Classification rationale
1
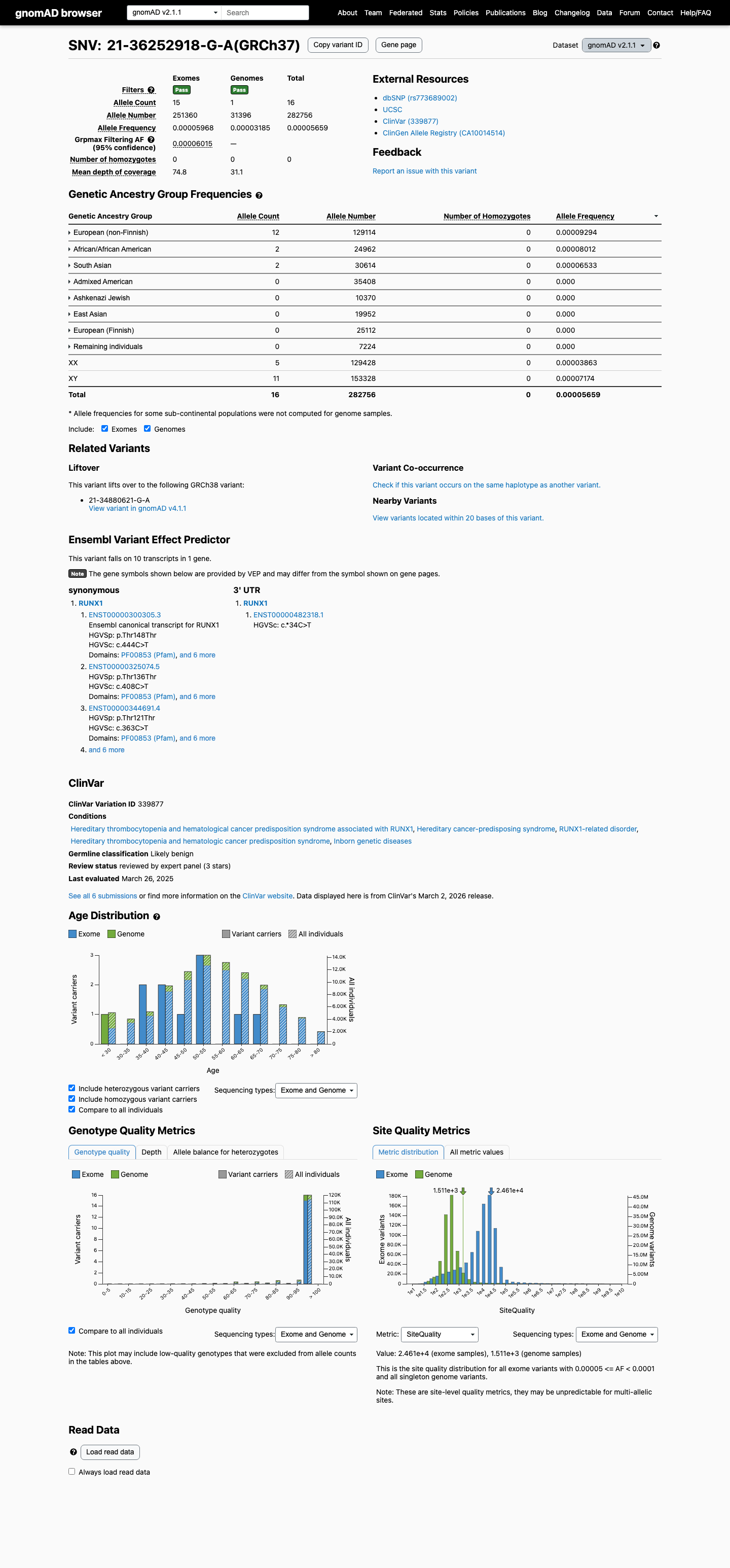
The RUNX1 c.444C>T (p.Thr148=) variant is reported in ClinVar and is currently classified as Likely Benign by the ClinGen Myeloid Malignancy Variant Curation Expert Panel.
