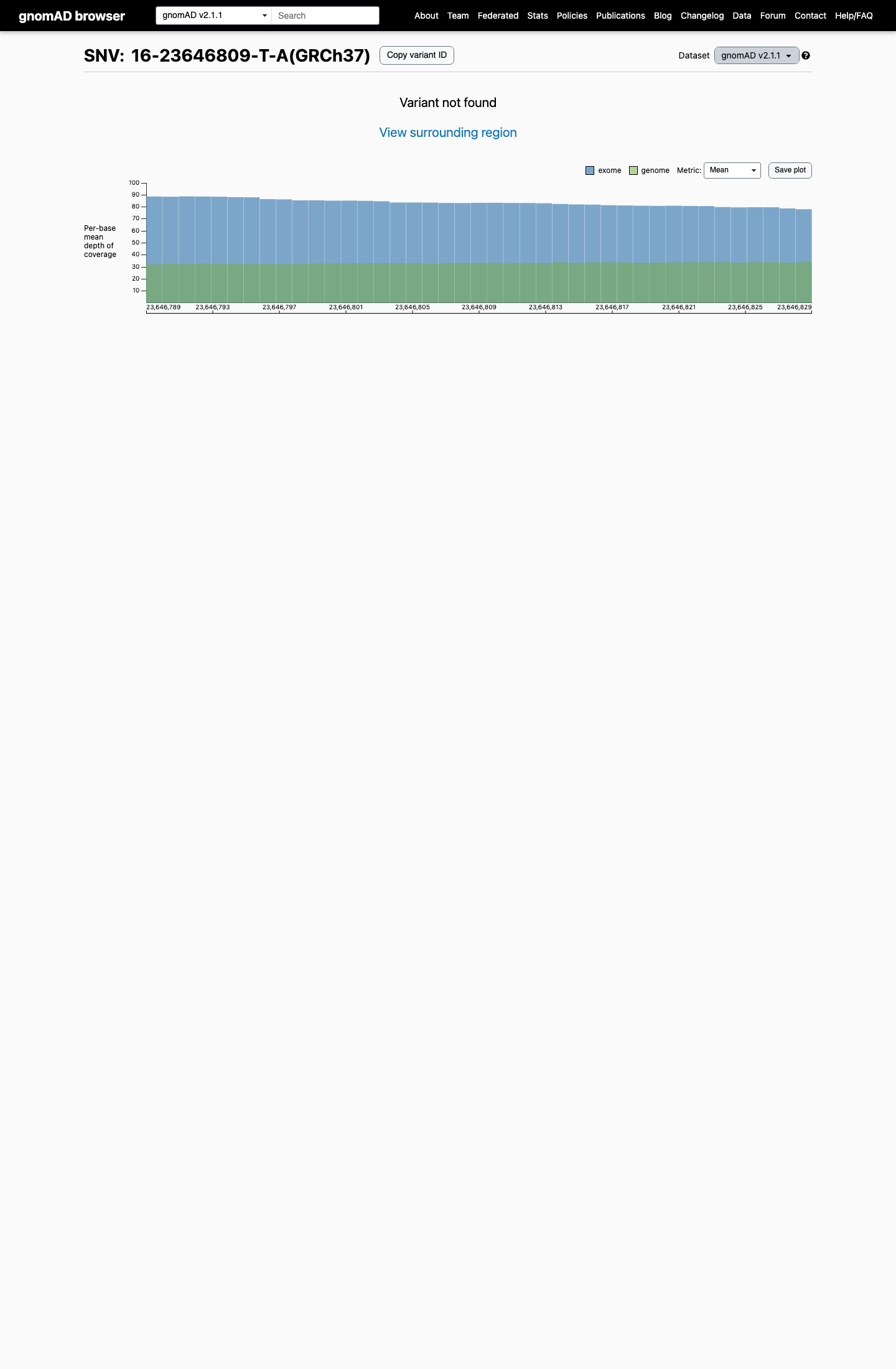
This variant is absent from gnomAD v2.1 and gnomAD v4.1, and its gnomAD v4.1 allele frequency is therefore below the PALB2 PM2_Supporting threshold of 0.000333%.
