Classification rationale
1
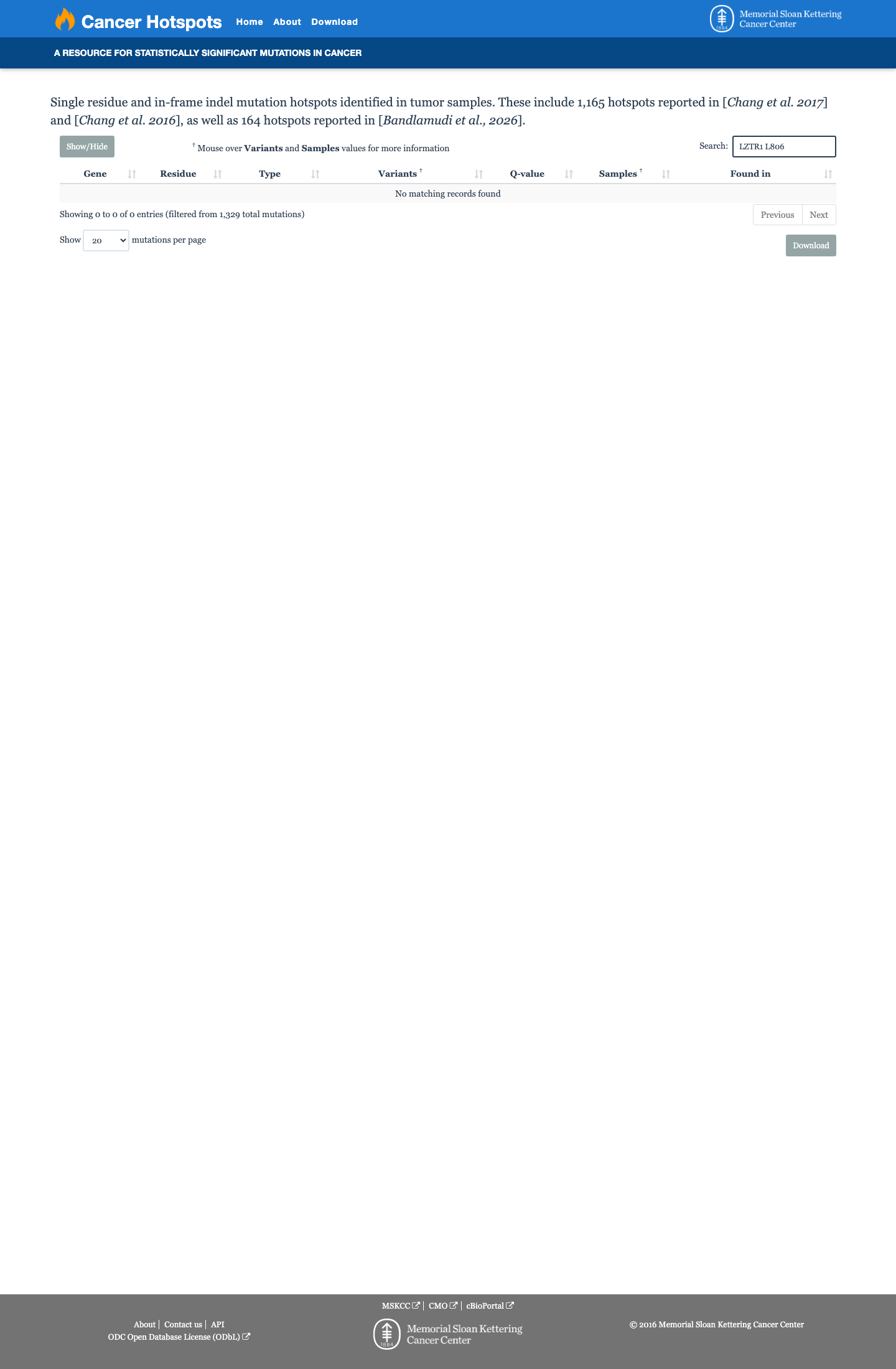
The LZTR1 c.2417T>G (p.Leu806Trp) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar as likely pathogenic by a single clinical laboratory.
