The TP53 c.221C>T (p.Ala74Val) variant has been reported in ClinVar, including a Likely Benign classification from the ClinGen TP53 Variant Curation Expert Panel, and available hotspot review did not identify it as a recurrent TP53 hotspot.
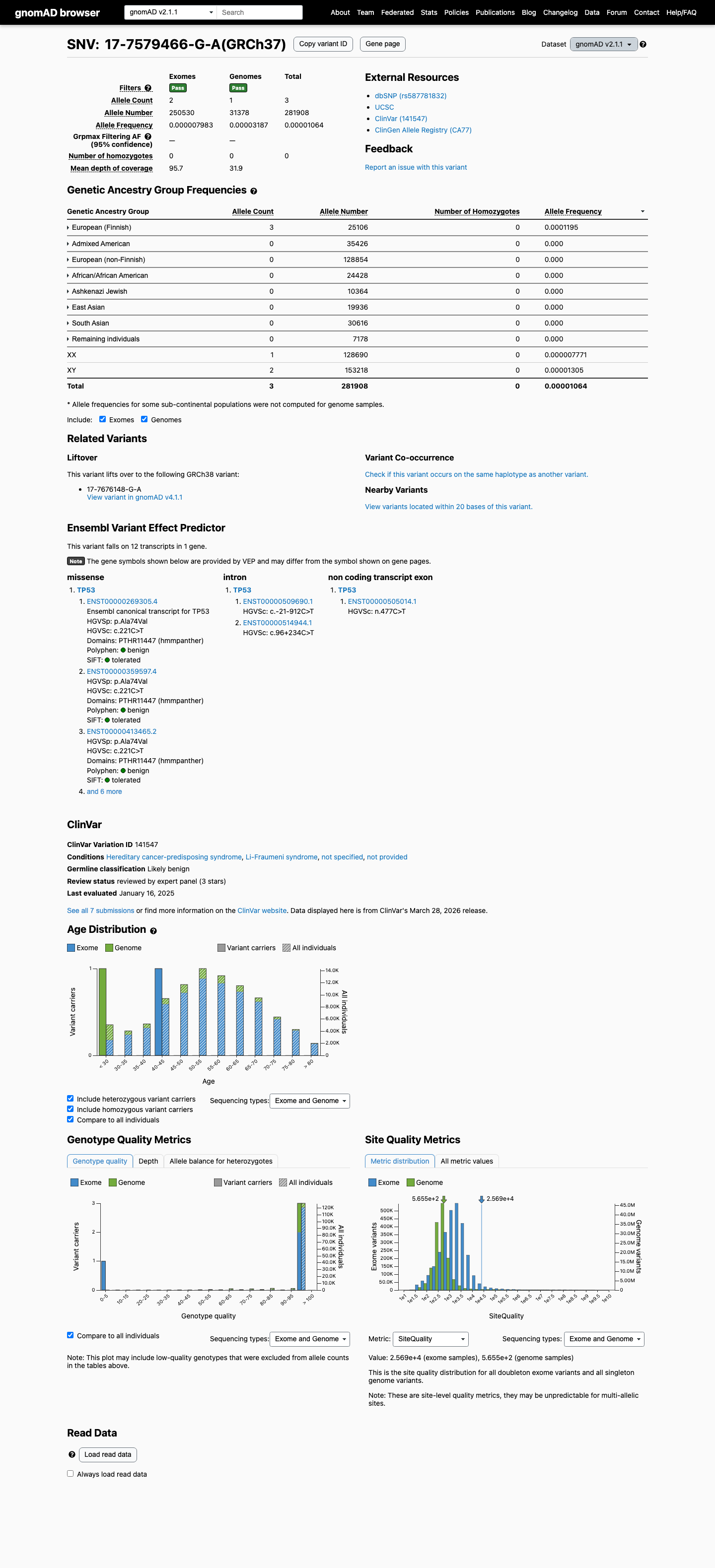
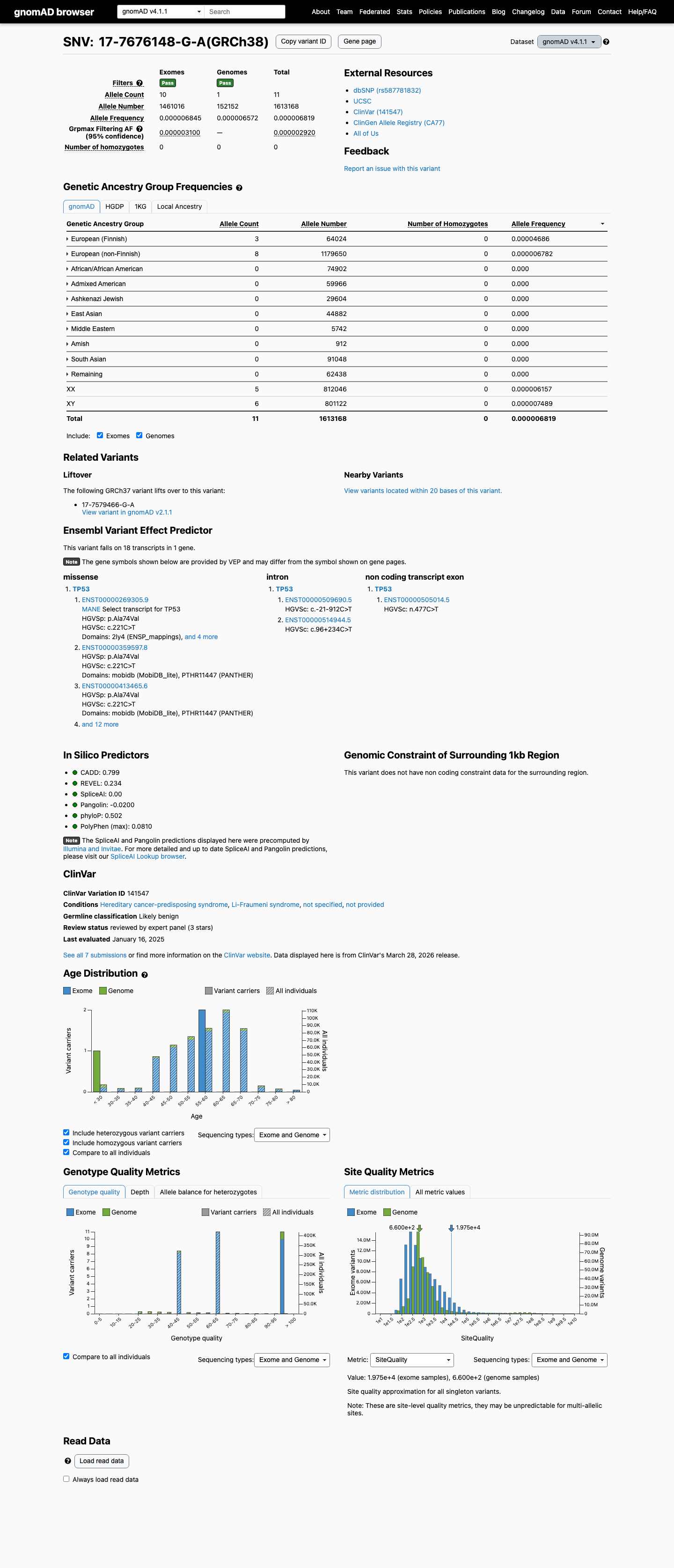
clinvar ↗ hotspots ↗This variant is rare in gnomAD v4.1 (11/1613168 alleles; AF 0.00068%; grpmax FAF 2.92e-06) and gnomAD v2.1 (3/281908 alleles; AF 0.00106%), which is below the TP53 PM2 threshold of 0.003% and below the BS1 and BA1 thresholds.
gnomad_v4 ↗ gnomad_v2 ↗ cspec ↗In TP53 functional data summarized by the TP53 VCEP, p.Ala74Val was functional and showed no loss of function in the available eligible assay summary, supporting BS3.
PMID:12826609 ↗ PMID:29979965 ↗ PMID:30224644 ↗TP53 VCEP bioinformatic calibration assigns BP4_moderate for c.221C>T; BayesDel is -0.213549, SpliceAI max delta is 0.00, and REVEL is 0.234, which together do not support a damaging computational effect.
spliceai ↗