The MAP2K2 c.383C>A (p.Pro128Gln, p.P128Q) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar as Pathogenic, including review by the ClinGen RASopathy Variant Curation Expert Panel.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity in population reference datasets.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In a published functional study, p.Pro128Gln increased downstream ERK phosphorylation relative to wild type, consistent with an activating effect, and RASopathy VCEP-approved functional assay tables include this variant among pathogenic validation controls for MEK and ERK activation assays.
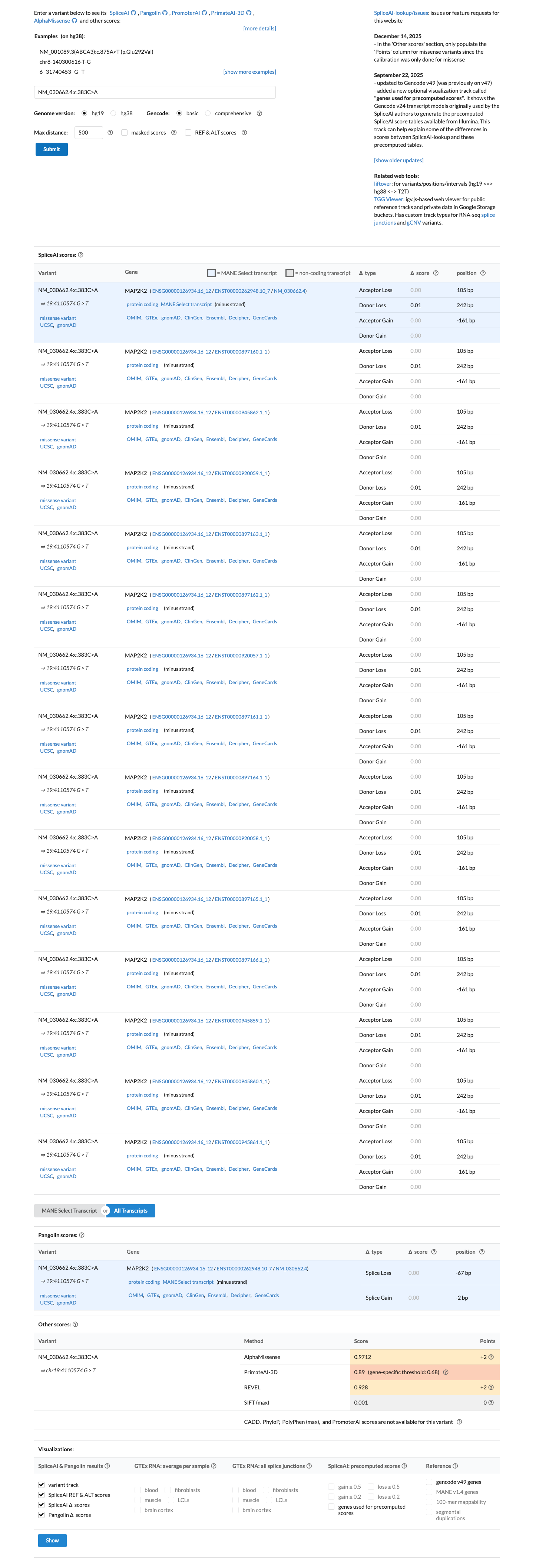
PMID:20358587 ↗Computational evidence supports a damaging effect, with REVEL 0.928 above the MAP2K2 RASopathy PP3 threshold of 0.7 and BayesDel 0.495827 in the damaging direction, while SpliceAI predicts no meaningful splice alteration with a maximum delta score of 0.01.
spliceai ↗ cspec ↗