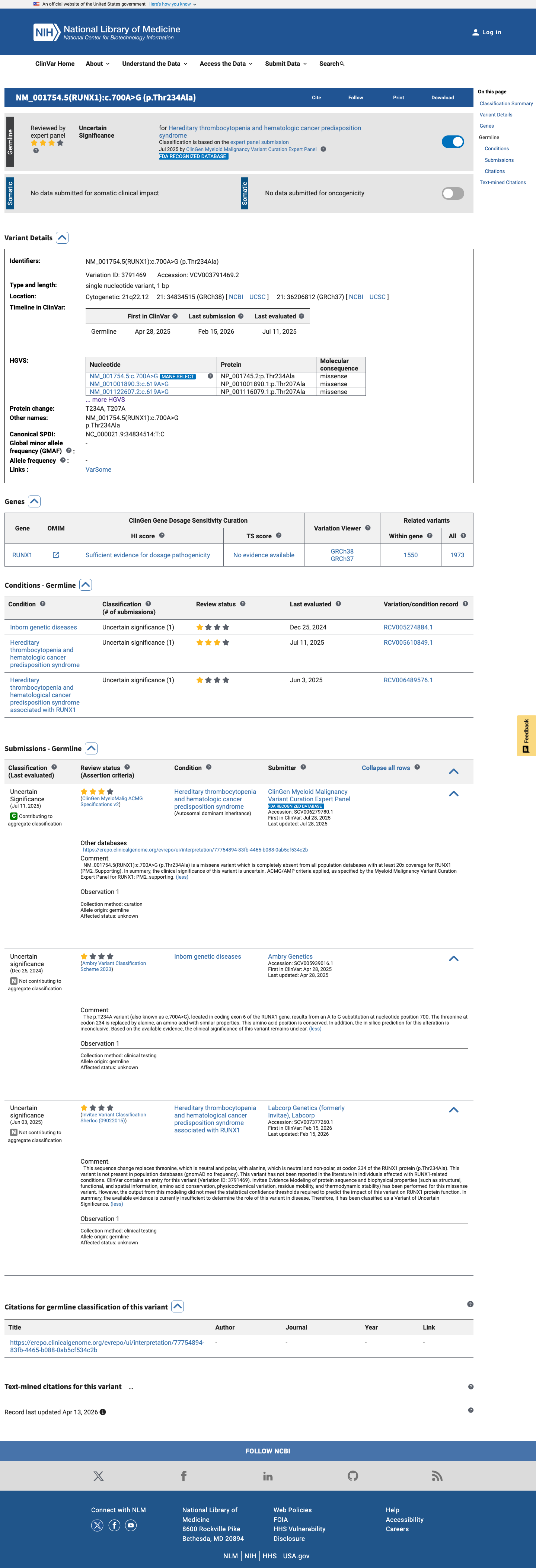
The RUNX1 NM_001001890.2:c.619A>G (p.(Thr207Ala)) variant has been reported in ClinVar and is currently classified there as uncertain significance, including an expert-panel submission from the ClinGen Myeloid Malignancy Variant Curation Expert Panel.
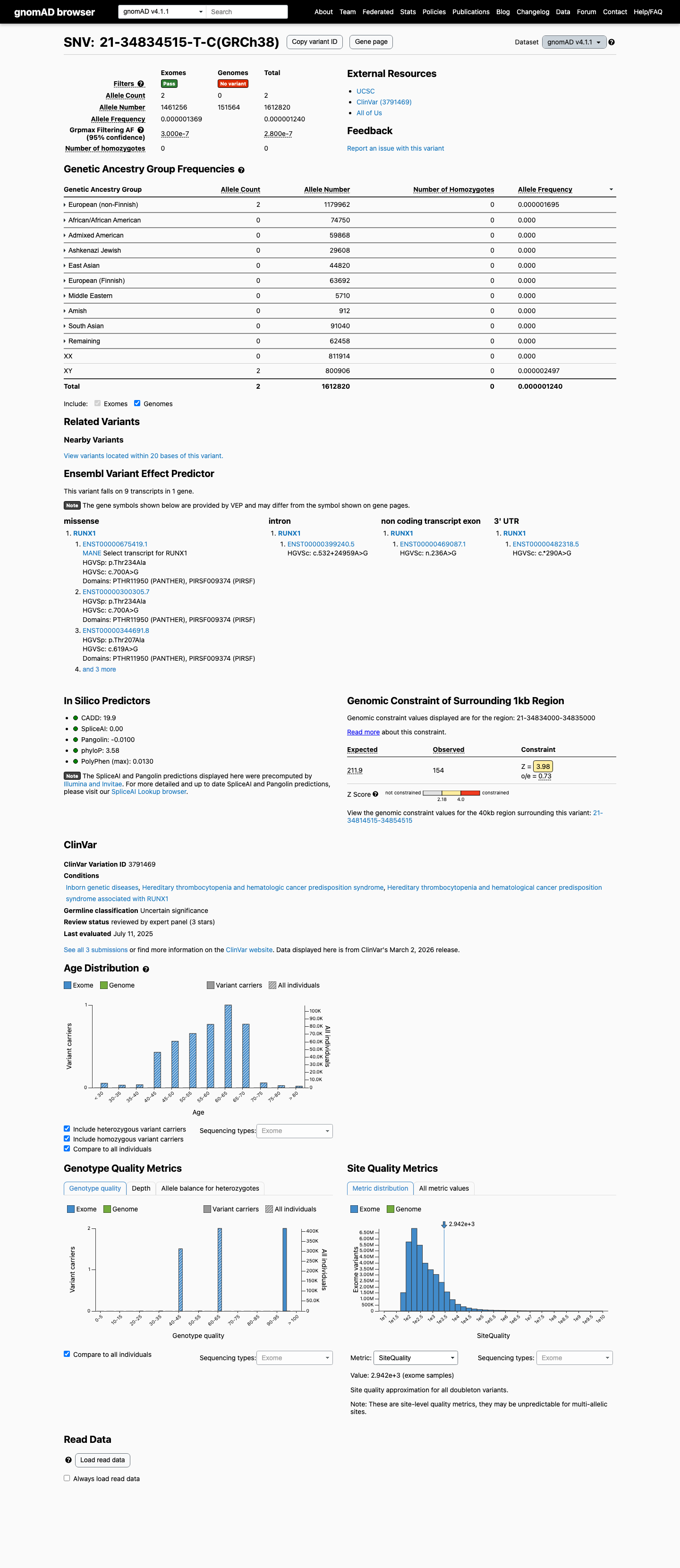
clinvar ↗This variant is absent from gnomAD v2.1 and is present at very low frequency in gnomAD v4.1, with 2/1,612,820 alleles overall and grpmax FAF 2.8e-07, which supports PM2 at supporting strength under the RUNX1 VCEP threshold of 0.00005.