cspec ↗ gnomad_v2 ↗ gnomad_v4 ↗3
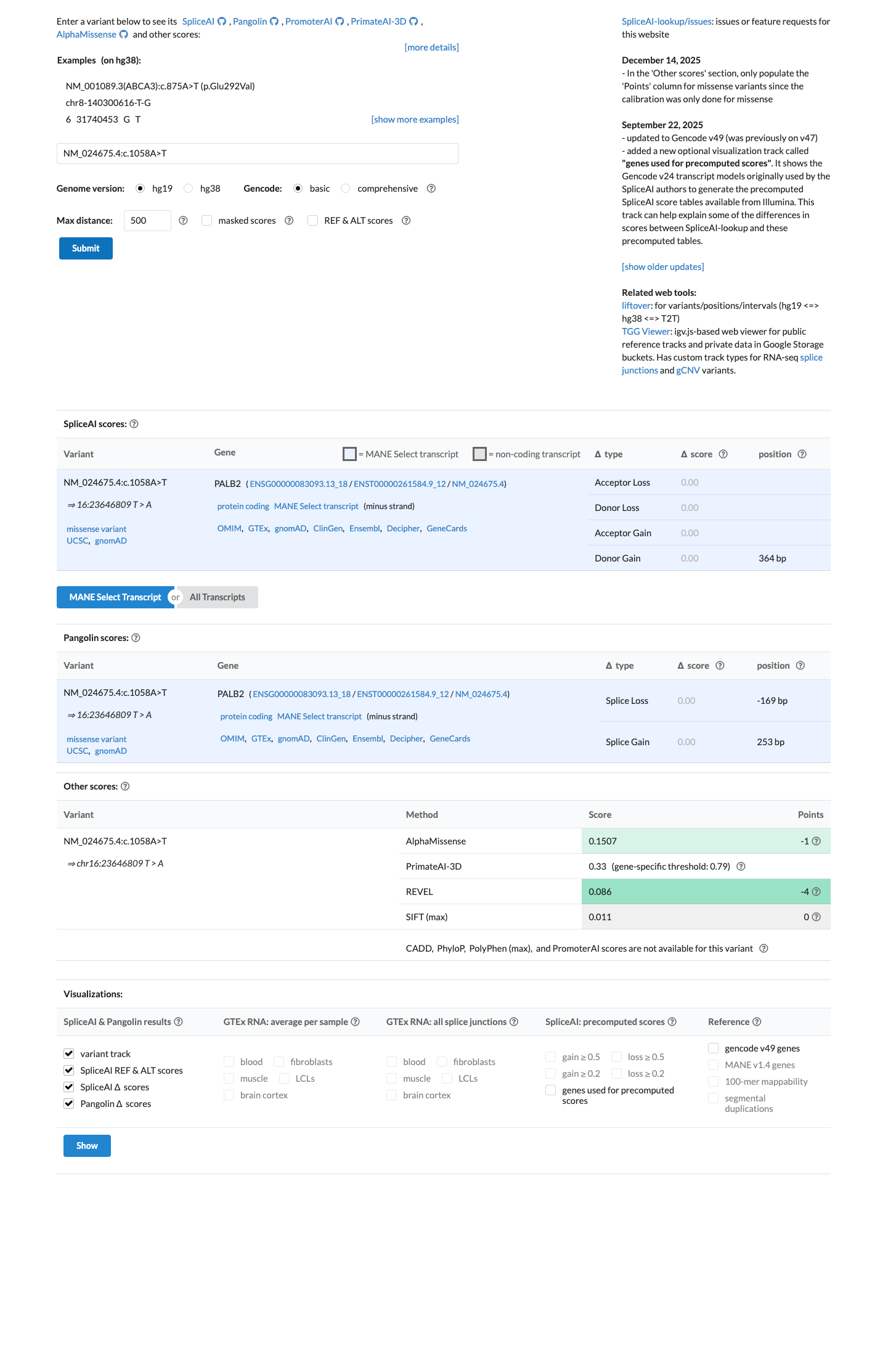
PALB2-specific rules support BP1 for this missense variant, while SpliceAI predicts no significant splice impact with a maximum delta score of 0.00 and PVS1, PP3, and BP4 are not applied in this missense setting.
cspec ↗ spliceai ↗