Classification rationale
1
2
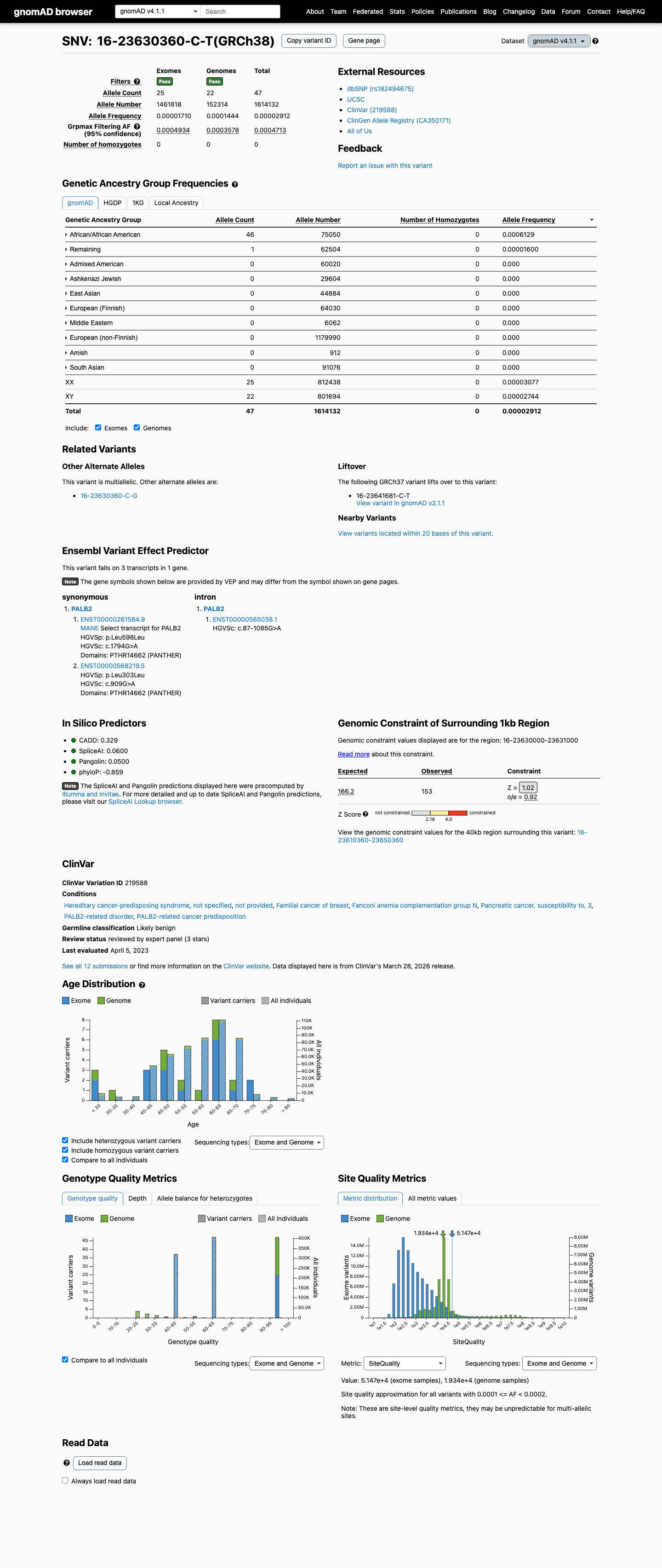
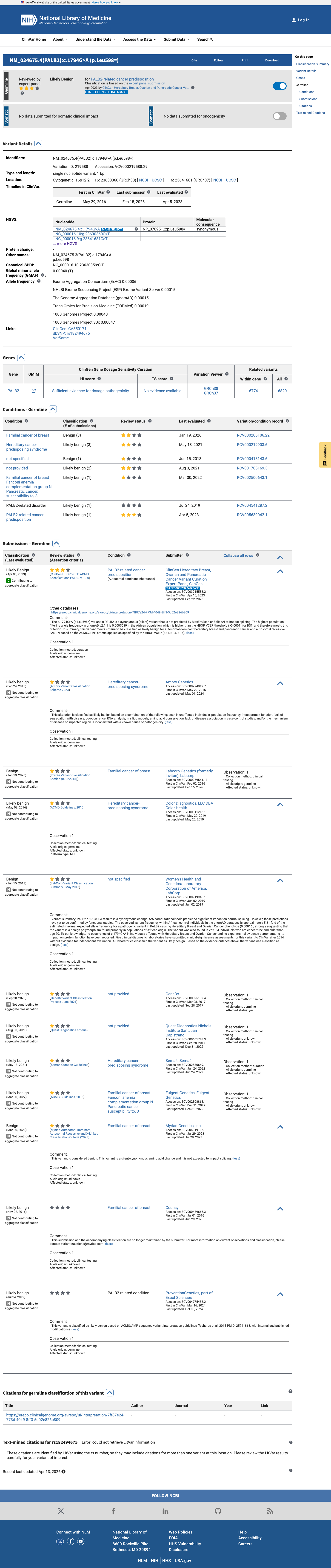
In gnomAD v4.1, this variant is present at 47/1614132 alleles (0.00291%) with a grpmax filtering allele frequency of 0.04713%, which is above the PALB2 BS1 threshold of 0.01%; gnomAD v2.1 also shows the variant in population databases, including African/African American individuals.
