Classification rationale
1
The NF1 c.2033C>T (p.Pro678Leu) variant has been reported in ClinVar, where most submissions classify it as likely benign or benign, with one submission classifying it as uncertain significance.
clinvar ↗2
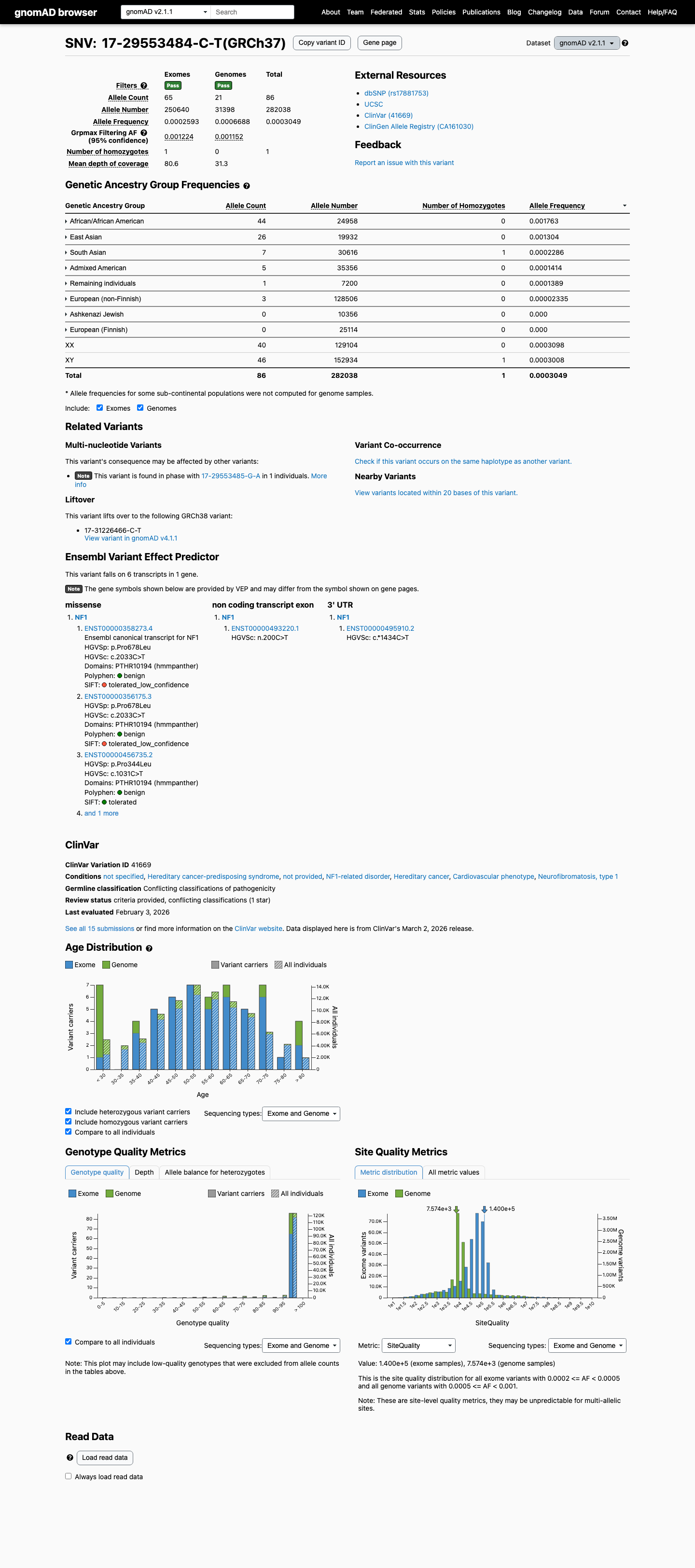
