Classification rationale
1
The KRAS c.451-14T>C (p.?) variant has been reported in ClinVar as Likely Benign, including an expert-panel likely benign assertion from the ClinGen RASopathy Variant Curation Expert Panel.
clinvar ↗2
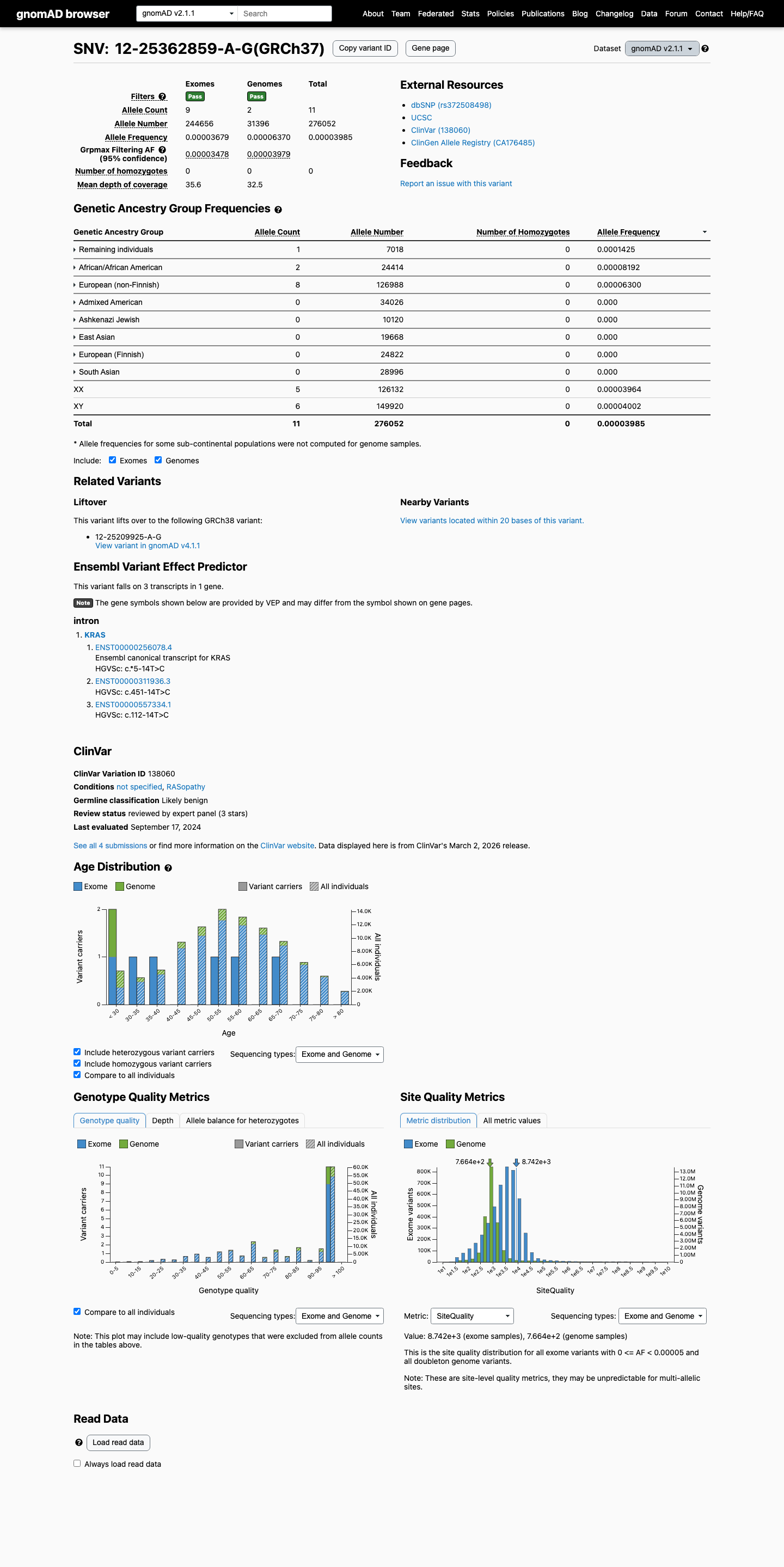
This variant is present in population databases, including gnomAD v2.1 at 0.00398% (11/276052 alleles) and gnomAD v4.1 at 0.01100% (175/1591272 alleles), which is above the PM2 absence requirement but below the KRAS RASopathy VCEP BS1 threshold of 0.025% and BA1 threshold of 0.05%.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
SpliceAI predicts no significant splice impact for this intronic change, with a maximum delta score of 0.01, which does not support PP3 and leaves BP7 incomplete without the additional conservation evidence required by the VCEP rule.
spliceai ↗ cspec ↗