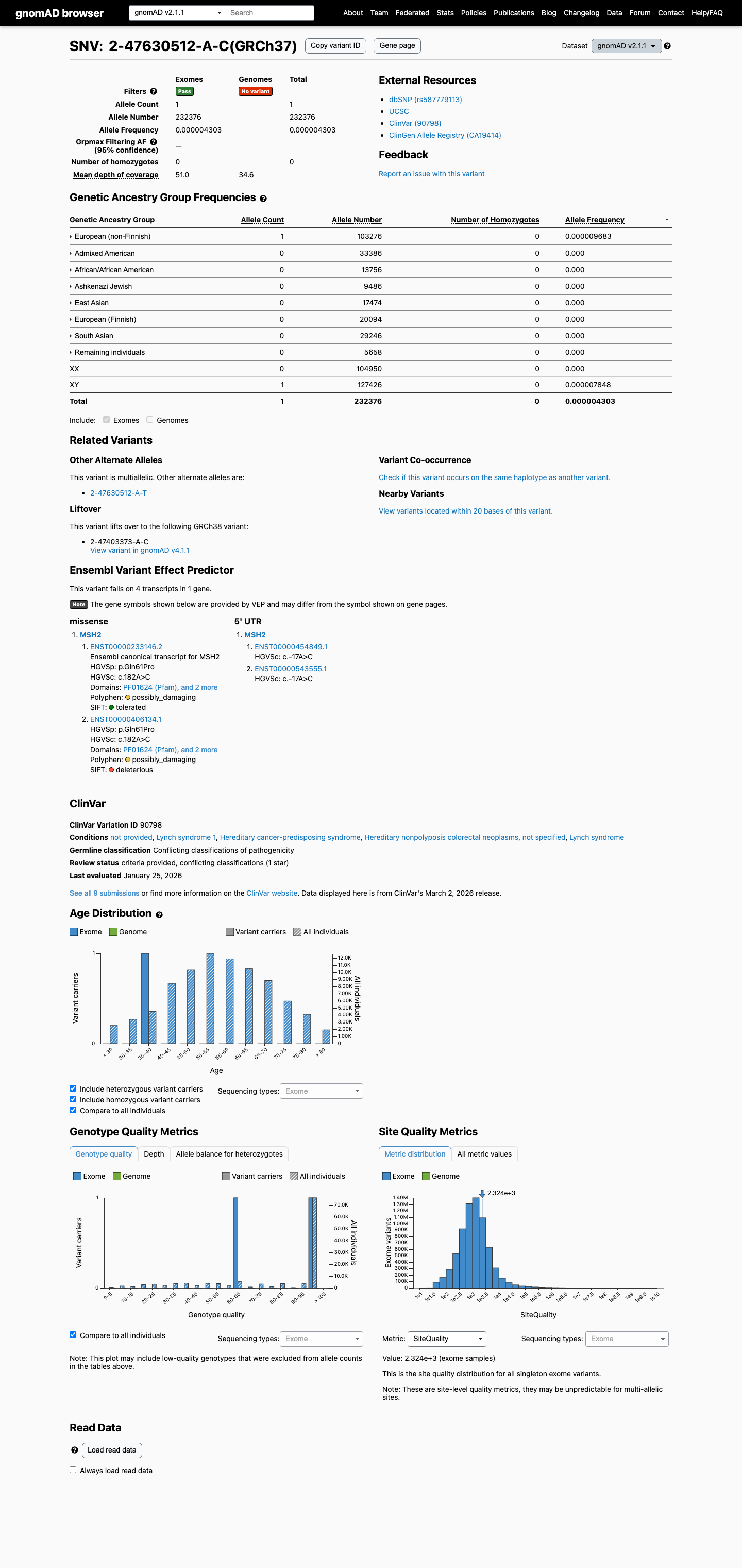
The MSH2 c.182A>C (p.Gln61Pro) variant has not been observed in COSMIC and has been reported in ClinVar with conflicting interpretations, including uncertain significance and likely benign submissions.
clinvar ↗This variant is present at extremely low frequency in gnomAD v4.1 (5/1,606,956 alleles; AF 3.11e-06; grpmax FAF 1.24e-06), which is below the InSiGHT MSH2 PM2_Supporting threshold of 0.00002.
gnomad_v4 ↗ cspec ↗Published MSH2 functional studies are cited for this variant class, but no verified variant-specific assay result or calibrated functional evidence was identified for p.(Gln61Pro), so PS3 and BS3 are not supported at this time.
PMID:17720936 ↗ PMID:20176959 ↗ PMID:33357406 ↗In silico data show no meaningful predicted splice effect by SpliceAI (max delta 0.01), while missense predictors show a damage signal (REVEL 0.72; BayesDel 0.0679979); however, the MSH2 PP3/BP4 framework is based on HCI prior probabilities, which were not identified here.
spliceai ↗ cspec ↗