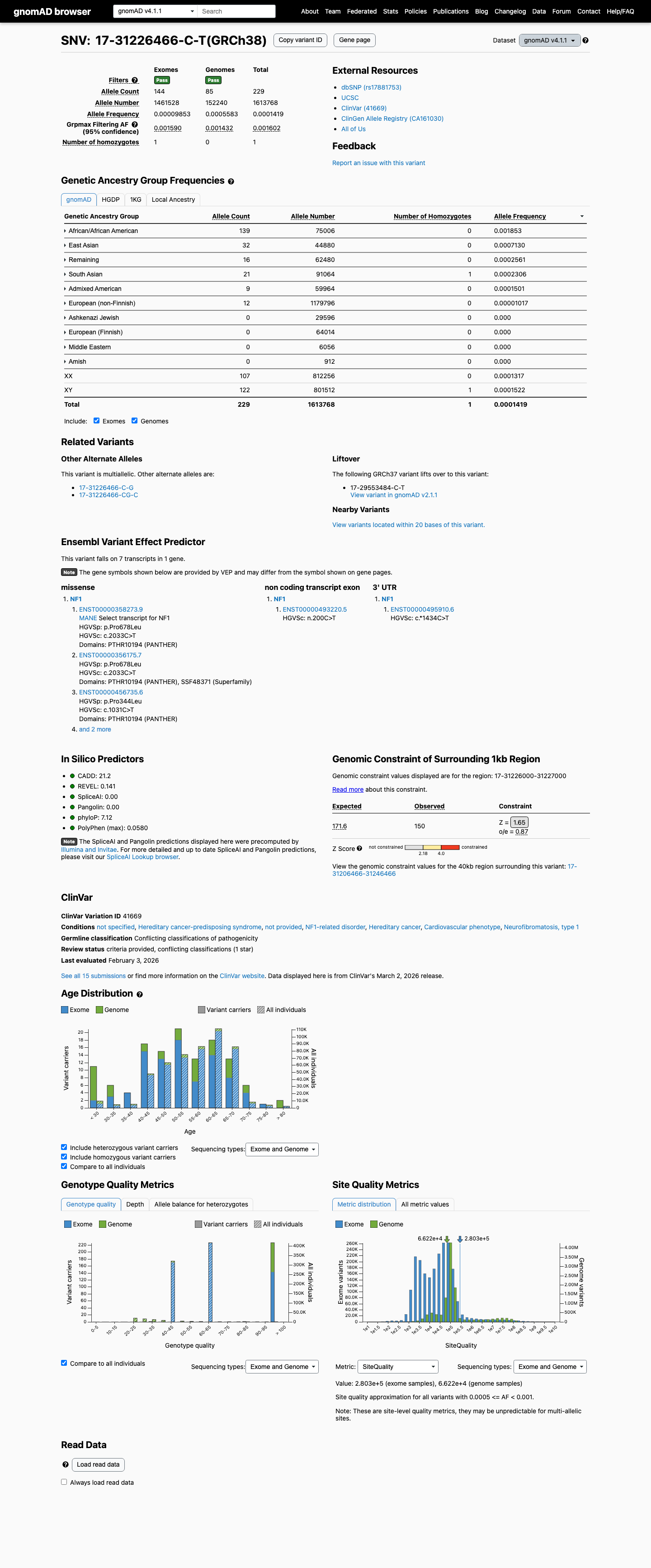
This variant is absent from gnomAD v2.1 but is present in gnomAD v4.1 at 229/1613768 alleles overall (AF 0.01419%), with the highest observed frequency in the African/African American population at 139/75006 alleles (AF 0.18532%) and one homozygote reported.
