Classification rationale
1
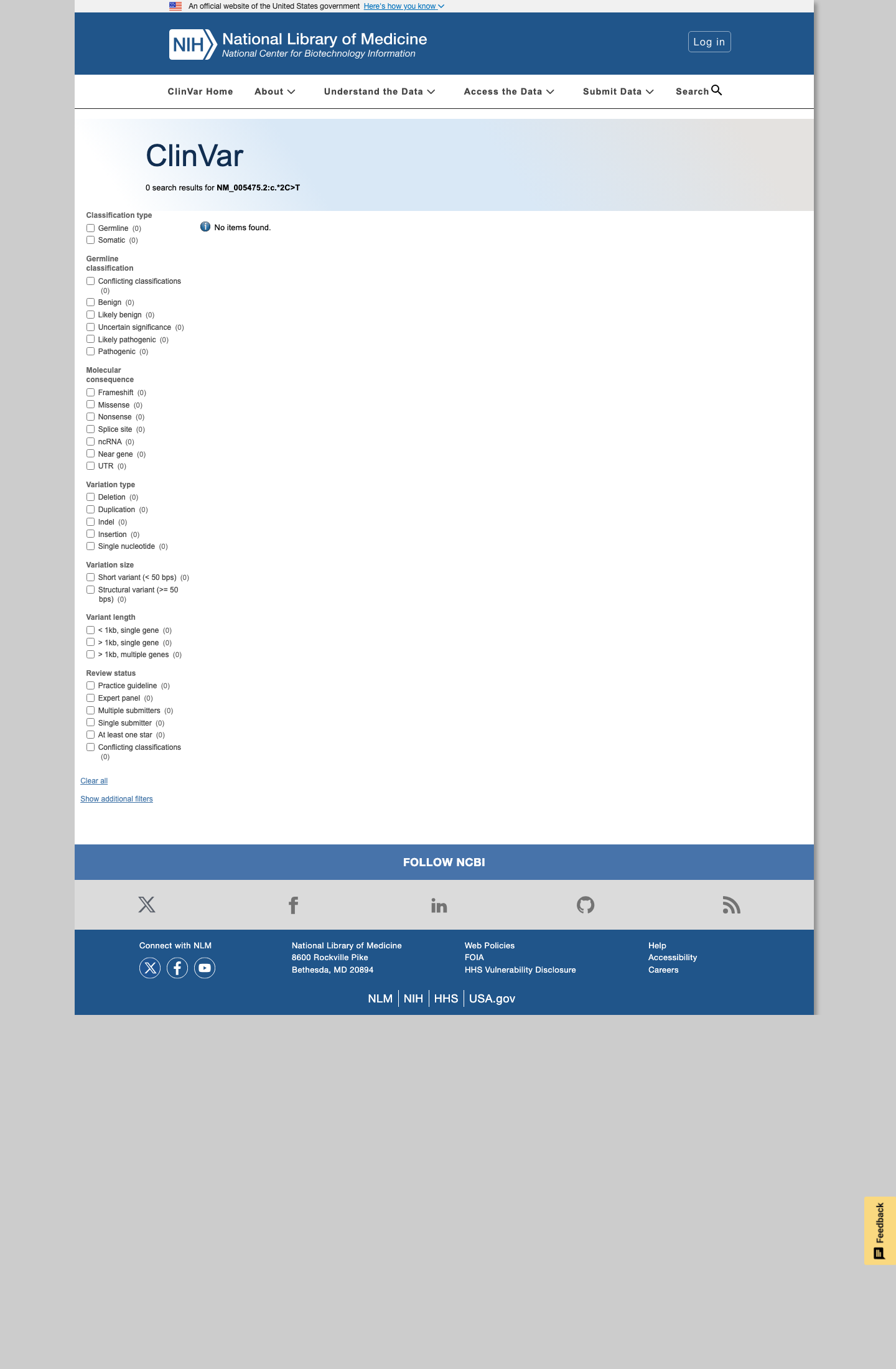
The SH2B3 c.*2C>T (NP_005466.1:p.?) variant has not been reported in ClinVar.
