Classification rationale
1
The LZTR1 c.2412dup (p.Lys805GlnfsTer46; p.K805Qfs*46) variant has not been observed in COSMIC and has been reported in ClinVar with likely pathogenic and pathogenic submissions.
clinvar ↗2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity in population databases and meeting the LZTR1 PM2_Supporting threshold of ≤0.0025%.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
OncoKB classifies this variant as likely oncogenic with a likely loss-of-function biological effect, but no approved RASopathy VCEP functional assay result was identified for this specific variant.
oncokb ↗4
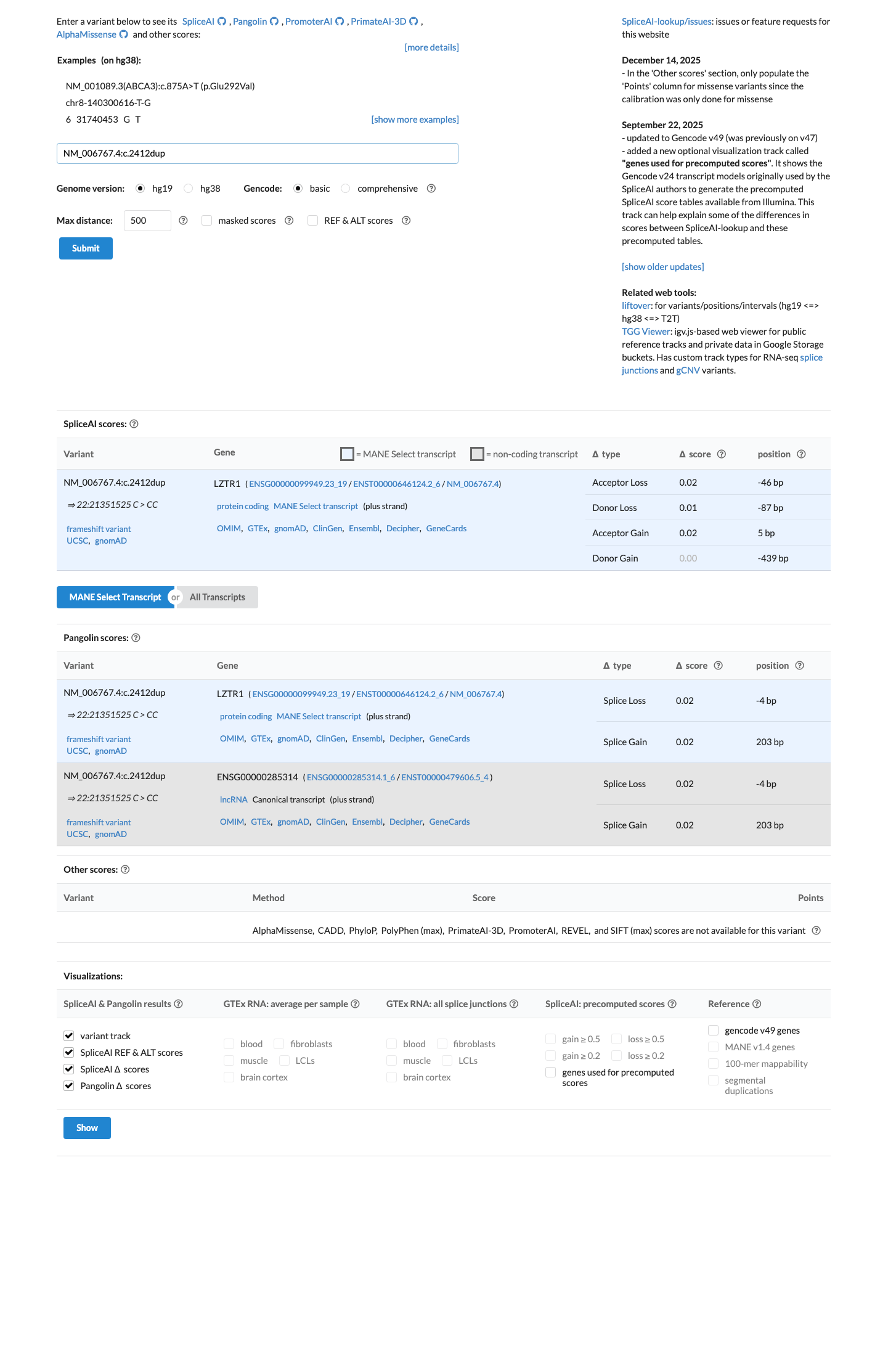
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.02, and no REVEL or BayesDel score was available because this is not a single-nucleotide variant.
spliceai ↗