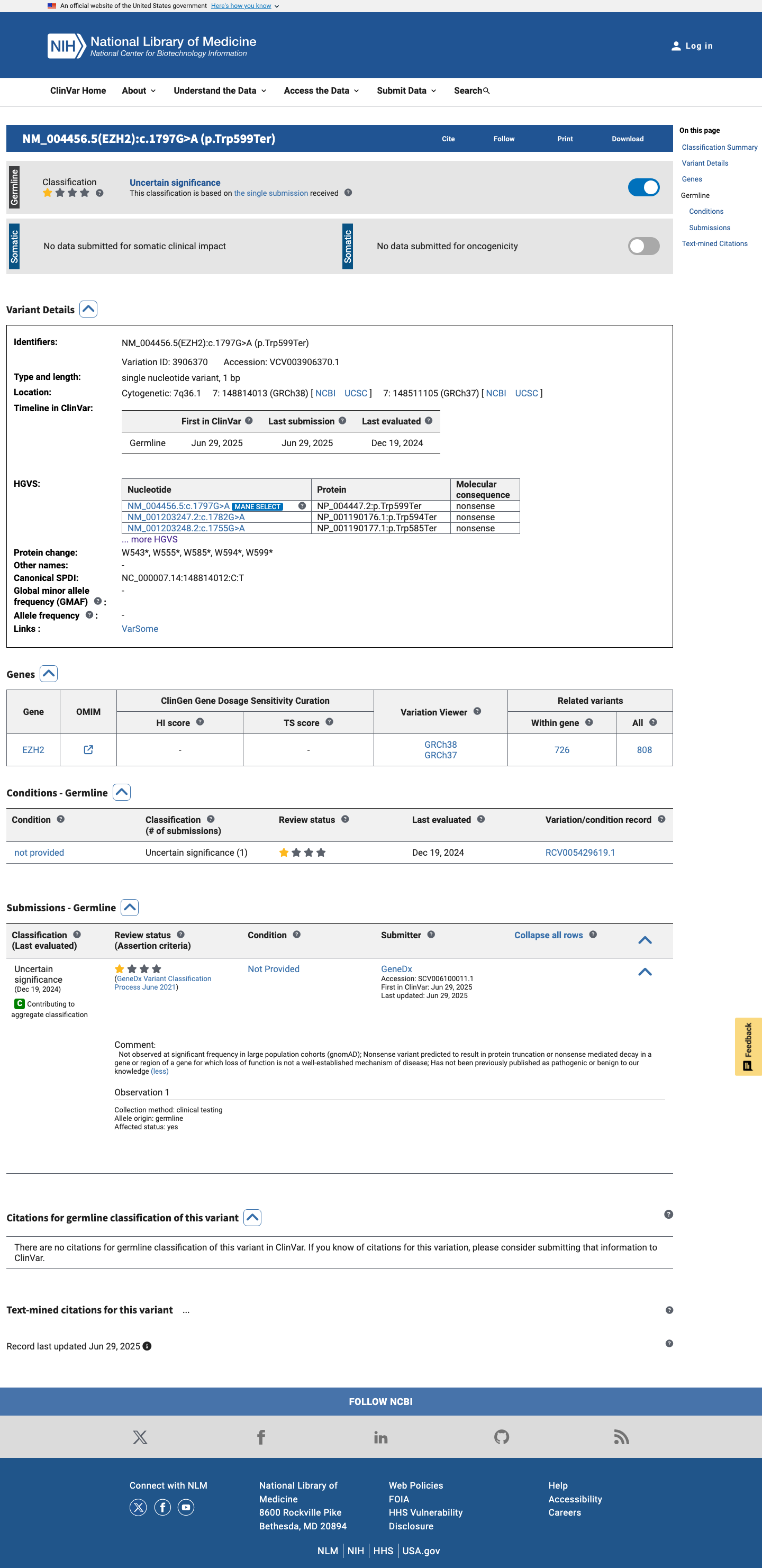
The EZH2 c.1797G>A (p.Trp599Ter; p.W599*) variant has been reported in ClinVar as a variant of uncertain significance with one clinical laboratory submission.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, which is below the usual PM2 population threshold of 0.1% for a rare germline disorder.
