The TP53 c.245C>T (p.Pro82Leu) variant has been reported in ClinVar, where the aggregate classification is Likely Benign with expert panel review.
clinvar ↗This variant is present at very low frequency in gnomAD v2.1 and v4.1 (overall AF 1.99644e-05 and 1.11599e-05; grpmax FAF 7.03e-06 and 8.76e-06), which is below the TP53 VCEP PM2_Supporting threshold of 0.00003 overall and below 0.00004 in any non-founder ancestry group with multiple alleles.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In the TP53 VCEP functional worksheet, p.Pro82Leu (P82L) is assigned BS3 and the summarized assay interpretation is functional with no loss-of-function, supporting retained protein function.
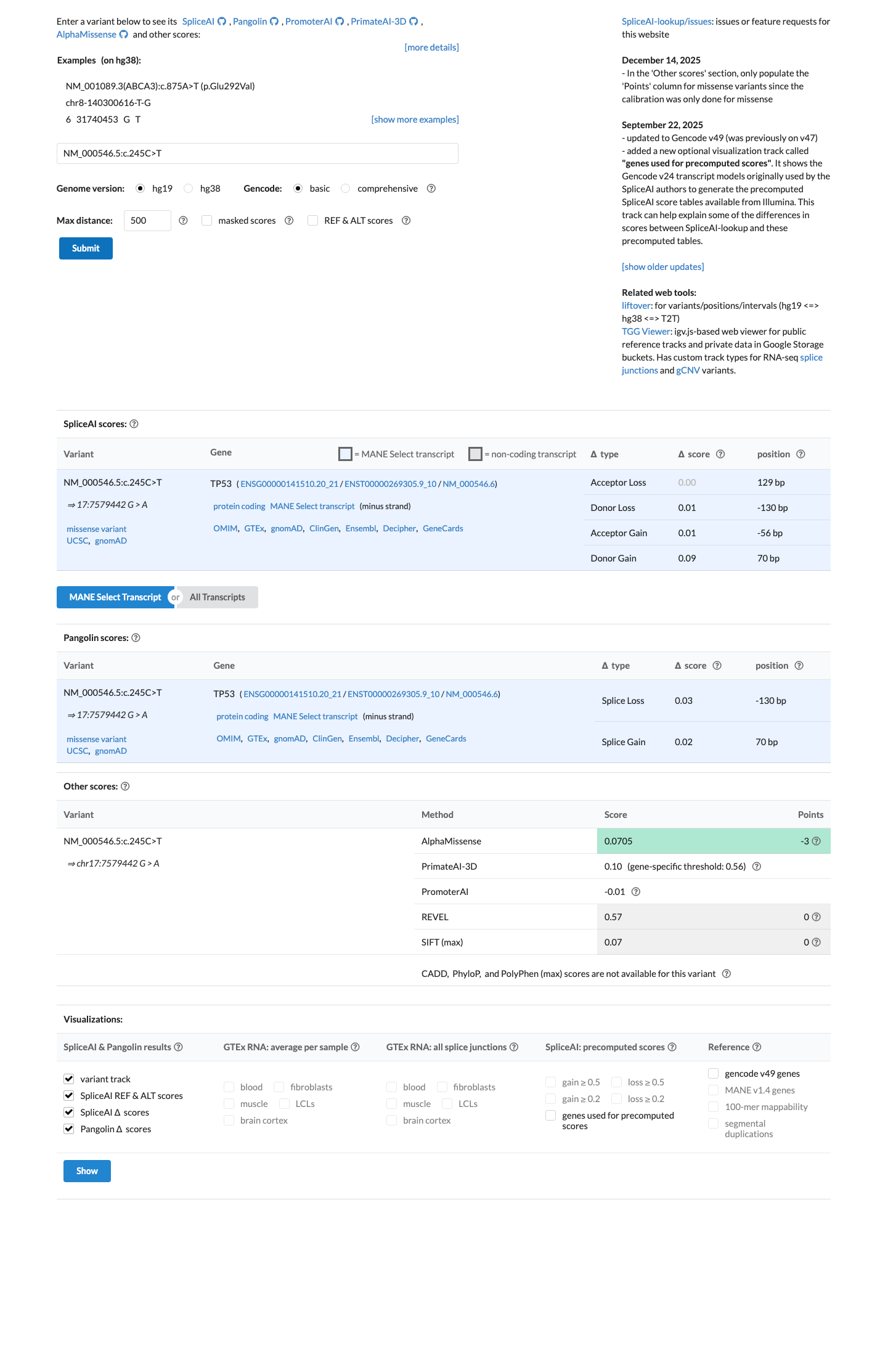
Computational assessment under the TP53 VCEP missense framework lists c.245C>T as Class C0 with BayesDel 0.0708215 and BP4, while SpliceAI predicts no splice impact (max delta 0.00); REVEL is 0.57 but does not override the TP53 VCEP benign computational assignment for this variant.
spliceai ↗