Classification rationale
1
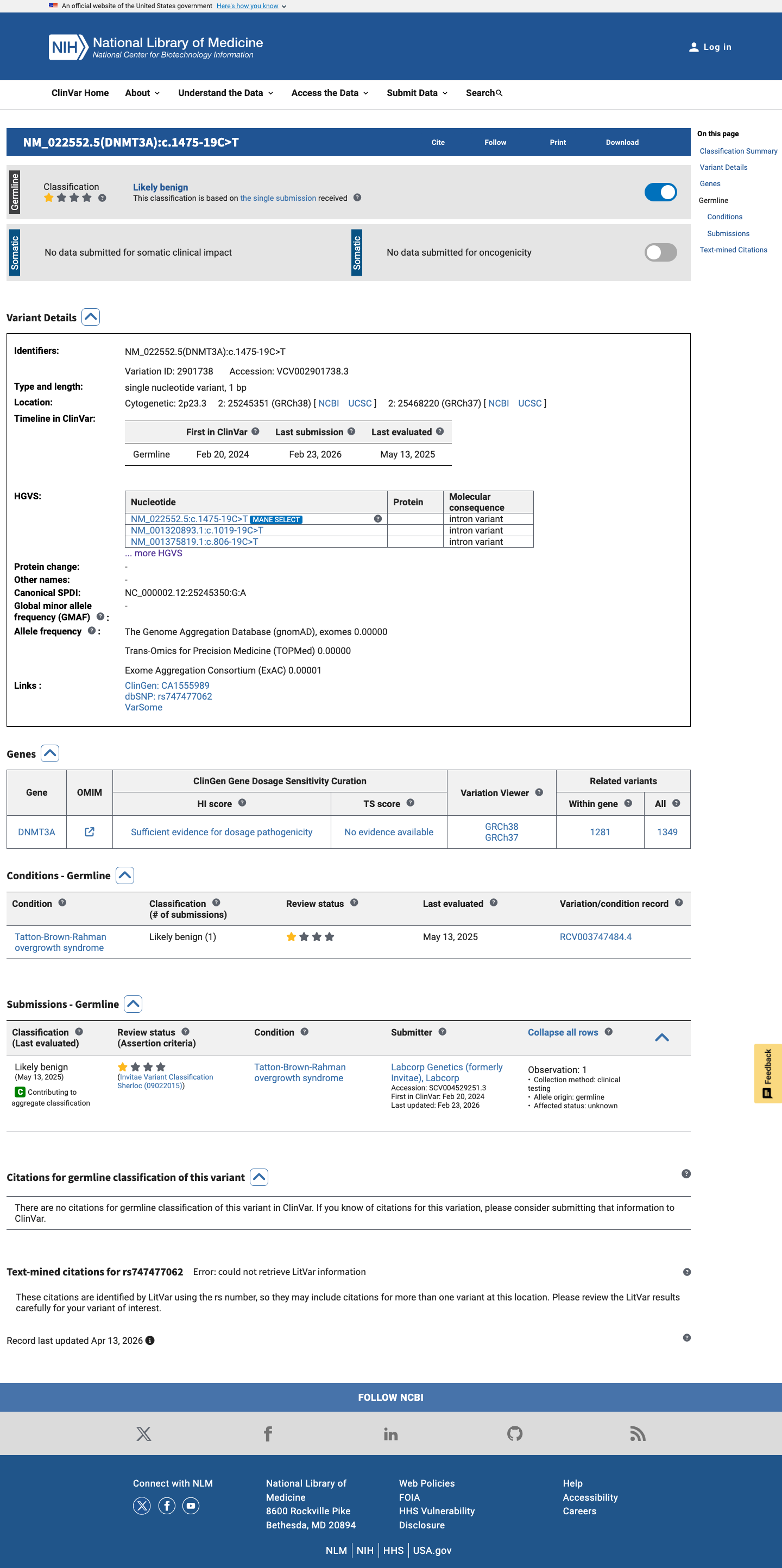
The DNMT3A c.1475-19C>T (p.?) variant has been reported in ClinVar with a Likely benign classification from a single clinical laboratory.
