Classification rationale
1
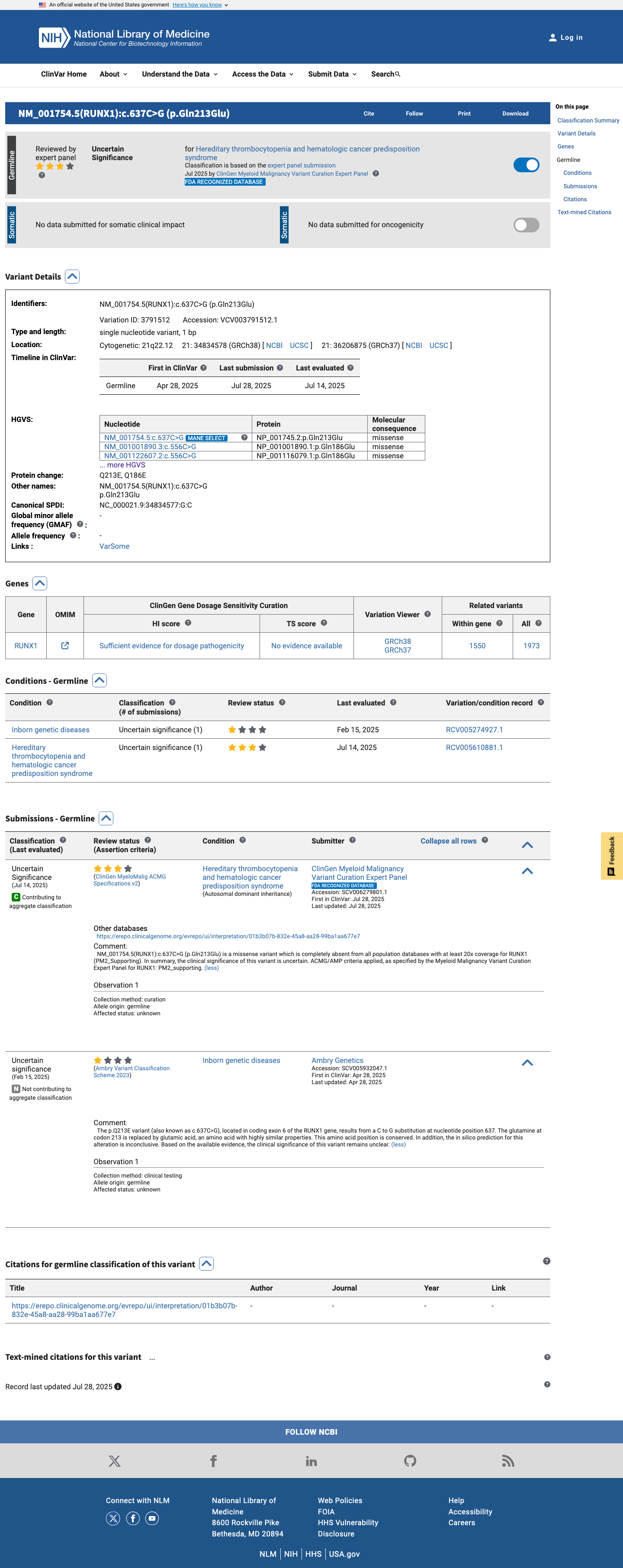
The RUNX1 c.556C>G (p.Gln186Glu) variant has not been observed in COSMIC and has been reported in ClinVar, where the ClinGen Myeloid Malignancy Variant Curation Expert Panel classified the canonical transcript change NM_001754.5:c.637C>G (p.Gln213Glu) as uncertain significance.
clinvar ↗2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting RUNX1 PM2_Supporting because the observed allele frequency is 0 and therefore below the VCEP threshold of 0.00005.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
Computational evidence does not meet the RUNX1 thresholds for either PP3 or BP4: SpliceAI predicts no significant splice effect with a max delta score of 0.01, REVEL is 0.59, and BayesDel is 0.0862396.
spliceai ↗ cspec ↗