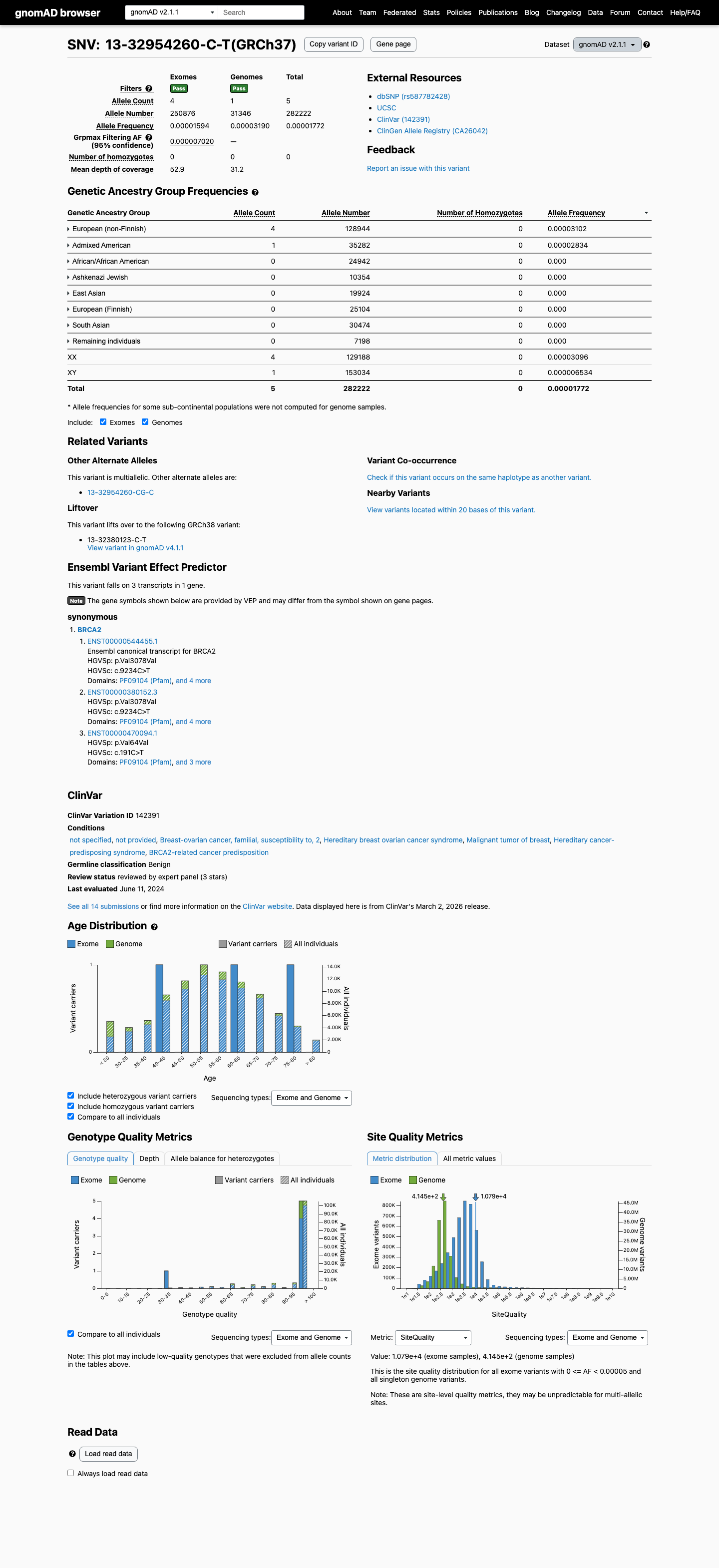
The BRCA2 c.9234C>T (p.Val3078=; p.V3078=) variant has been reported in ClinVar and is classified as Benign by the ClinGen ENIGMA BRCA1/BRCA2 expert panel.
