The PIK3CA NM_006218.2:c.2015+9A>G (NP_006209.2:p.?) variant has been reported in ClinVar, where the aggregate classification is likely benign and the ClinGen Brain Malformations Variant Curation Expert Panel has classified it as likely benign.
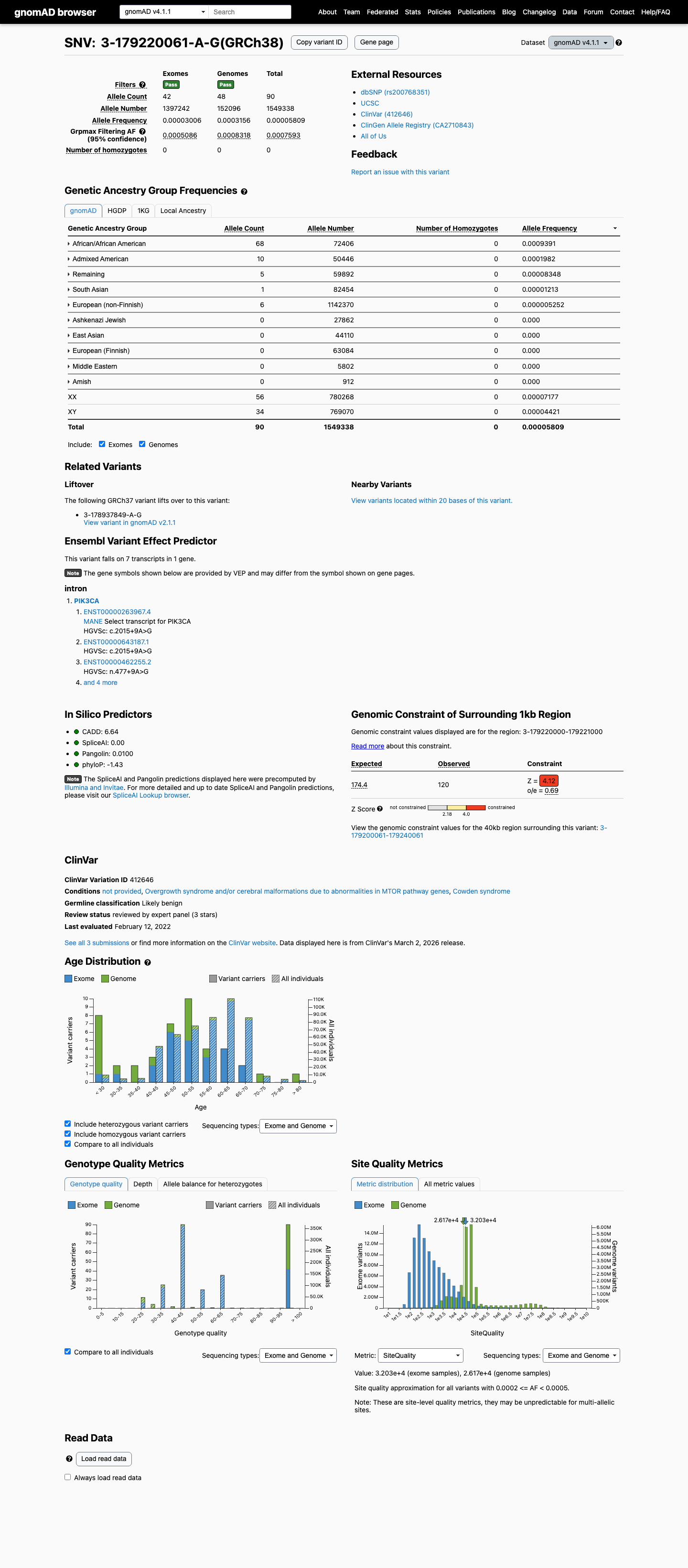
clinvar ↗This variant is present in population databases at a frequency above the Brain Malformations VCEP BS1 threshold of 0.0185%, including 31/249678 alleles in gnomAD v2.1 (AF 0.01242%) and a highest observed African/African American frequency of 0.09391% in gnomAD v4.1.
gnomad_v2 ↗ gnomad_v4 ↗No variant-specific functional study was identified showing either an abnormal or a normal effect on PIK3CA splicing or function.
cspec ↗Available computational splicing evidence could not be independently confirmed as a complete 2-of-3 benign prediction set, so BP4 and BP7 were not applied from the available records.
spliceai ↗ cspec ↗