clinvar ↗2
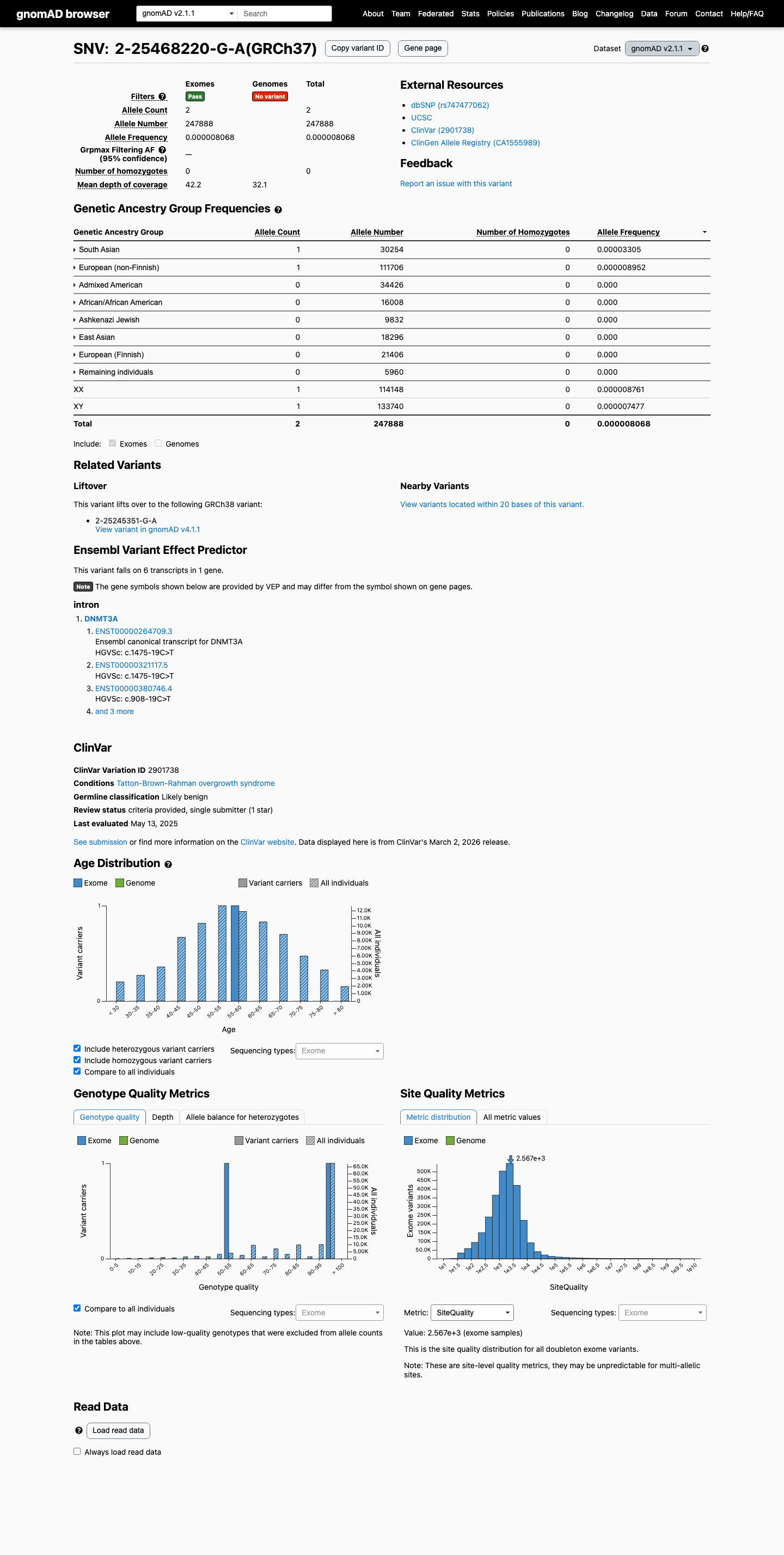
This variant is present at very low overall frequency in population databases, with gnomAD v2.1 AF 0.00081% (2/247888 alleles) and gnomAD v4.1 AF 0.00043% (7/1611438 alleles); the highest observed frequency is 0.04936% in the Middle Eastern population in gnomAD v4.1, which is below the default BS1 threshold of 0.3% and below the BA1 threshold of 1.0%.