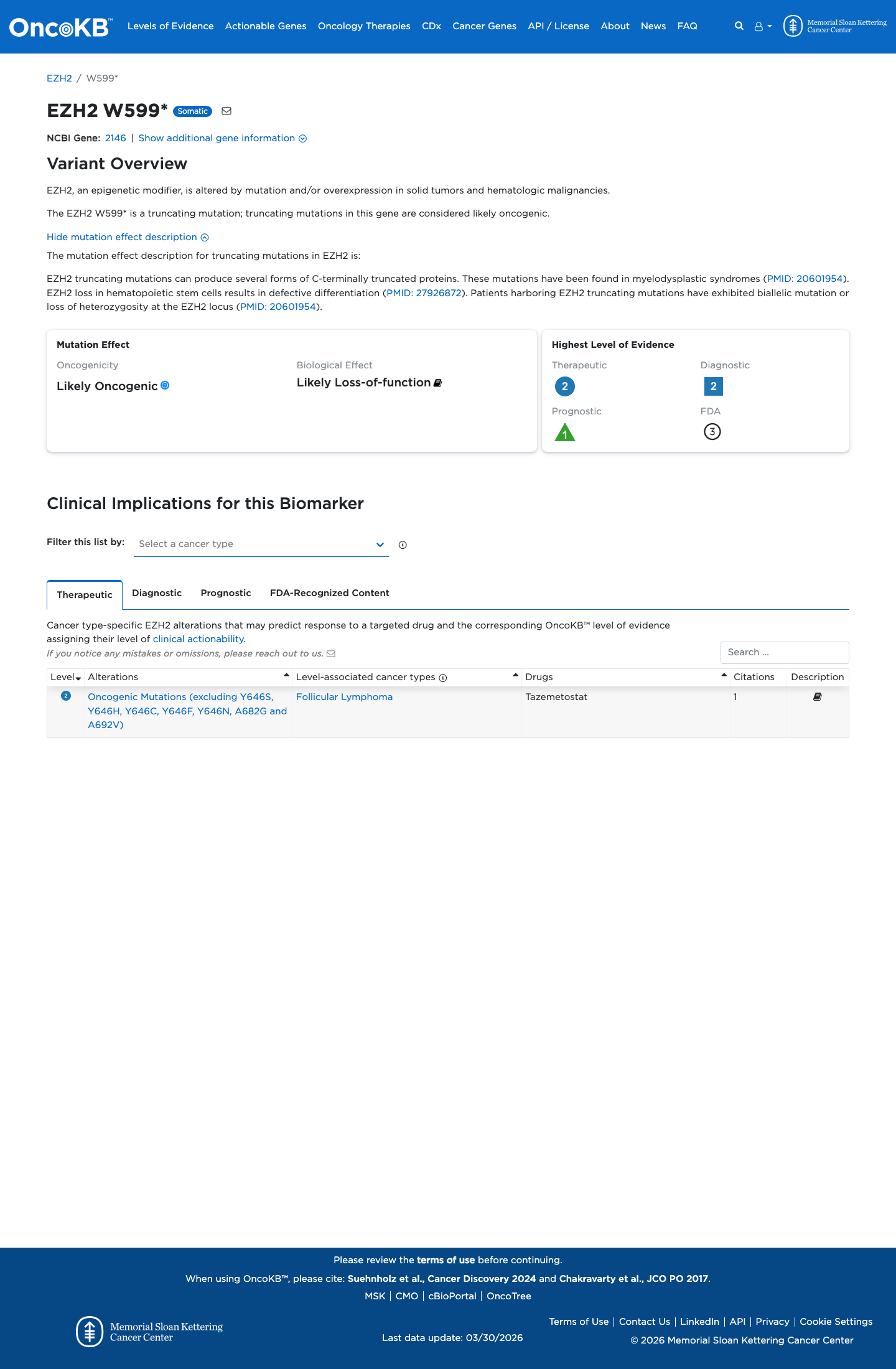
gnomad_v2 ↗ gnomad_v4 ↗This nonsense variant is predicted to introduce a premature termination codon in exon 15 with downstream coding exons remaining, and generic PVS1 review indicates that EZH2 is eligible for loss-of-function assessment under the ClinGen SVI framework.
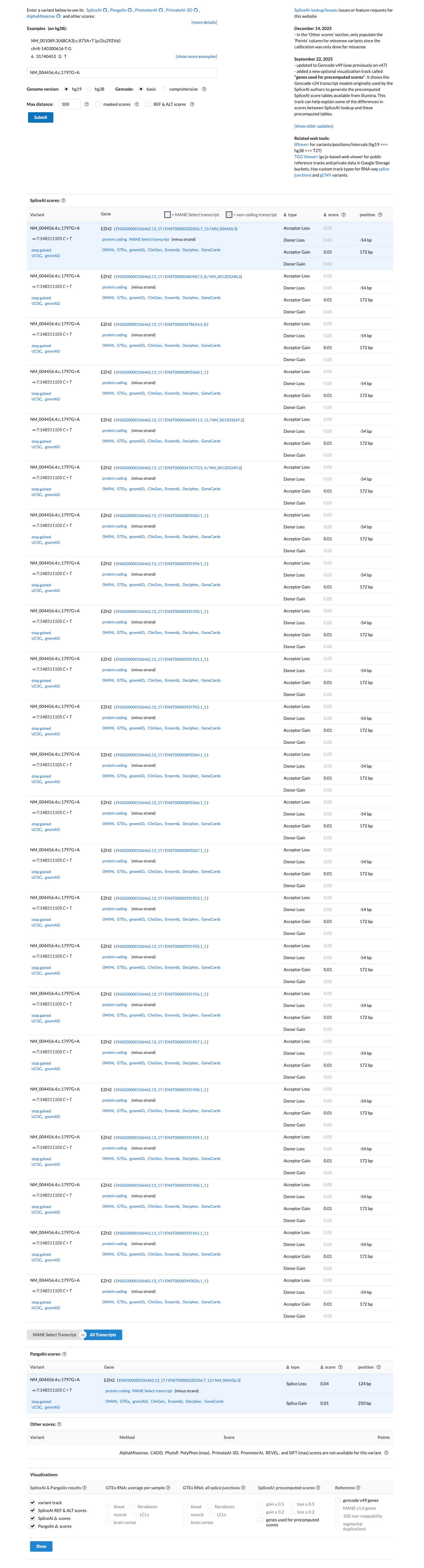
pvs1_generic_framework ↗SpliceAI predicts no significant splice impact for this variant (maximum delta score 0.01), while BayesDel is 0.655873; these computational findings do not provide standalone benign support for this truncating variant.
spliceai ↗