Classification rationale
1
2
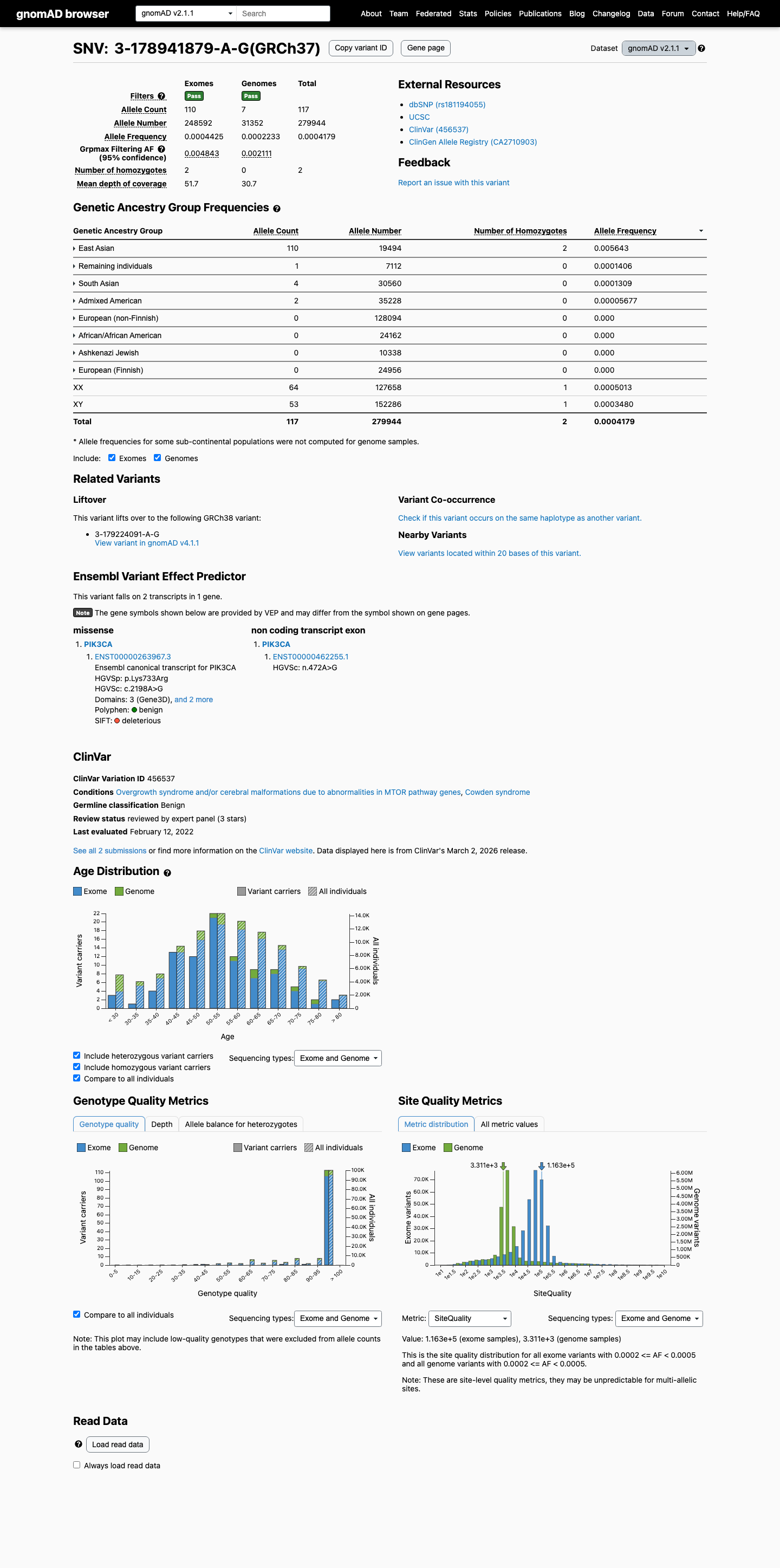
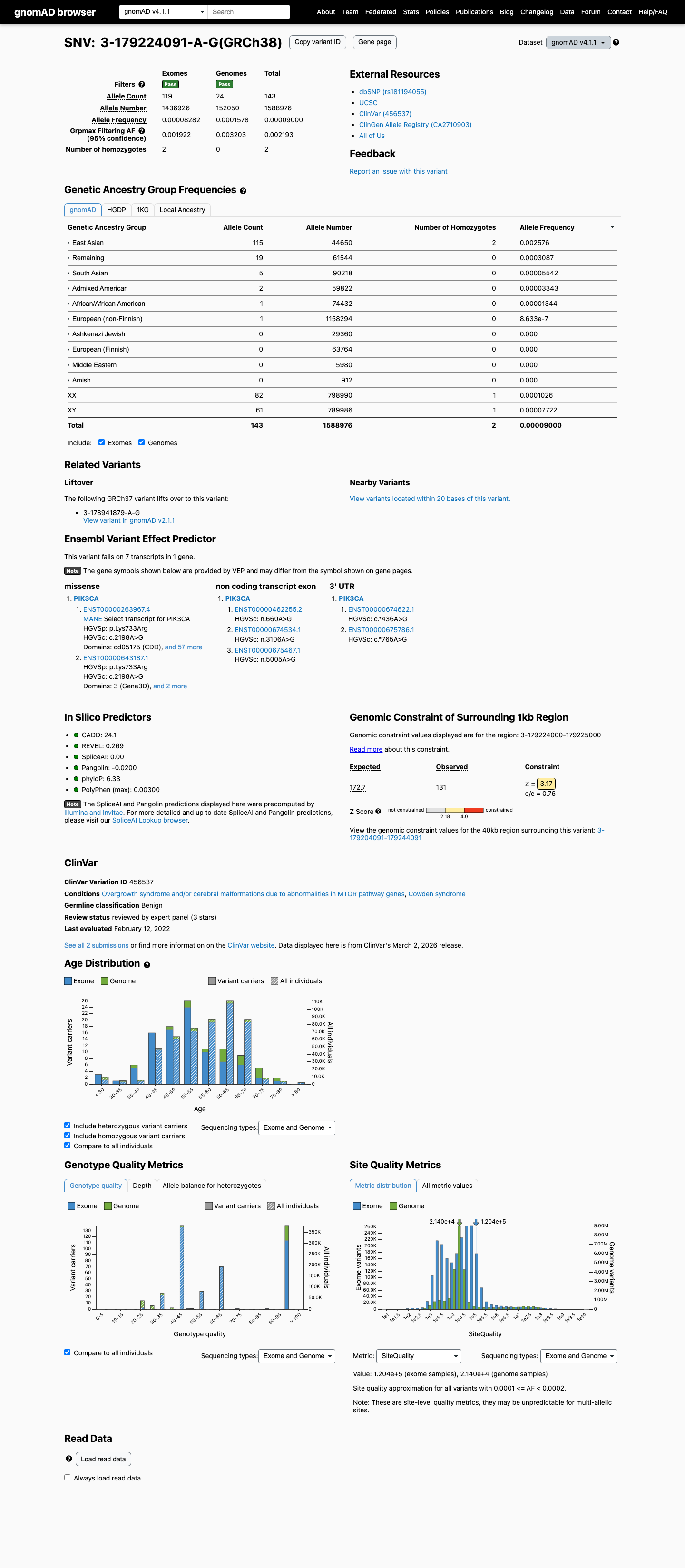
This variant is present in population databases at a frequency above the Brain Malformations VCEP benign thresholds, with East Asian AF 0.56428% in gnomAD v2.1 and 0.25756% in gnomAD v4.1, exceeding both the BS1 threshold of 0.0185% and the BA1 threshold of 0.0926%.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
In silico review does not support a splice-disrupting effect, with SpliceAI max delta score 0.12; REVEL was 0.269 and BayesDel was -0.264459, although PP3 and BP4 are not applicable to this missense gain-of-function interpretation under the Brain Malformations VCEP.
spliceai ↗ cspec ↗