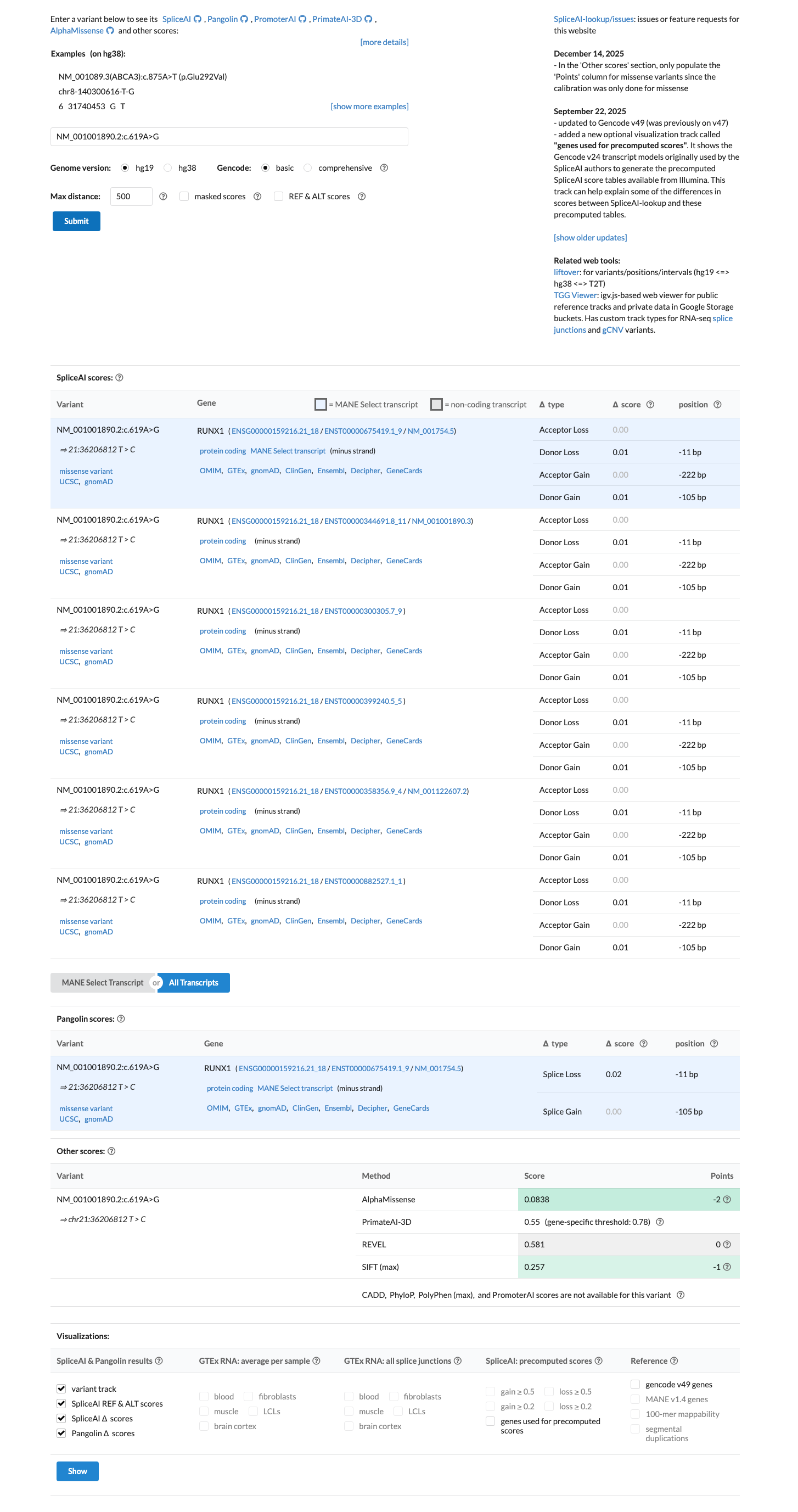
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Computational data do not support a splice effect, with SpliceAI max delta score 0.01, and the missense prediction is indeterminate under the RUNX1 VCEP computational rules because REVEL is 0.581, below the PP3 threshold of 0.88 and above the BP4 threshold of 0.50; BayesDel is 0.0617795.
spliceai ↗ cspec ↗