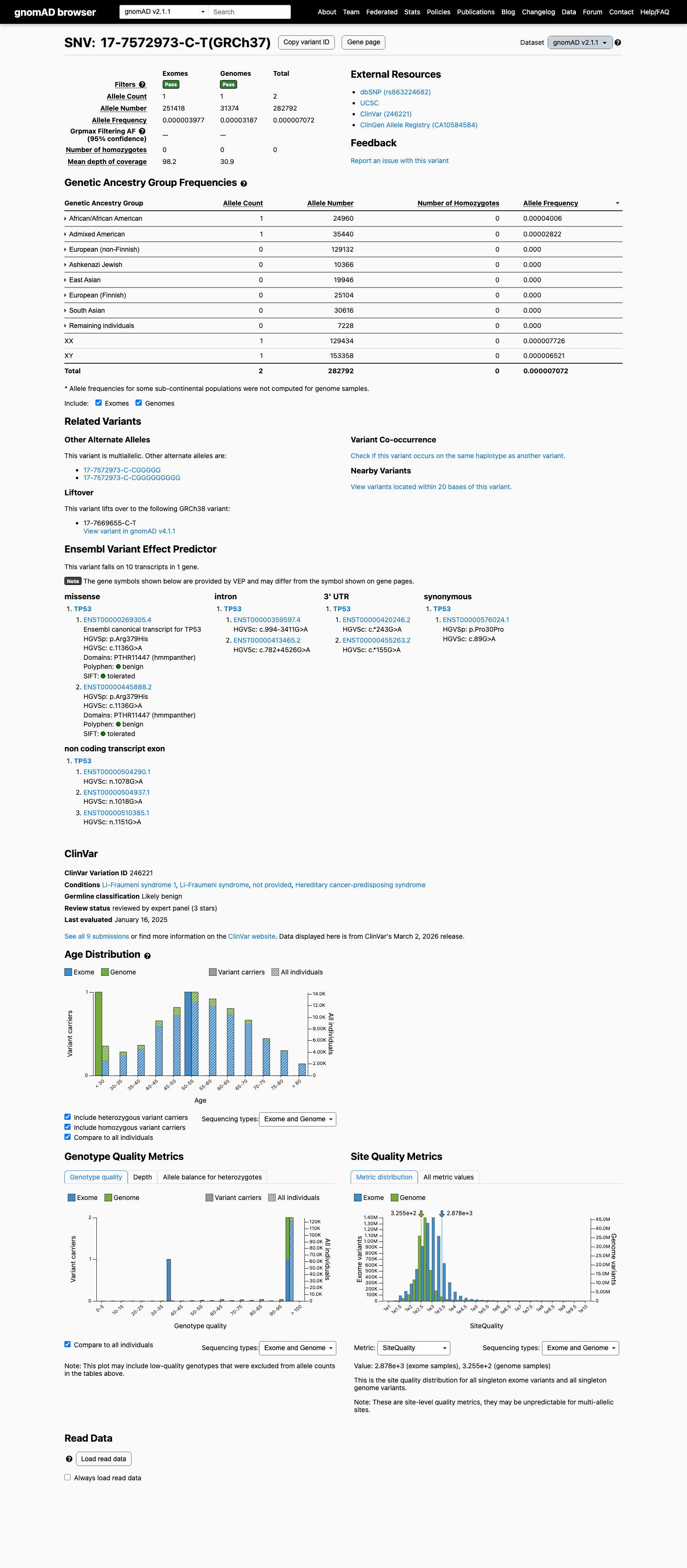
The TP53 c.1136G>A (p.Arg379His) variant has been reported in ClinVar, including a Likely Benign expert-panel assertion from the ClinGen TP53 Variant Curation Expert Panel.
clinvar ↗This variant is rare in population databases, with a gnomAD v4.1 total allele frequency of 4.95699e-06 and joint grpmax filtering allele frequency of 4.43e-06, which supports PM2_Supporting and is below the BS1 and BA1 thresholds.
