Classification rationale
1
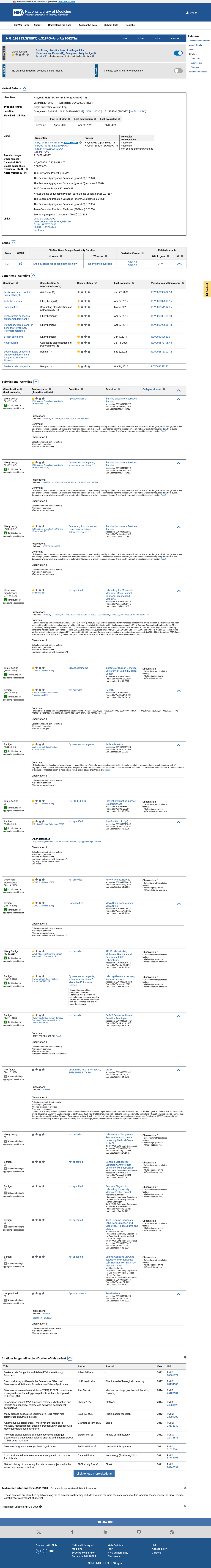
The TERT NM_198253.2:c.3184G>A (p.Ala1062Thr) variant has been reported in ClinVar, where most submissions classify it as benign or likely benign.
clinvar ↗2
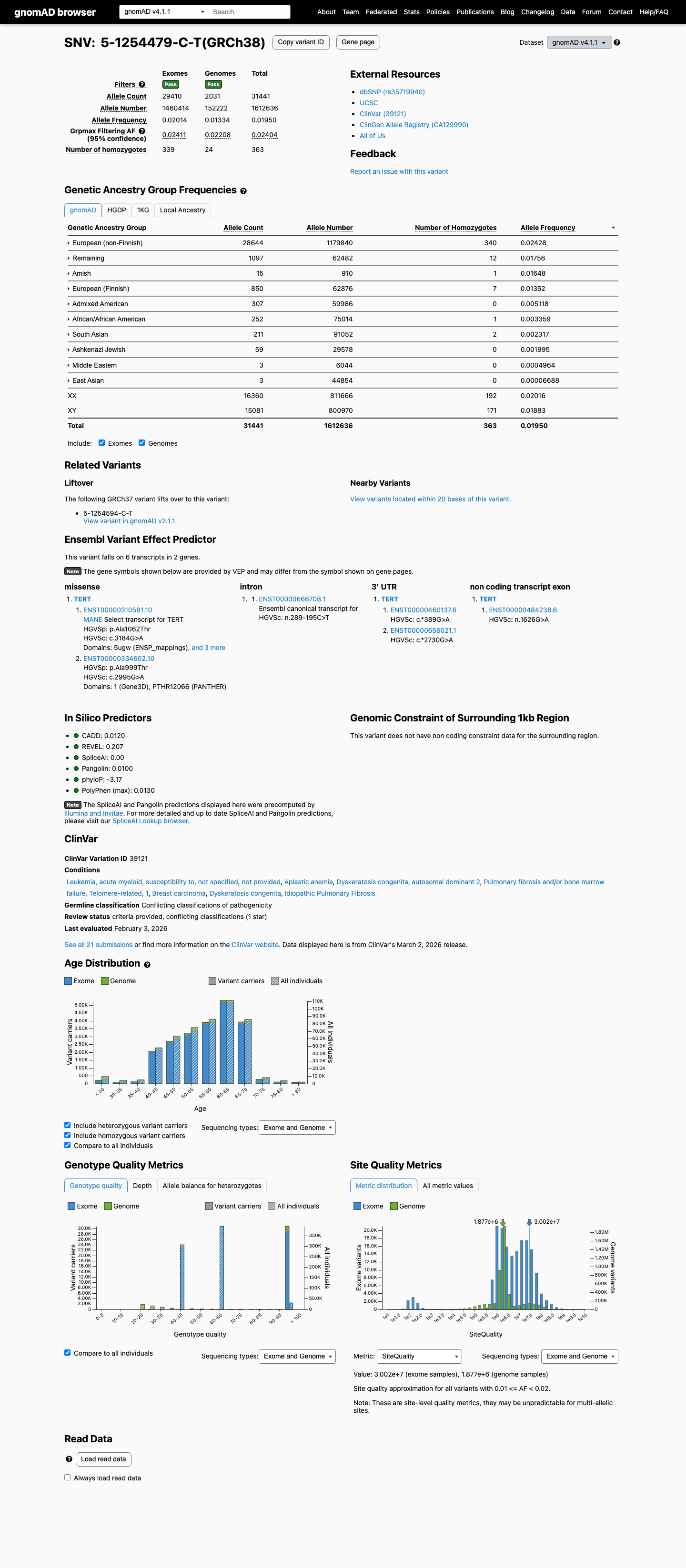
This variant is common in population databases, with allele frequencies of 1.24601% in gnomAD v2.1 and 1.94967% in gnomAD v4.1, which are above the benign stand-alone threshold of 1.0% and above the BS1 threshold of 0.3%.
gnomad_v2 ↗ gnomad_v4 ↗3
In silico prediction does not support a damaging effect, with REVEL 0.207, BayesDel -0.373519, and SpliceAI maximum delta score 0.01 predicting no significant splice impact.
spliceai ↗