gnomad_v4 ↗ gnomad_v2 ↗ cspec ↗3
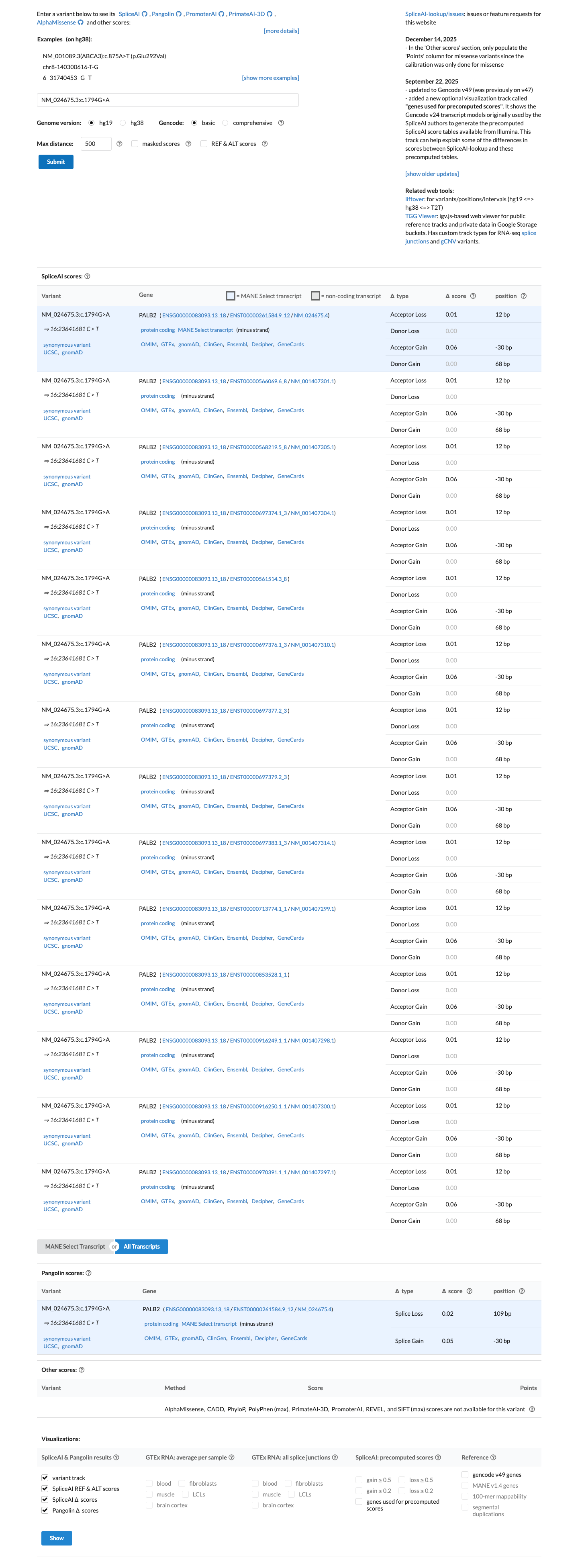
SpliceAI predicts no significant splice impact for this synonymous variant, with a maximum delta score of 0.06, which is below the PALB2 BP4 threshold of 0.1 and below the PP3 threshold of 0.2.
spliceai ↗ cspec ↗