Classification rationale
1
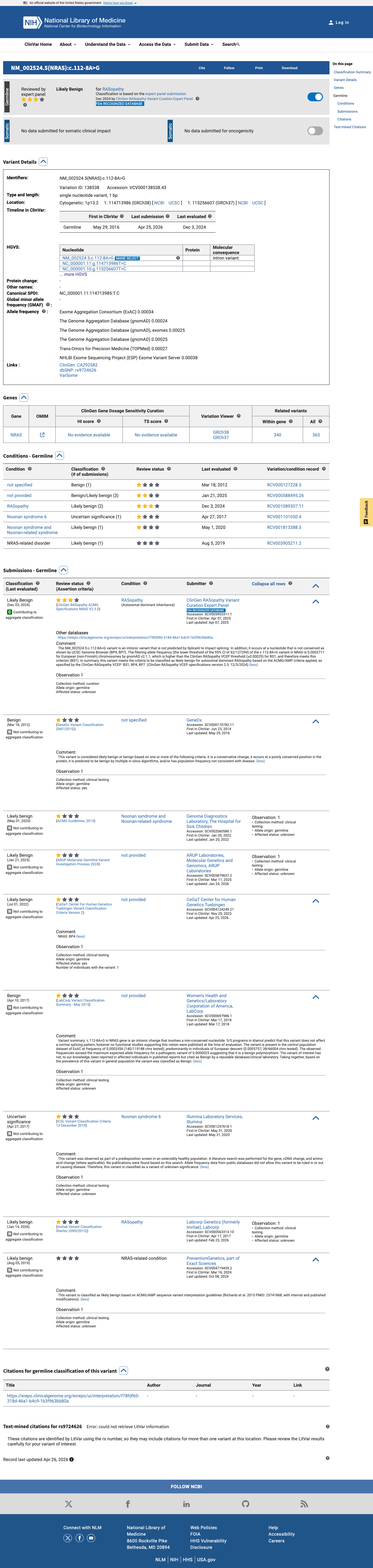
The NRAS c.112-8A>G (p.?) variant has been reported in ClinVar, where the aggregate classification is likely benign and expert panel review is present.
clinvar ↗2
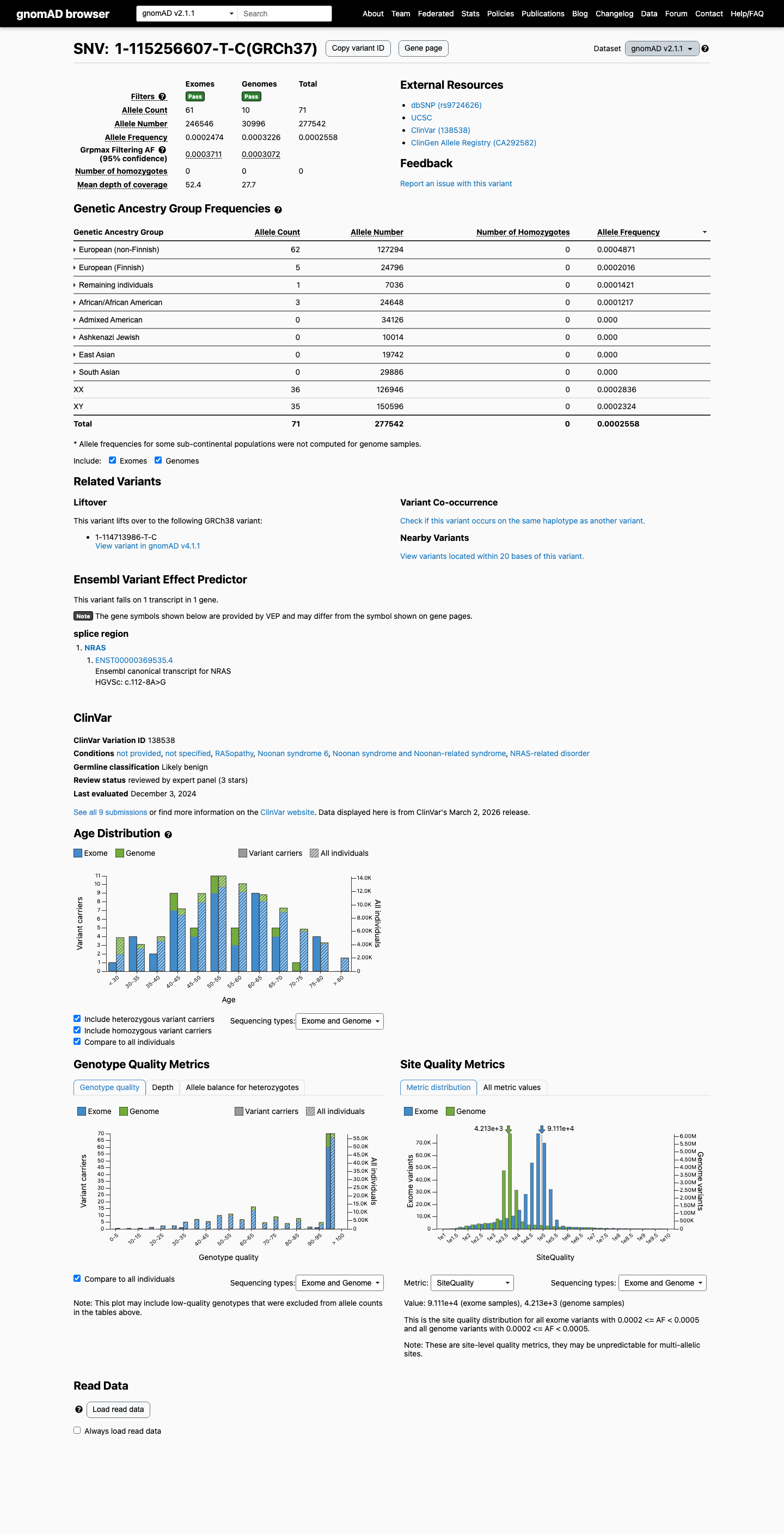