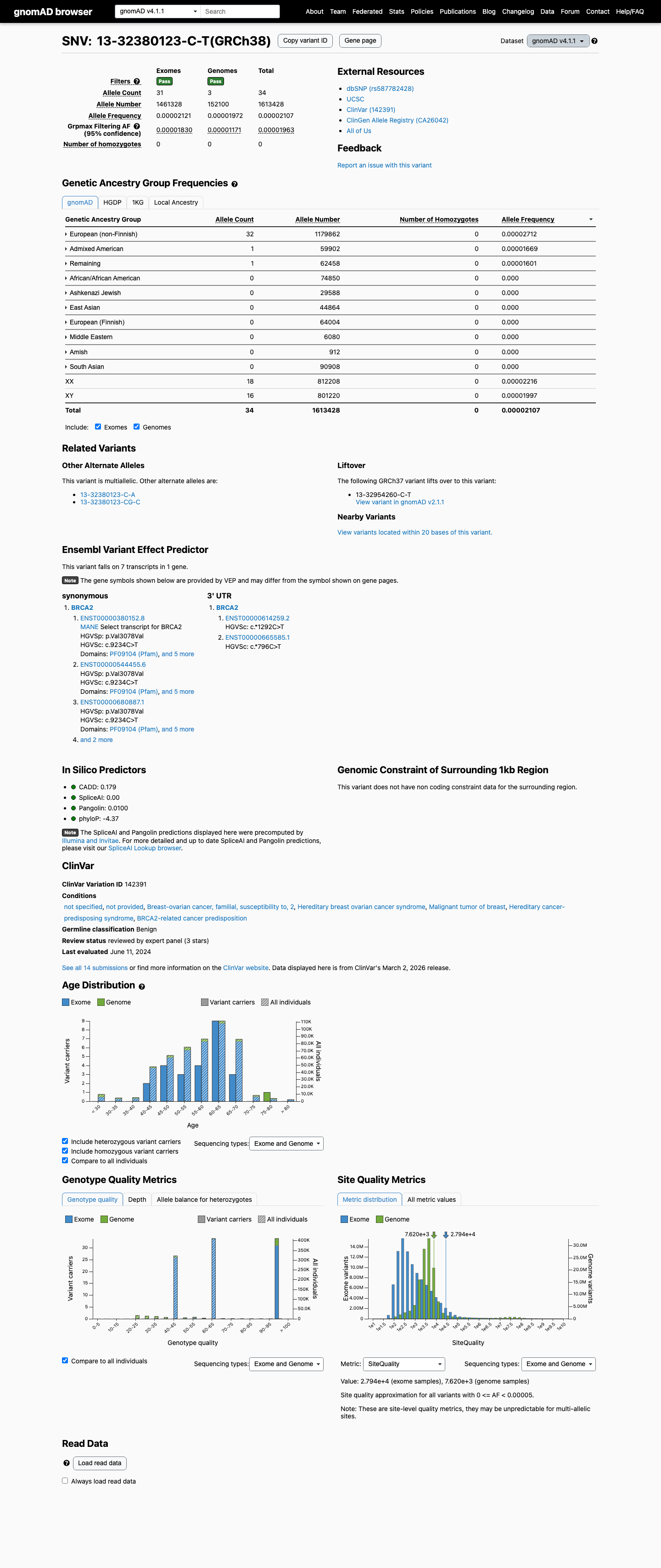
clinvar ↗This variant is present at low frequency in gnomAD, with AF 1.77e-05 in v2.1 and AF 2.11e-05 in v4.1; these frequencies are above the ENIGMA PM2 absence-from-controls requirement and below the BS1 and BA1 frequency thresholds.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In the BRCA2 clinical-history likelihood-ratio dataset, this variant has an LR of 0.284 from 12 probands, which is in the benign direction and meets ENIGMA BP5 at Supporting strength.
PMID:31853058 ↗ cspec ↗Computational splice prediction does not support splice disruption: SpliceAI shows a maximum delta score of 0.06, below the ENIGMA BP4/BP7 threshold of 0.1 and below the PP3 splice threshold of 0.2.
spliceai ↗ cspec ↗