Classification rationale
1
The TET2 c.1452T>A (p.(Cys484Ter)) variant has not been reported in ClinVar.
clinvar ↗2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, which is below the 0.1% rarity threshold used for non-VCEP PM2 assessment.
gnomad_v2 ↗ gnomad_v4 ↗3
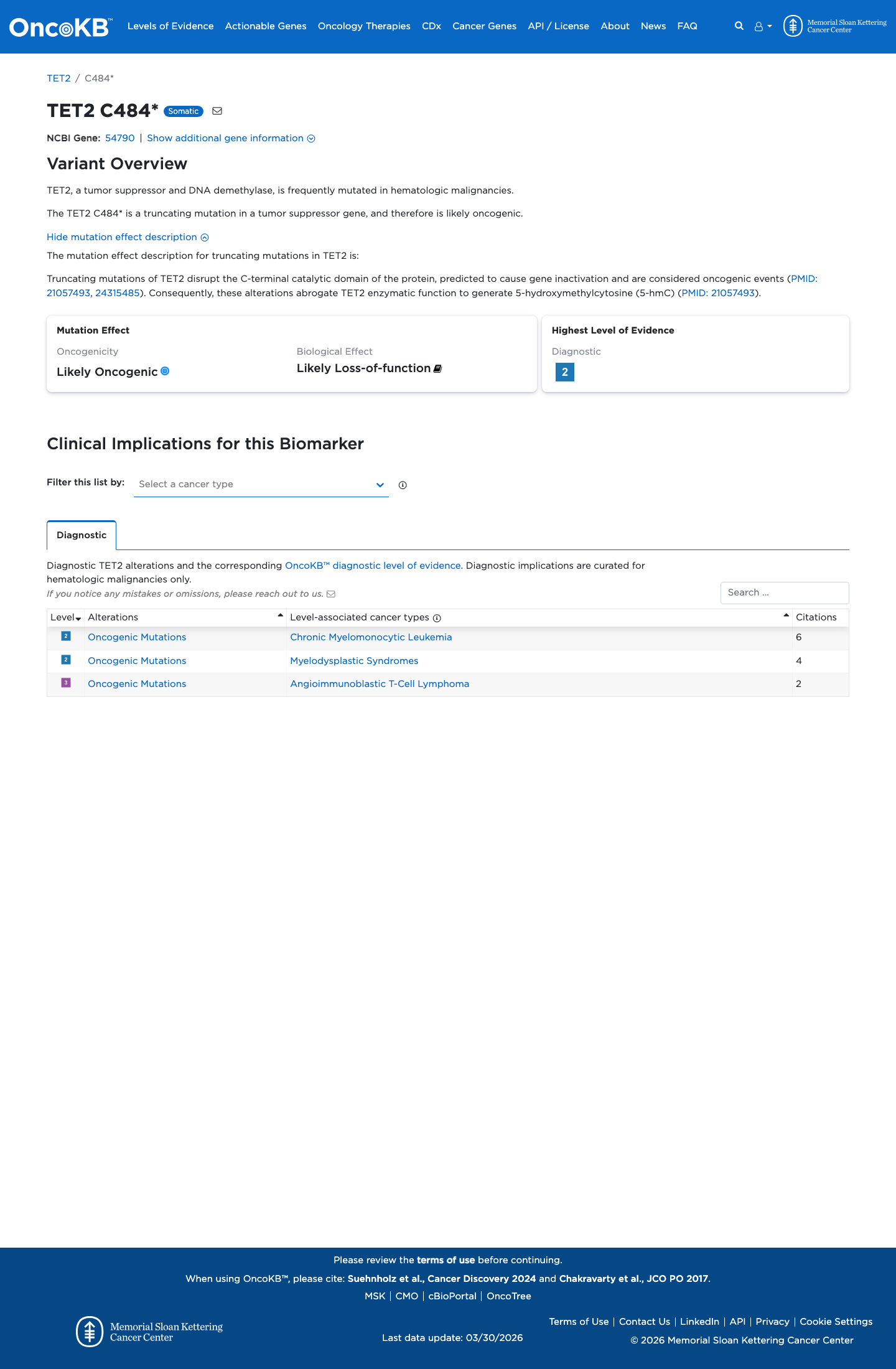
This nonsense variant introduces a premature stop codon early in the coding sequence, and available gene-level evidence supports germline TET2 loss of function as a disease mechanism, supporting PVS1 application under the generic ClinGen SVI framework.
