The ATM c.4397_4398delinsCG (p.Arg1466Pro) variant has been reported in ClinVar, where the overall expert-panel classification is uncertain significance and additional submissions range from uncertain significance to likely pathogenic and pathogenic.
clinvar ↗This variant is absent from gnomAD v4.1 and gnomAD v2.1, which is below the ATM VCEP PM2_supporting population threshold of less than or equal to 0.001%.
gnomad_v4 ↗ gnomad_v2 ↗ cspec ↗In an ATM VCEP supplementary variant table, the equivalent single-nucleotide representation c.4397G>C (p.Arg1466Pro) is listed as non-functional with high confidence, supporting an abnormal ATM protein effect.
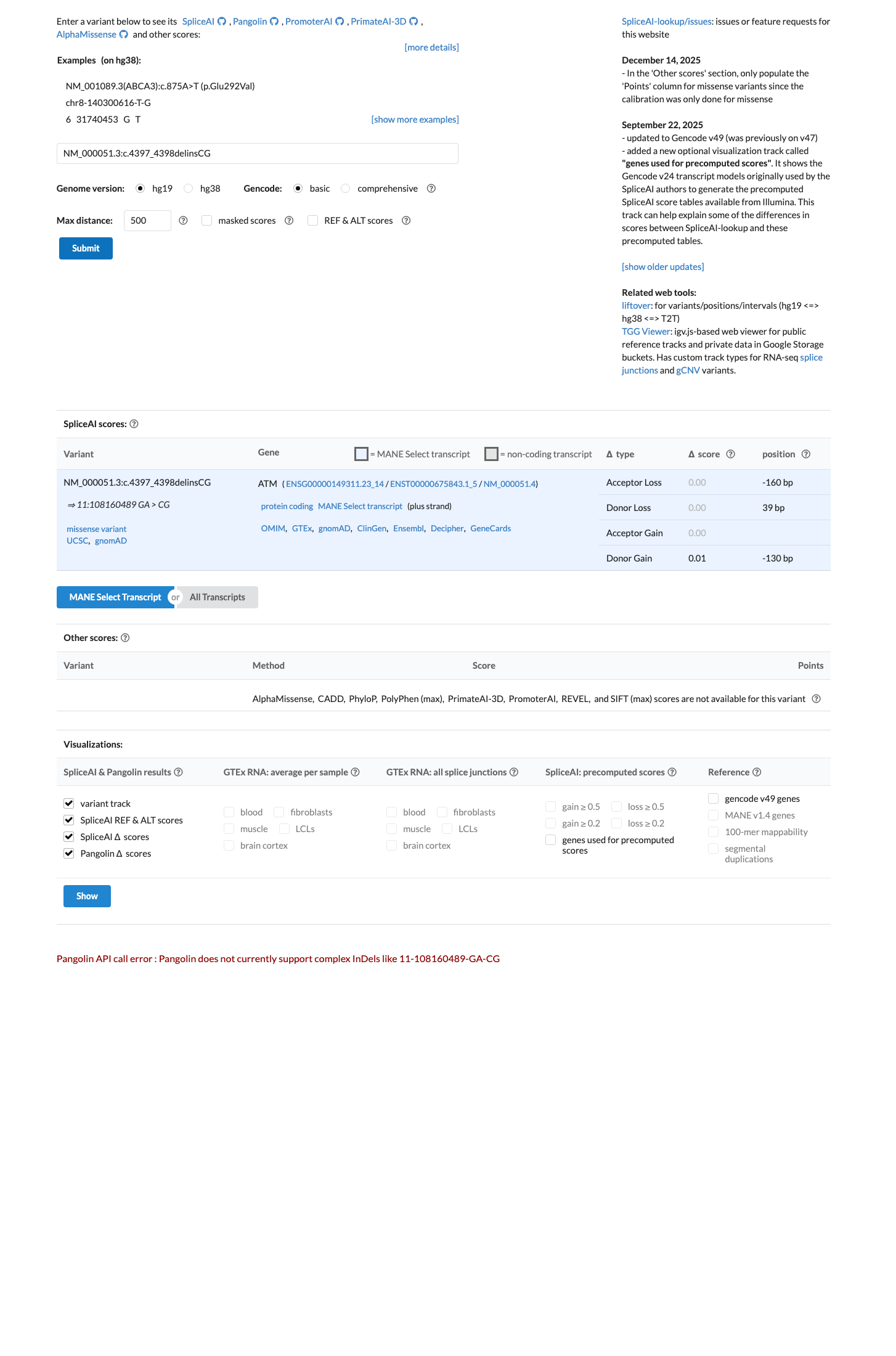
cspec ↗SpliceAI predicts no significant splice impact for this variant with a max delta score of 0.01, and the equivalent c.4397G>C missense representation has a REVEL score of 0.677 in the ATM VCEP supplementary variant table, which is below the ATM PP3 threshold of greater than 0.7333 and above the BP4 threshold of less than or equal to 0.249.
spliceai ↗ cspec ↗