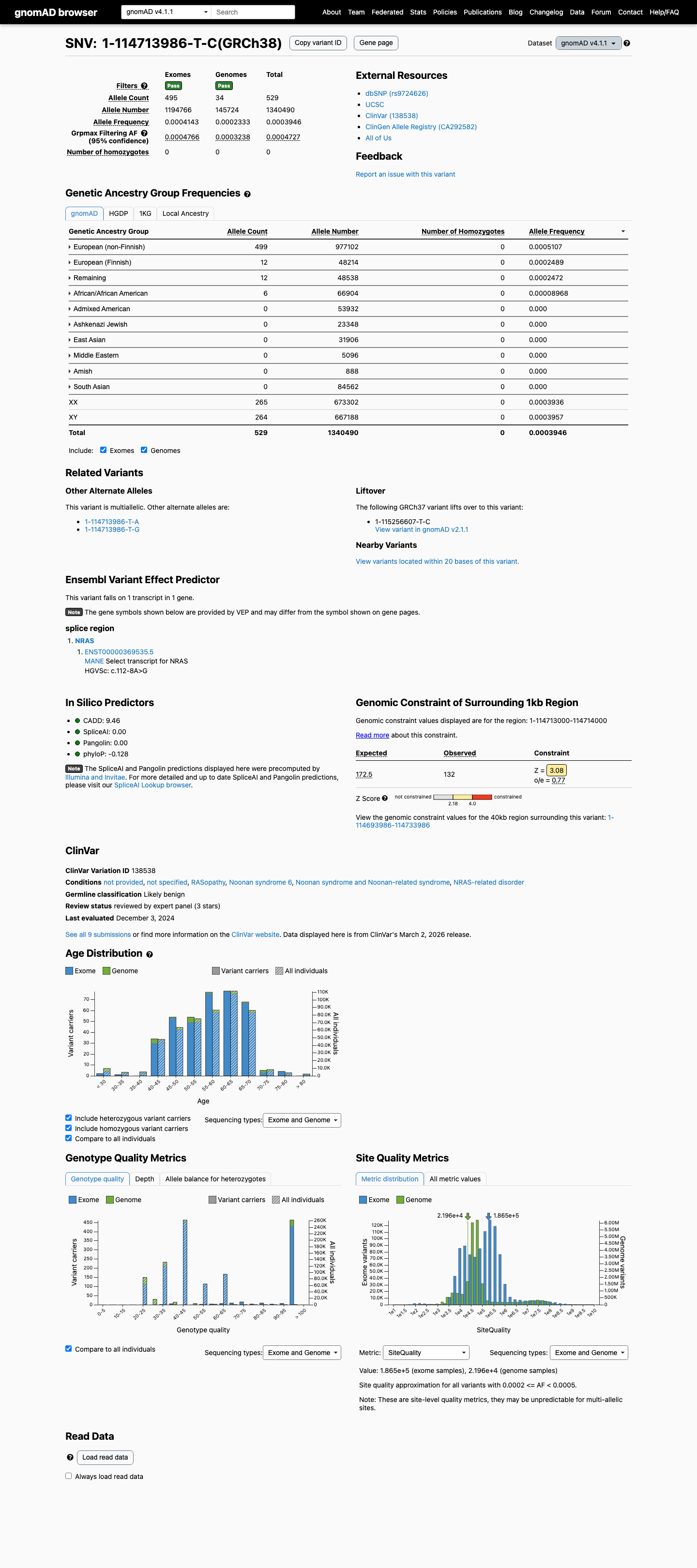
This variant is present in population databases, with gnomAD v2.1 total AF 0.02558% and grpmax FAF 0.03711%, and gnomAD v4.1 total AF 0.03946% and grpmax FAF 0.04727%, which is above the NRAS RASopathy BS1 threshold of 0.025% but below the BA1 threshold of 0.05%.