clinvar ↗2
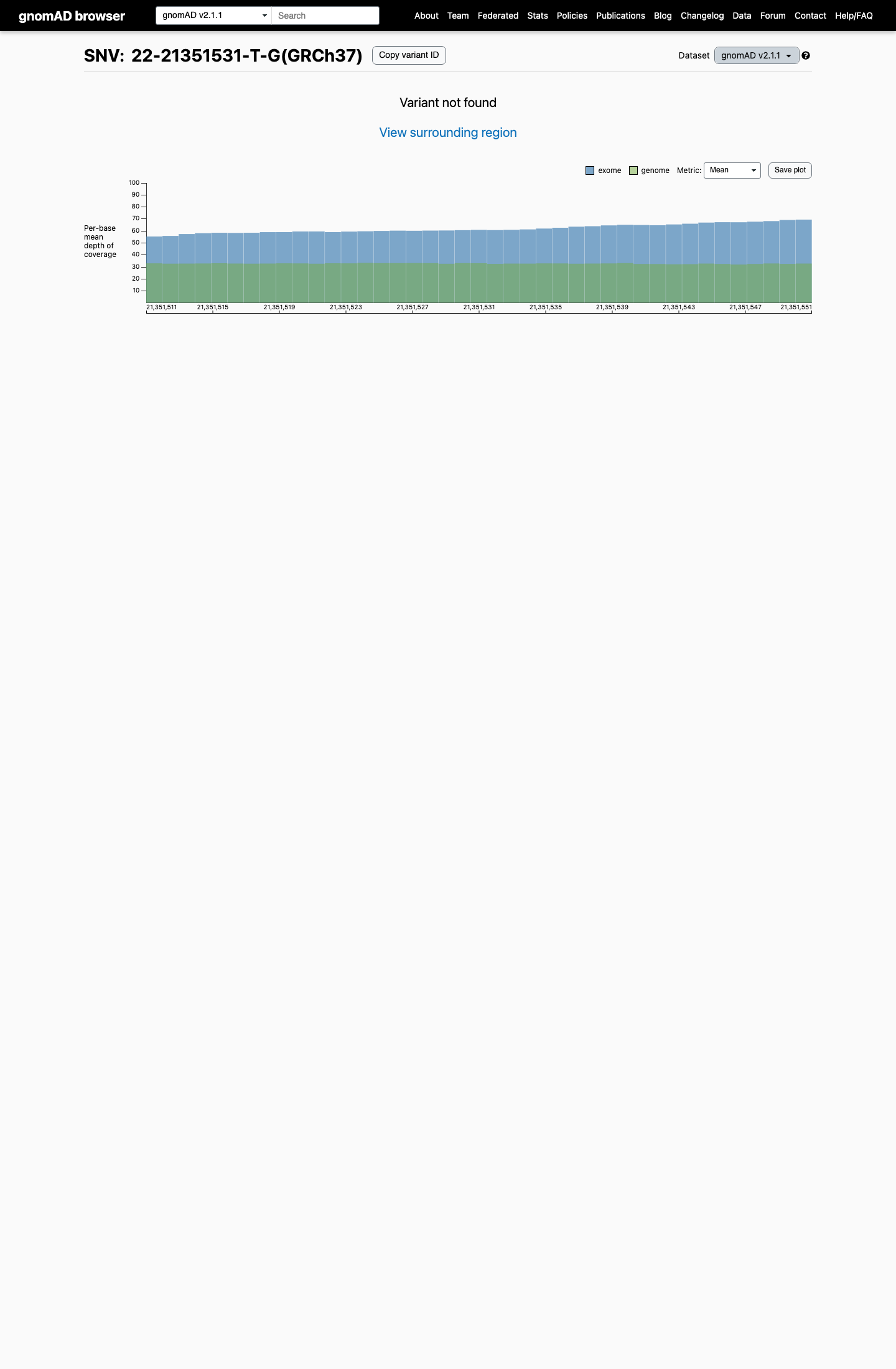
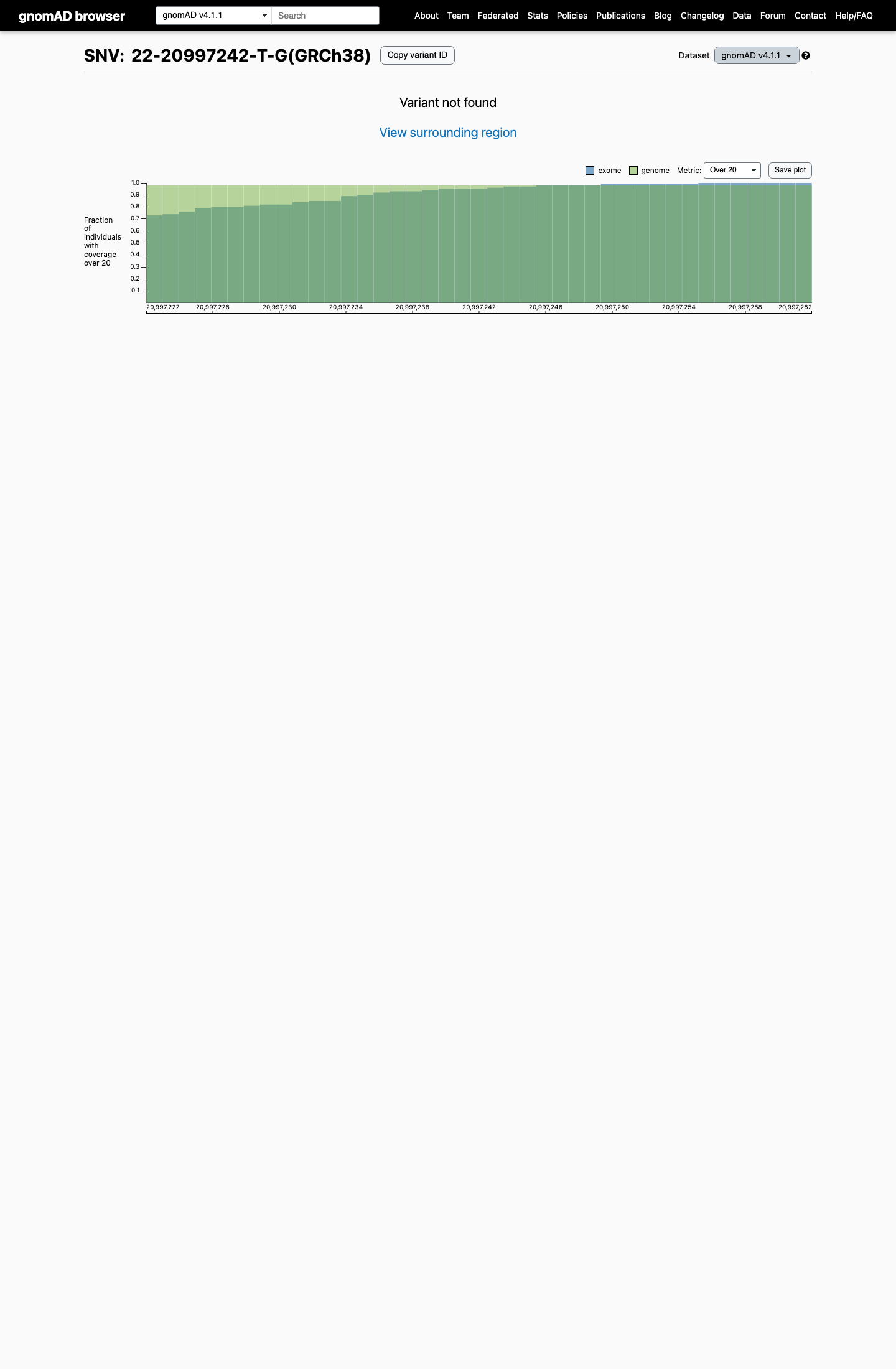
This variant is absent from gnomAD v2.1 and gnomAD v4.1, corresponding to an observed allele frequency of 0 in both datasets.
gnomad_v2 ↗ gnomad_v4 ↗3
Computational evidence supports a benign interpretation because REVEL is 0.174, below the LZTR1 BP4 threshold of 0.3, SpliceAI predicts no splice effect with a maximum delta score of 0.00, and BayesDel is -0.202983.
cspec ↗ spliceai ↗