Classification rationale
1
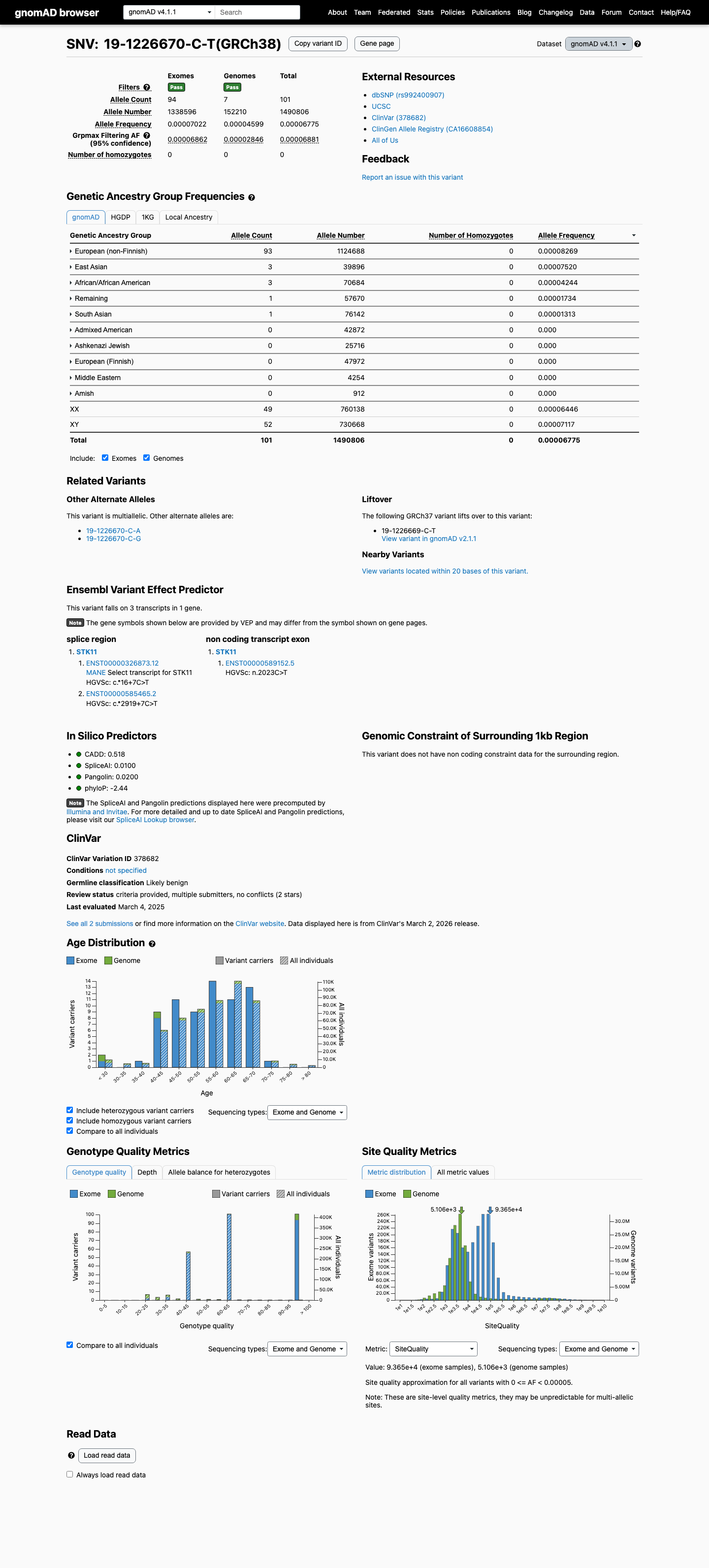
The STK11 c.*16+7C>T (p.?) variant has been reported in ClinVar as likely benign by two clinical laboratories, and no expert-panel review was identified.