clinvar ↗2
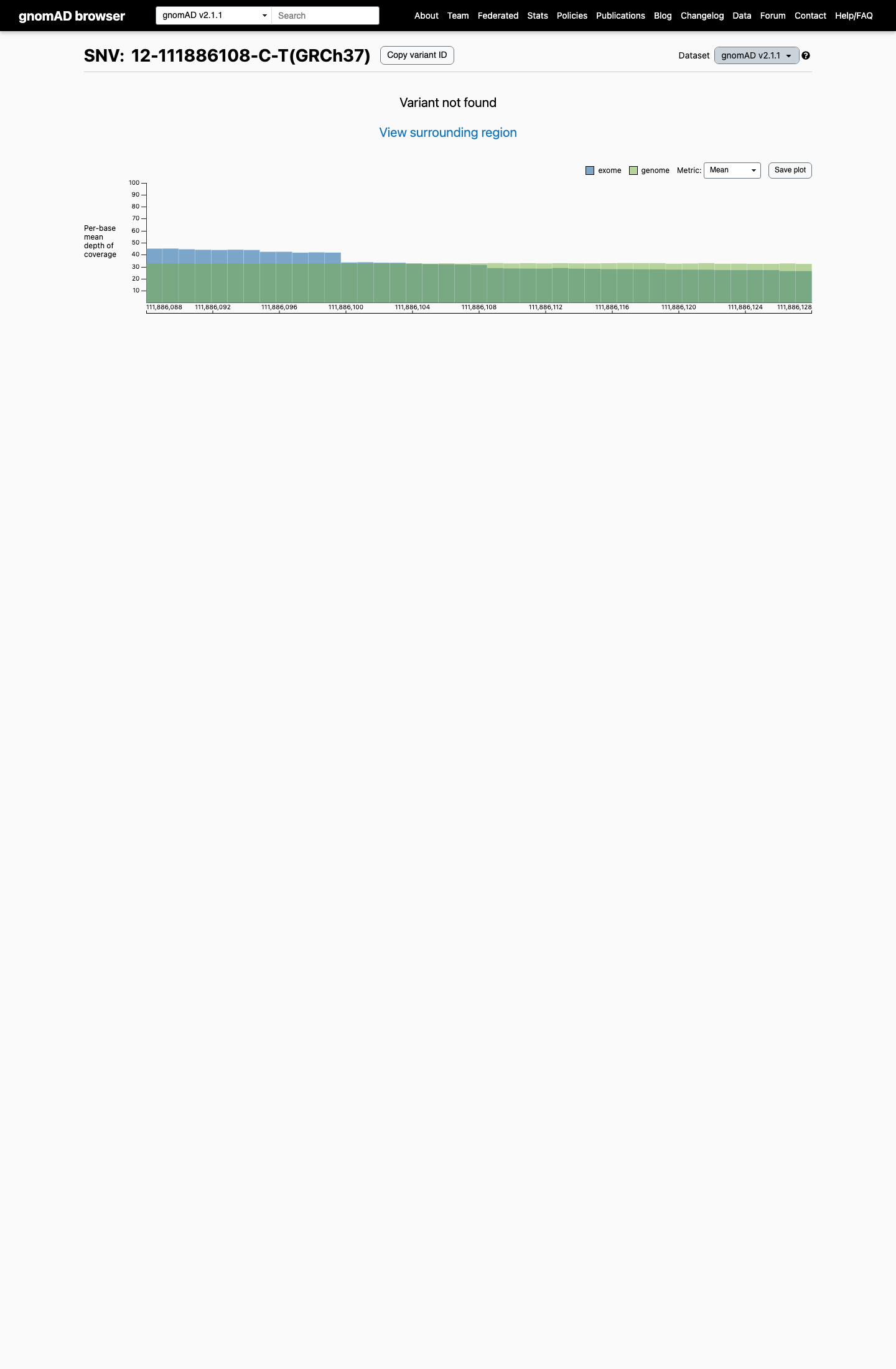
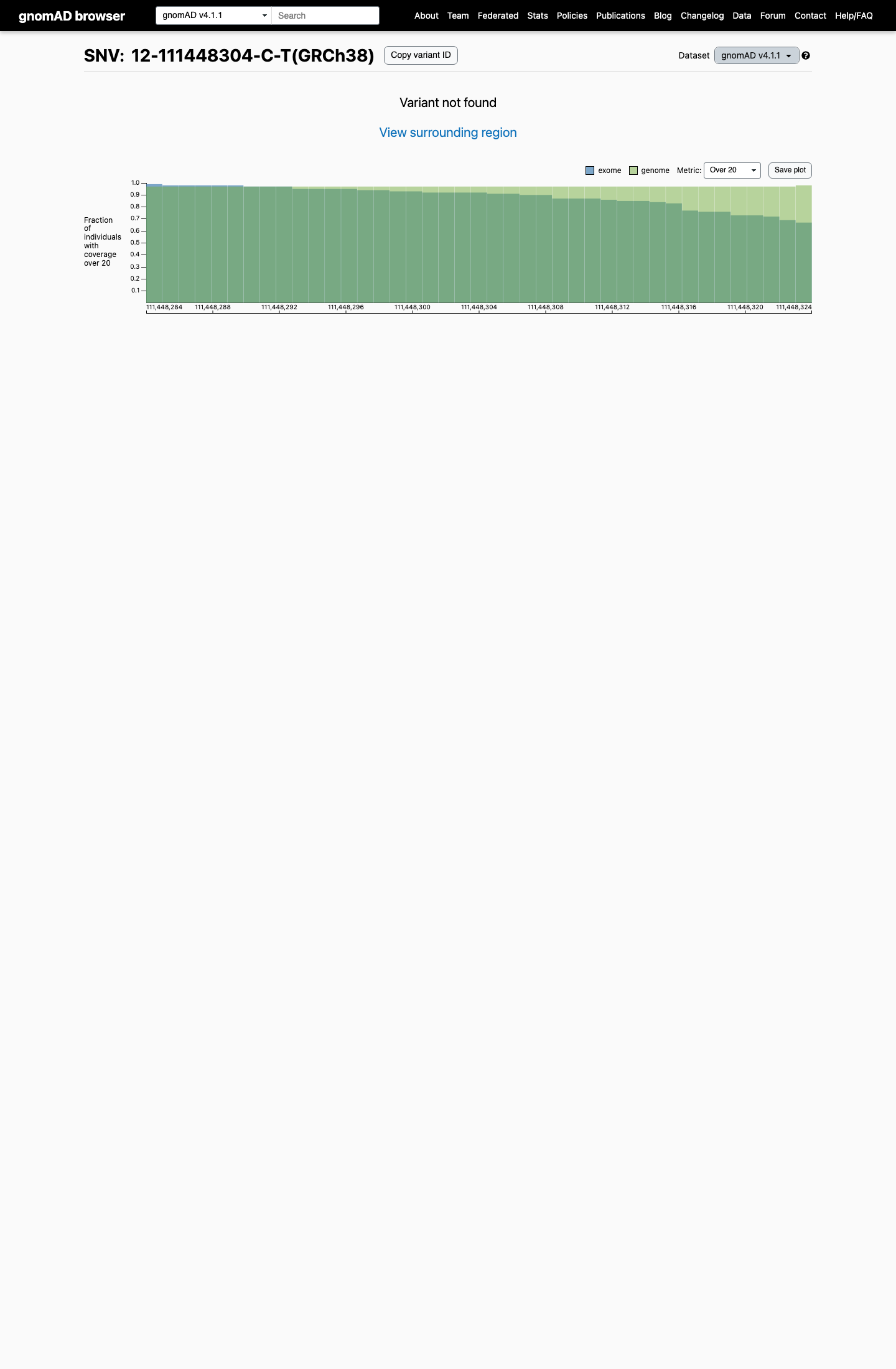
This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity below the usual PM2 threshold of 0.001 (0.1%).
gnomad_v2 ↗ gnomad_v4 ↗3
Although SH2B3 loss of function is an established disease mechanism, this variant is a 3'UTR substitution rather than a nonsense, frameshift, or canonical splice variant, so generic PVS1 criteria do not apply.