clinvar ↗2
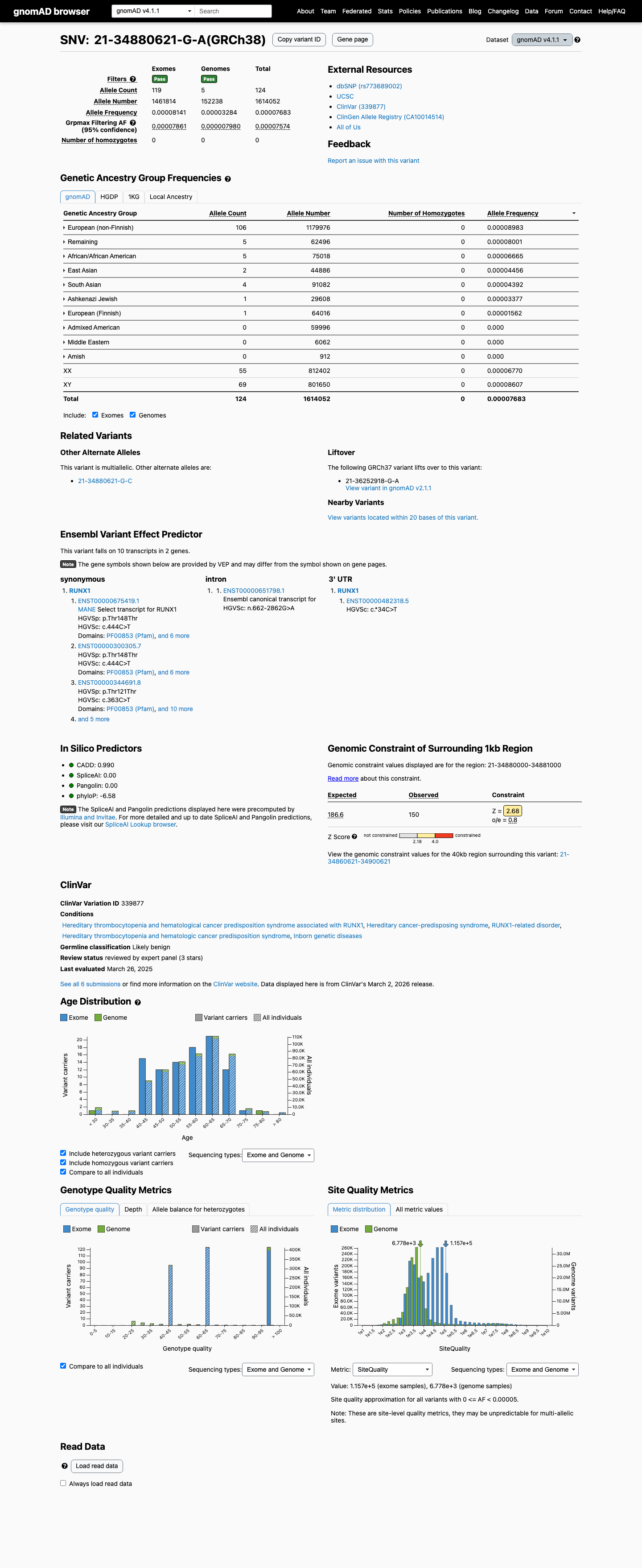
This variant is present in population databases, including gnomAD v4.1 with grpmax FAF 7.574e-05 and gnomAD v2.1 with grpmax FAF 6.015e-05; these values are above the RUNX1 PM2_Supporting threshold of 0.00005 and below the RUNX1 BS1 threshold of 0.00015.
gnomad_v4 ↗ gnomad_v2 ↗ cspec ↗3
In silico splicing prediction does not support a splice-disrupting effect, with SpliceAI max delta score 0.01, which is below the RUNX1 PP3 threshold of 0.38 and within the BP4 and BP7 benign threshold of 0.20.
spliceai ↗ cspec ↗