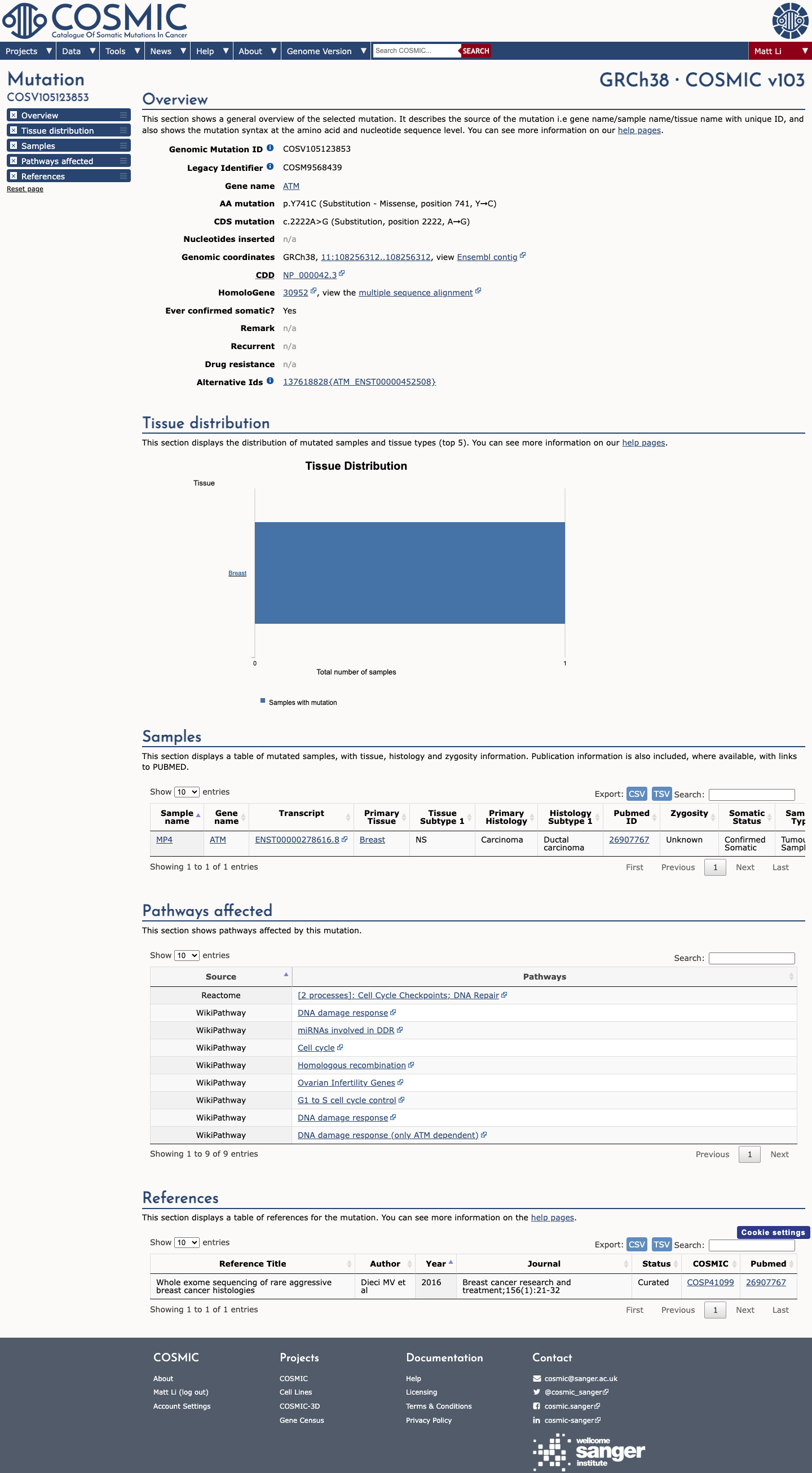
The ATM c.2222A>G (p.Tyr741Cys) variant has been observed once in somatic cancer in COSMIC and has been reported in ClinVar with conflicting germline classifications, including uncertain significance and likely benign submissions.
clinvar ↗This variant is present in population databases at low frequency, including gnomAD v4.1 at 0.00217% overall (35/1,611,122 alleles) with a highest observed subpopulation frequency of 0.00447% in East Asian individuals, which is above the ATM PM2_Supporting threshold of <=0.001% and below the BS1 and BA1 thresholds.
gnomad_v4 ↗ gnomad_v2 ↗ cspec ↗Computational evidence is mixed: SpliceAI predicts possible splice impact with a maximum delta score of 0.29, supporting ATM PP3_Supporting, whereas REVEL is low at 0.153 and BayesDel is negative at -0.288378, which argues against a damaging missense effect but does not resolve the predicted splice concern.
spliceai ↗ cspec ↗