gnomad_v2 ↗ gnomad_v4 ↗3
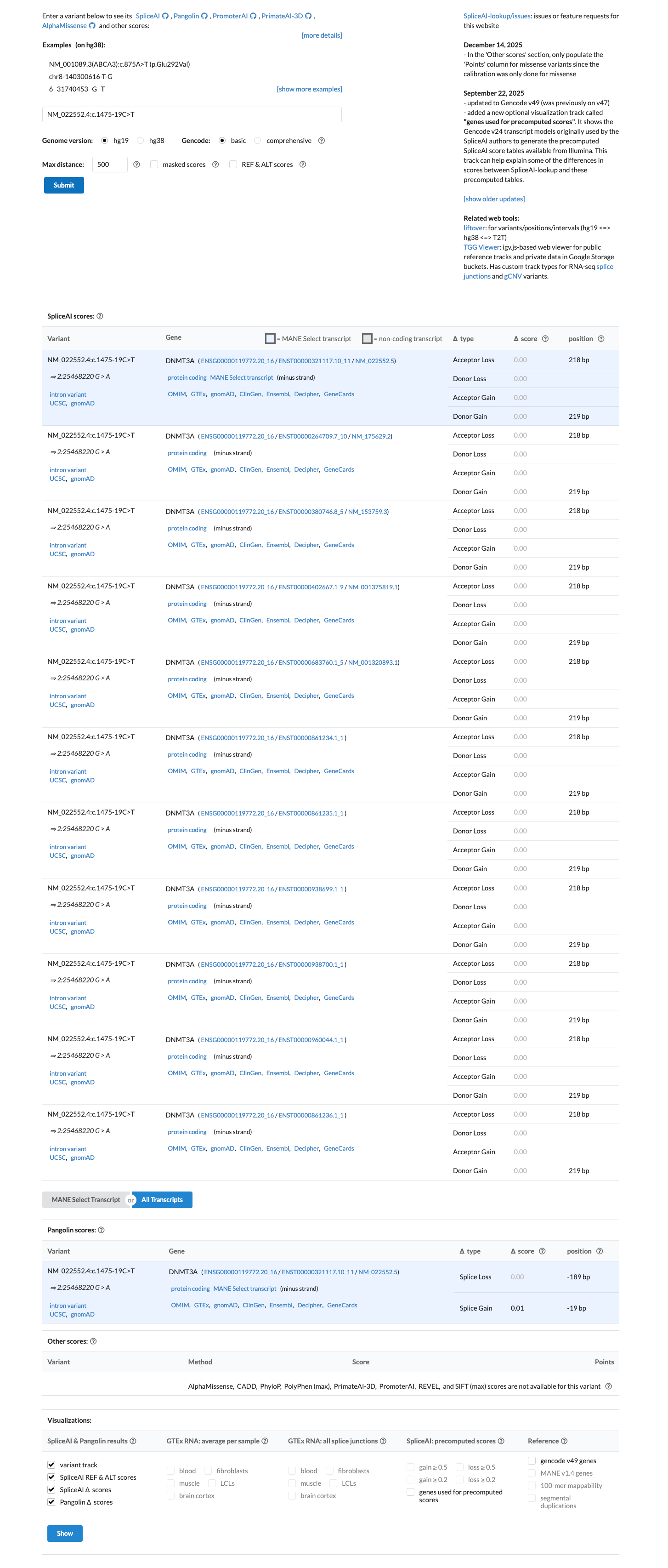
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.00, which supports a benign computational interpretation rather than evidence for abnormal splicing.
spliceai ↗