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
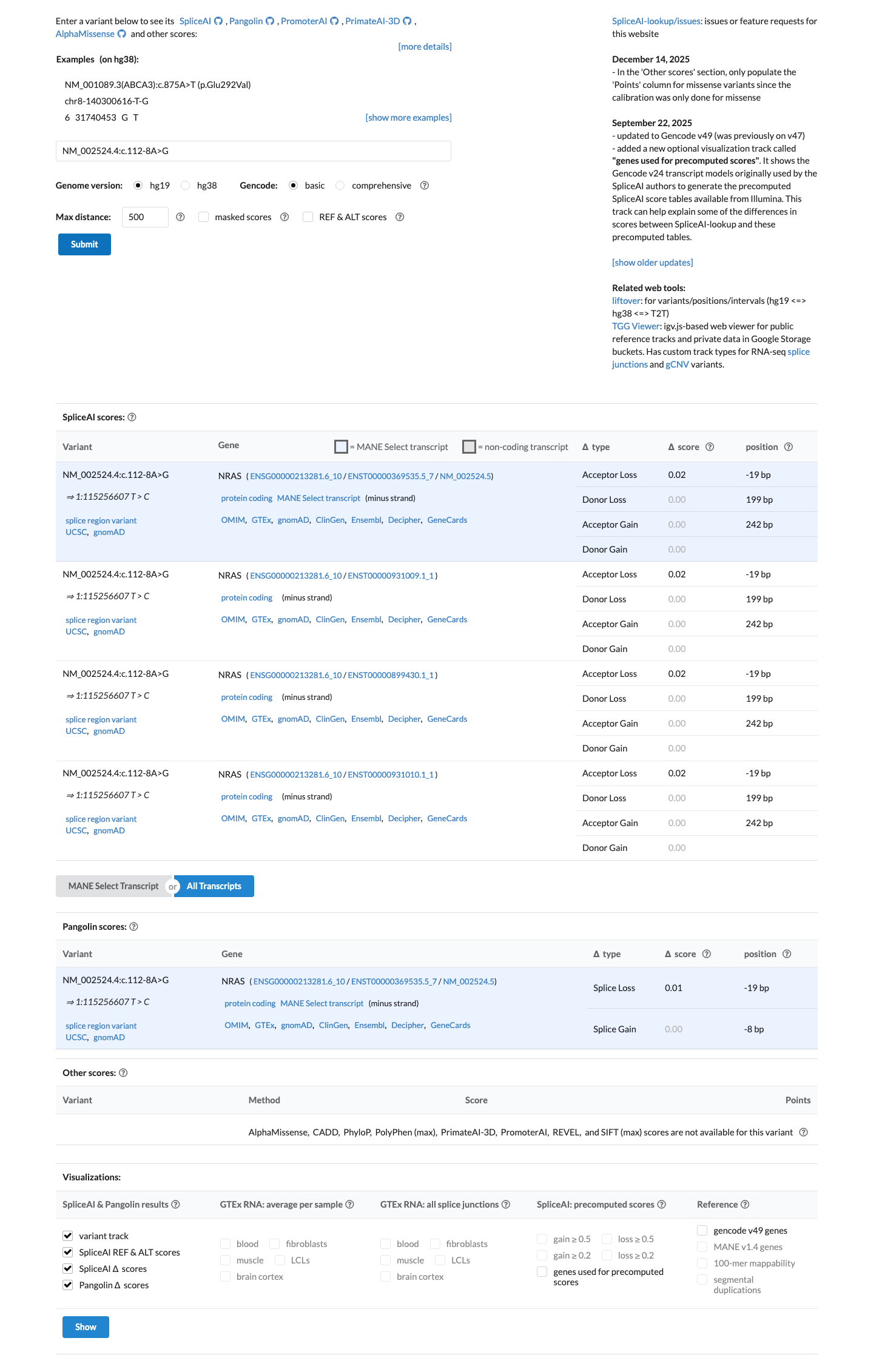
SpliceAI predicts no significant splice effect for this variant, with a maximum delta score of 0.02, which does not support a deleterious splicing prediction.
spliceai ↗ cspec ↗