pvs1_generic_framework ↗4
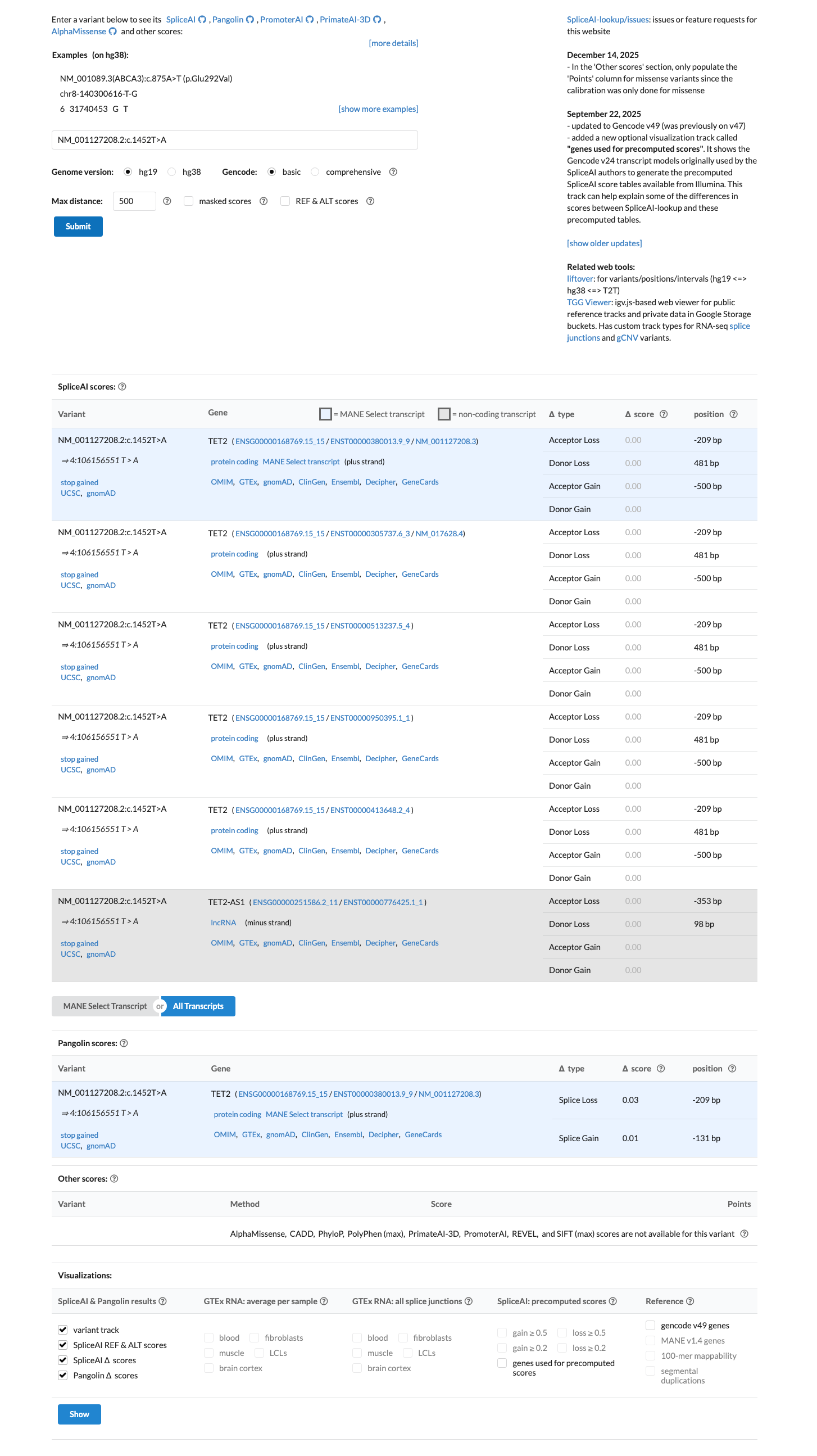
SpliceAI predicts no significant splice impact with a maximum delta score of 0.00, no REVEL score was available, and the available BayesDel score was 0.122979; these computational data were reviewed but do not outweigh the independent truncating effect of the variant.
spliceai ↗