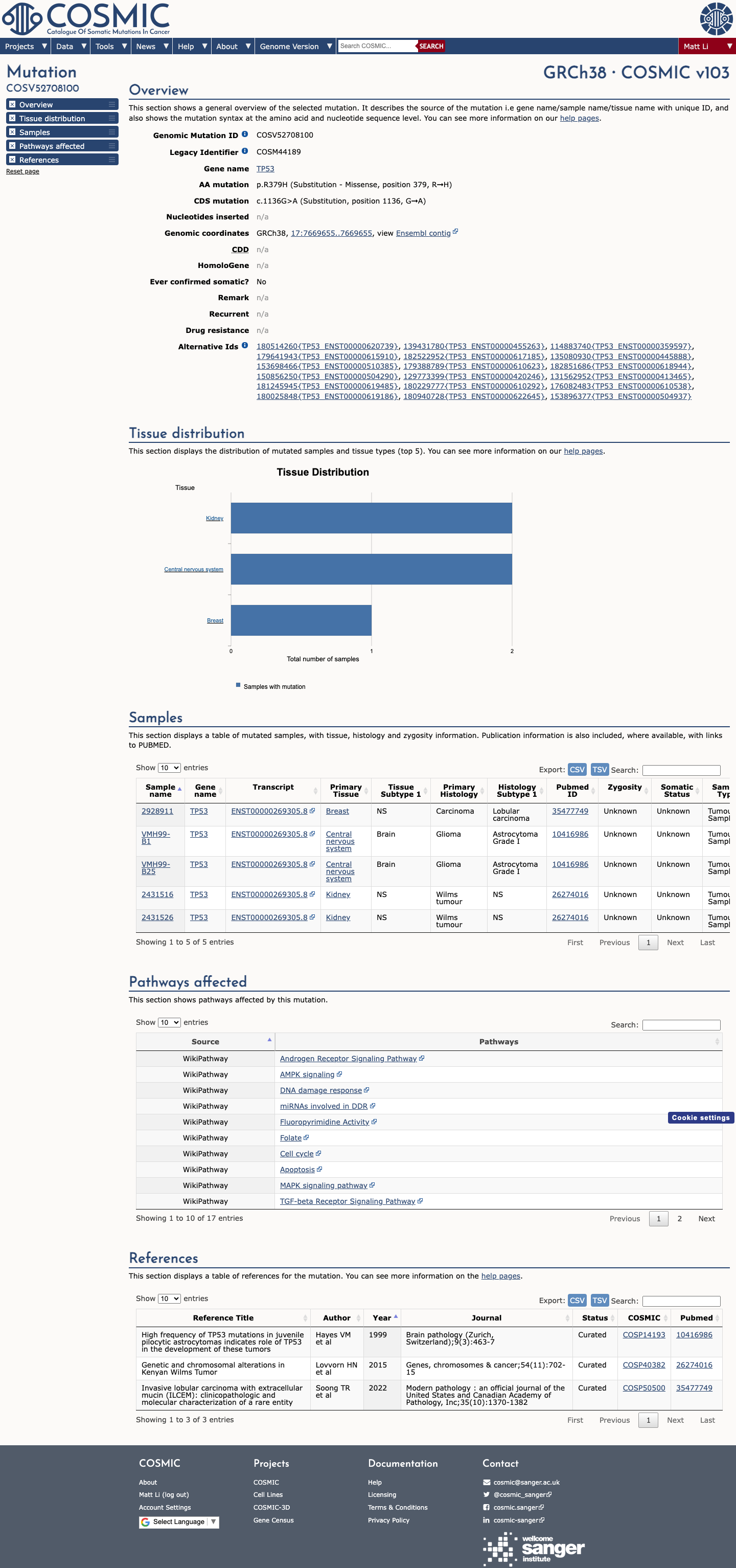
gnomad_v4 ↗ gnomad_v2 ↗ cspec ↗In published TP53 functional studies summarized by the TP53 VCEP functional worksheet, p.Arg379His was functional in the Kato assay set and showed no loss of function in the Giacomelli assay set, supporting BS3 and arguing against PS3.
PMID:12826609 ↗ PMID:30224644 ↗ cspec ↗TP53 VCEP in silico assessment supports a benign computational effect: the variant is assigned BP4_moderate in the TP53 bioinformatic worksheet, BayesDel is -0.0720003, SpliceAI shows no significant splice effect, and REVEL is 0.338.
spliceai ↗ cspec ↗