gnomad_v2 ↗ gnomad_v4 ↗3
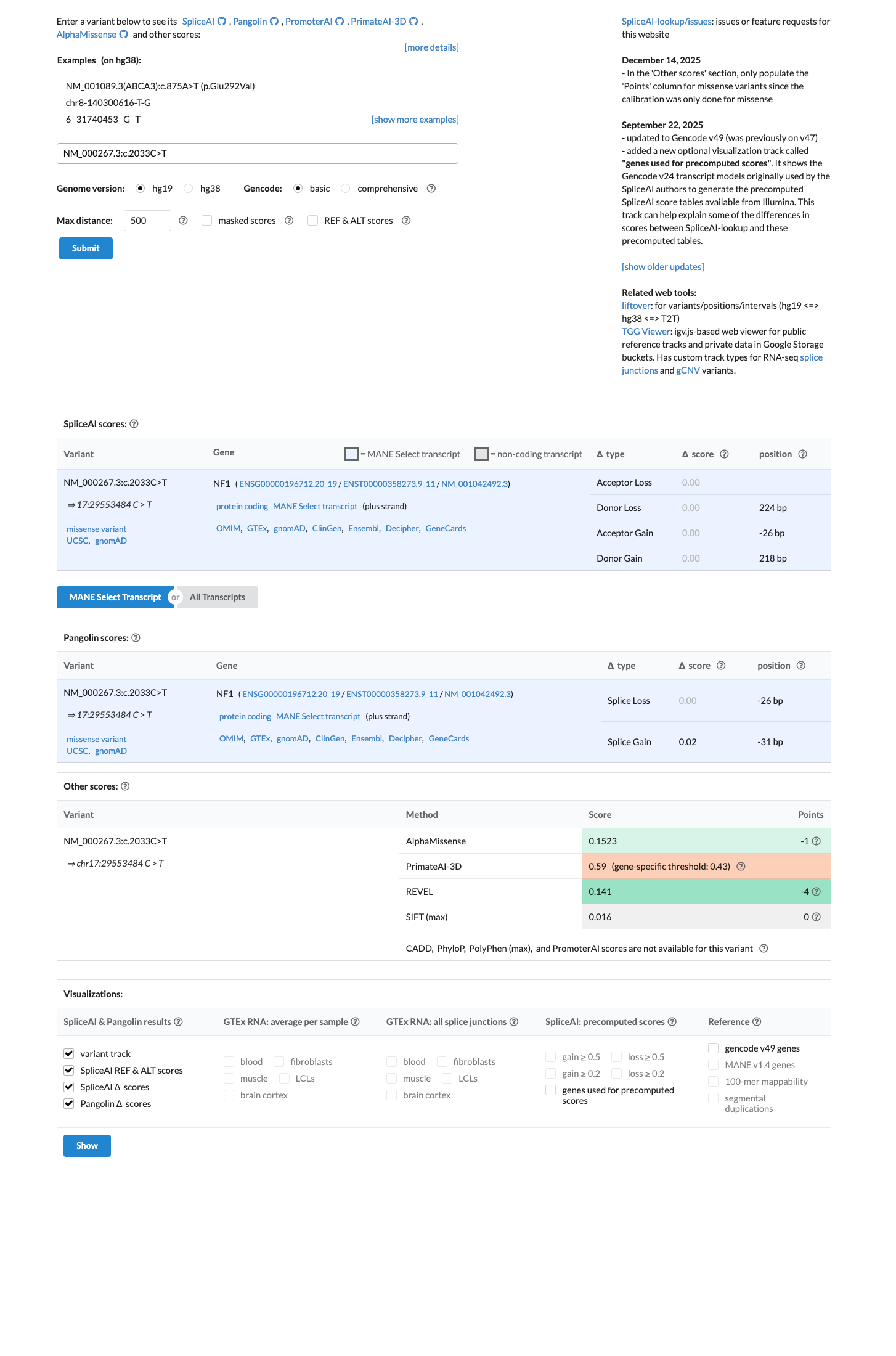
Available computational evidence supports no meaningful functional impact, with SpliceAI showing no predicted splice effect (maximum delta score 0.00), REVEL at 0.141, and BayesDel at -0.334865.
spliceai ↗