pvs1_generic_framework ↗4
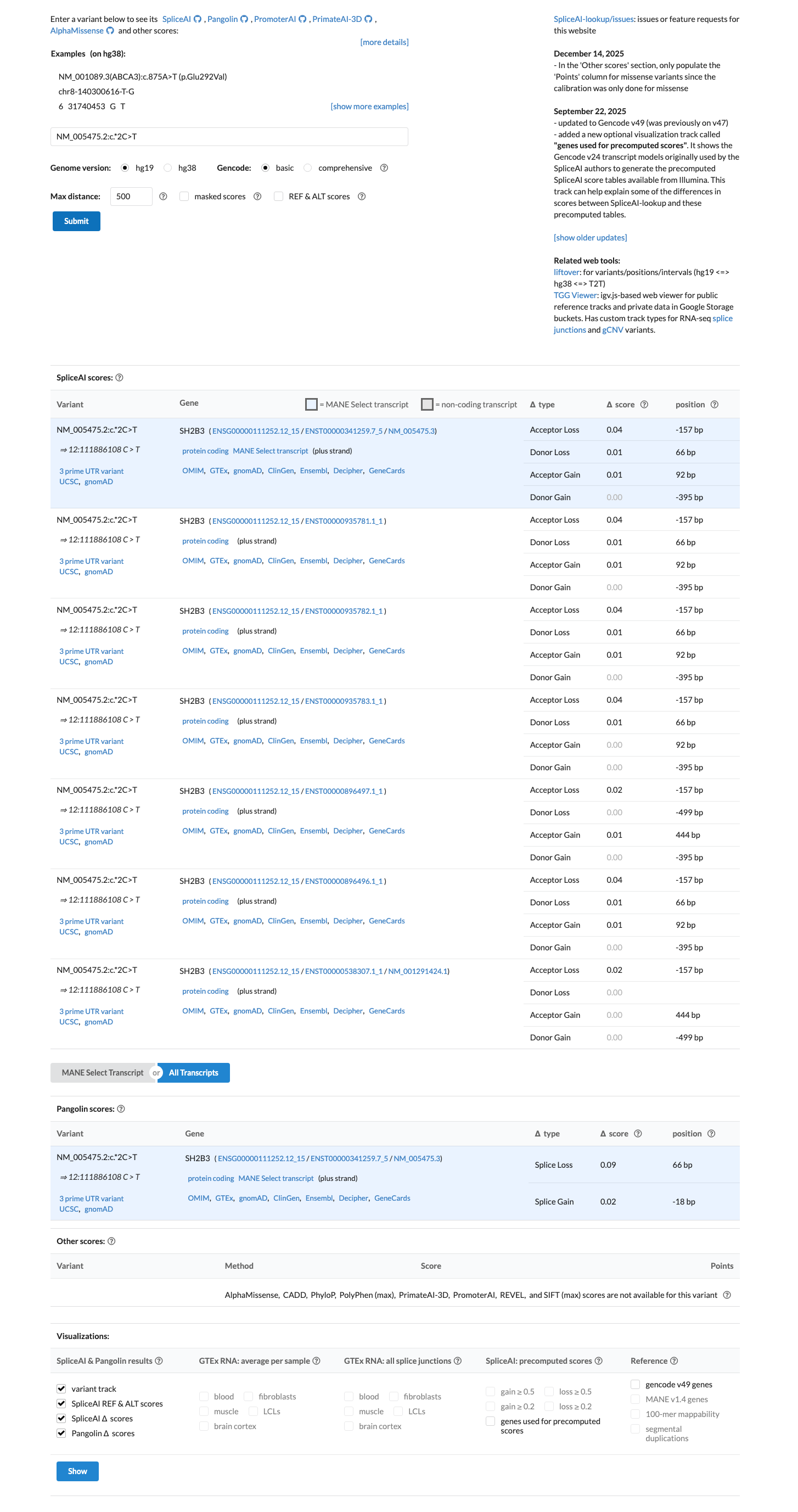
In silico splice prediction does not support a splice-altering effect, with a SpliceAI maximum delta score of 0.04.
spliceai ↗