clinvar ↗2
This variant is present at low frequency in gnomAD, with AF 0.00639% in v2.1 and AF 0.00677% in v4.1; these values remain below the 0.1% PM2 threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
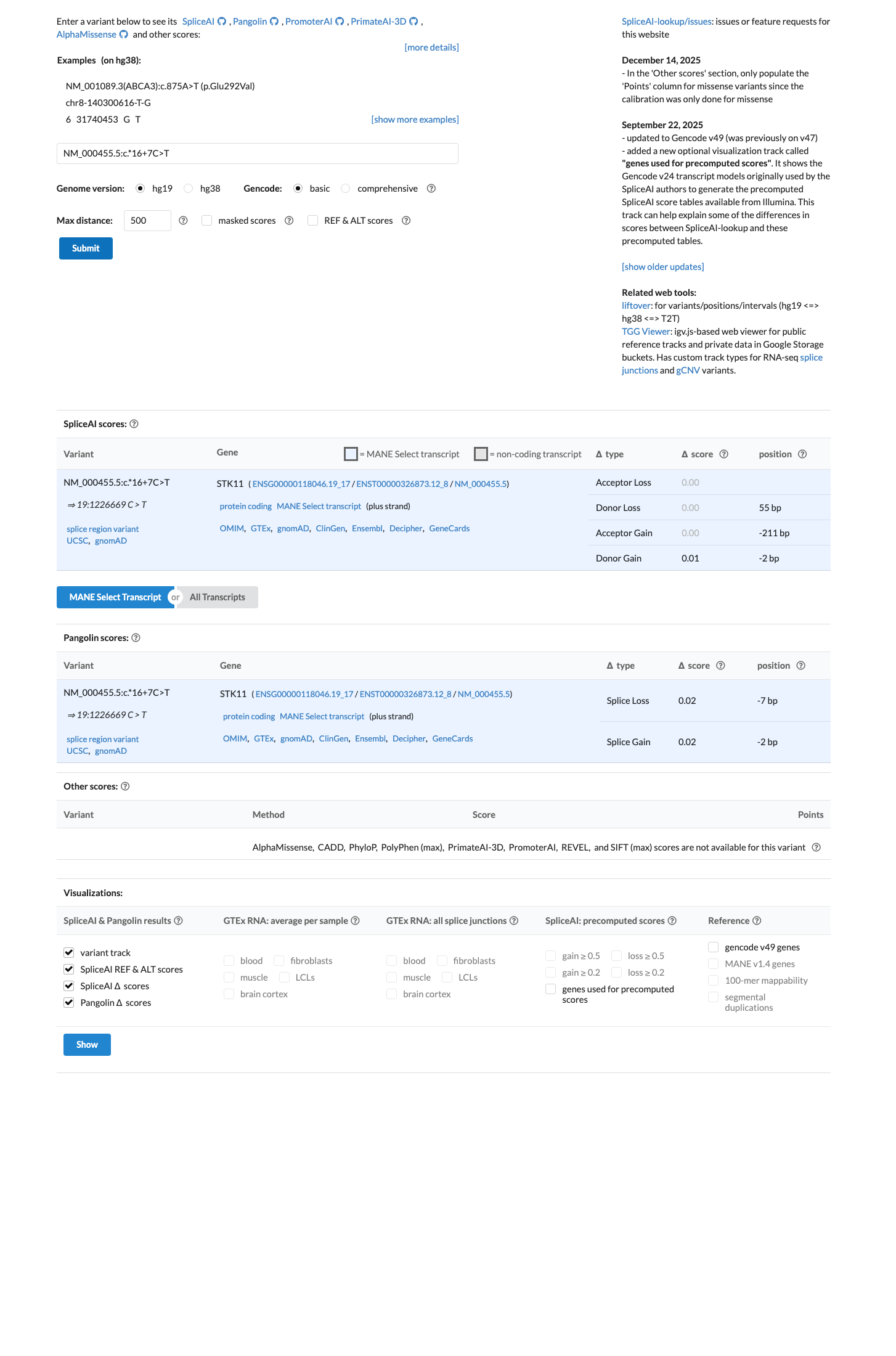
In silico splice prediction does not support a damaging effect, with SpliceAI showing a maximum delta score of 0.01, consistent with no significant splice impact.
spliceai ↗